"Leung": models, code, and papers
Local Risk Bounds for Statistical Aggregation
Jun 29, 2023In the problem of aggregation, the aim is to combine a given class of base predictors to achieve predictions nearly as accurate as the best one. In this flexible framework, no assumption is made on the structure of the class or the nature of the target. Aggregation has been studied in both sequential and statistical contexts. Despite some important differences between the two problems, the classical results in both cases feature the same global complexity measure. In this paper, we revisit and tighten classical results in the theory of aggregation in the statistical setting by replacing the global complexity with a smaller, local one. Some of our proofs build on the PAC-Bayes localization technique introduced by Catoni. Among other results, we prove localized versions of the classical bound for the exponential weights estimator due to Leung and Barron and deviation-optimal bounds for the Q-aggregation estimator. These bounds improve over the results of Dai, Rigollet and Zhang for fixed design regression and the results of Lecu\'e and Rigollet for random design regression.
Weighted regret-based likelihood: a new approach to describing uncertainty
Sep 05, 2013Recently, Halpern and Leung suggested representing uncertainty by a weighted set of probability measures, and suggested a way of making decisions based on this representation of uncertainty: maximizing weighted regret. Their paper does not answer an apparently simpler question: what it means, according to this representation of uncertainty, for an event E to be more likely than an event E'. In this paper, a notion of comparative likelihood when uncertainty is represented by a weighted set of probability measures is defined. It generalizes the ordering defined by probability (and by lower probability) in a natural way; a generalization of upper probability can also be defined. A complete axiomatic characterization of this notion of regret-based likelihood is given.
Evolutionary Algorithms: Concepts, Designs, and Applications in Bioinformatics: Evolutionary Algorithms for Bioinformatics
Aug 03, 2015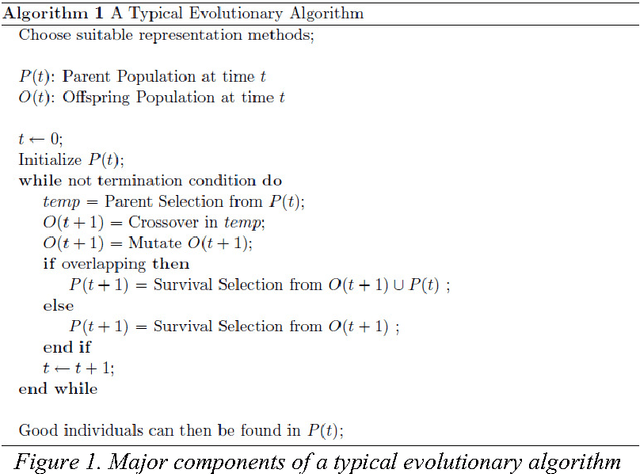
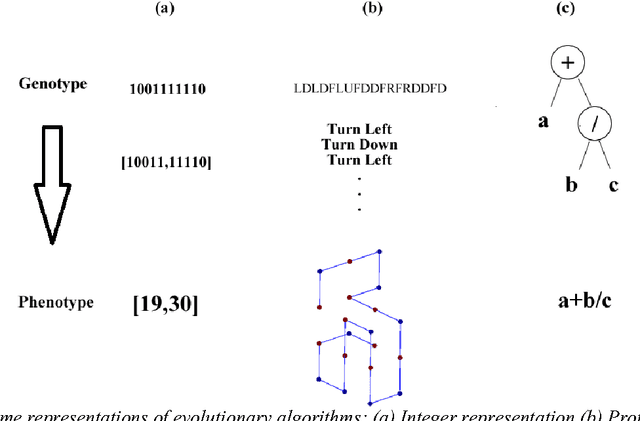
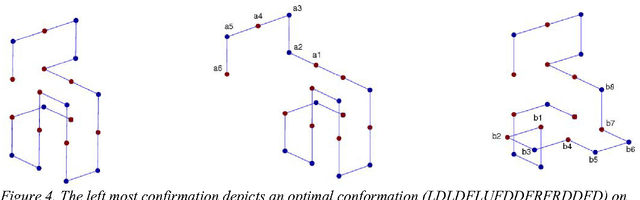
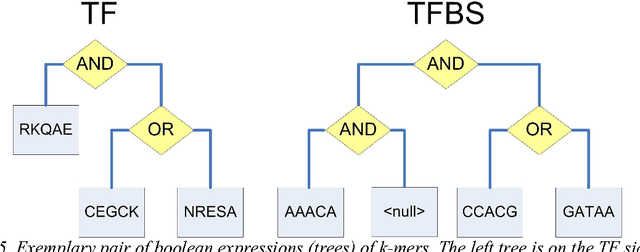
Since genetic algorithm was proposed by John Holland (Holland J. H., 1975) in the early 1970s, the study of evolutionary algorithm has emerged as a popular research field (Civicioglu & Besdok, 2013). Researchers from various scientific and engineering disciplines have been digging into this field, exploring the unique power of evolutionary algorithms (Hadka & Reed, 2013). Many applications have been successfully proposed in the past twenty years. For example, mechanical design (Lampinen & Zelinka, 1999), electromagnetic optimization (Rahmat-Samii & Michielssen, 1999), environmental protection (Bertini, Felice, Moretti, & Pizzuti, 2010), finance (Larkin & Ryan, 2010), musical orchestration (Esling, Carpentier, & Agon, 2010), pipe routing (Furuholmen, Glette, Hovin, & Torresen, 2010), and nuclear reactor core design (Sacco, Henderson, Rios-Coelho, Ali, & Pereira, 2009). In particular, its function optimization capability was highlighted (Goldberg & Richardson, 1987) because of its high adaptability to different function landscapes, to which we cannot apply traditional optimization techniques (Wong, Leung, & Wong, 2009). Here we review the applications of evolutionary algorithms in bioinformatics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge