Evaluating Large Language Models for Health-Related Text Classification Tasks with Public Social Media Data
Mar 27, 2024Yuting Guo, Anthony Ovadje, Mohammed Ali Al-Garadi, Abeed Sarker
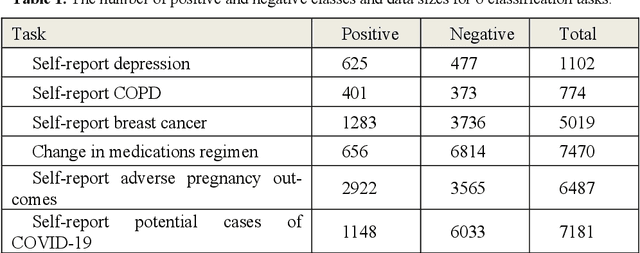
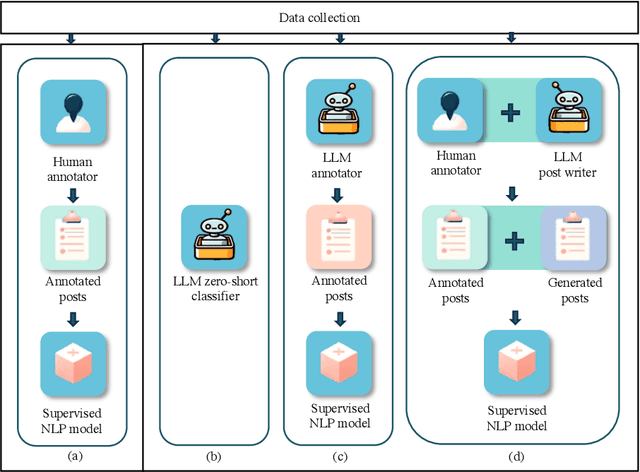
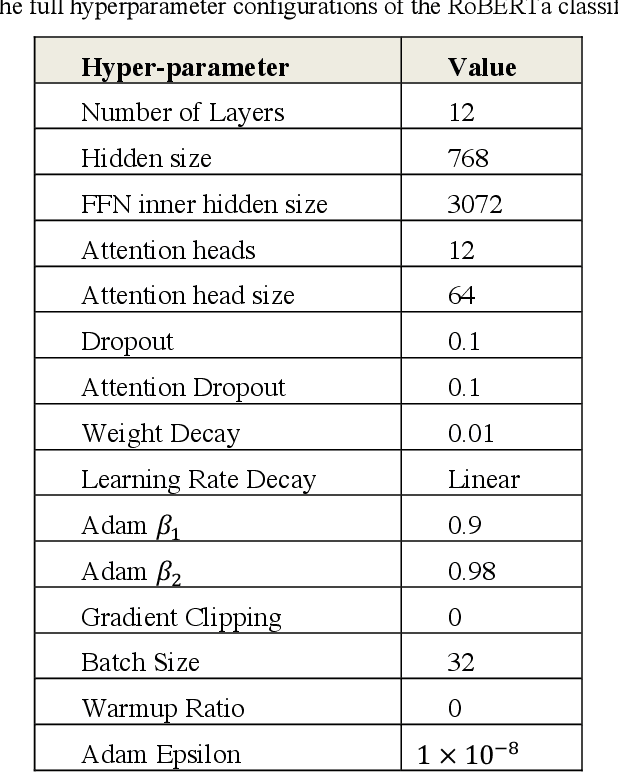
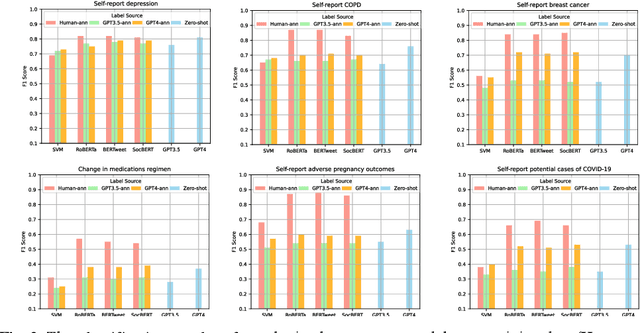
Large language models (LLMs) have demonstrated remarkable success in NLP tasks. However, there is a paucity of studies that attempt to evaluate their performances on social media-based health-related natural language processing tasks, which have traditionally been difficult to achieve high scores in. We benchmarked one supervised classic machine learning model based on Support Vector Machines (SVMs), three supervised pretrained language models (PLMs) based on RoBERTa, BERTweet, and SocBERT, and two LLM based classifiers (GPT3.5 and GPT4), across 6 text classification tasks. We developed three approaches for leveraging LLMs for text classification: employing LLMs as zero-shot classifiers, us-ing LLMs as annotators to annotate training data for supervised classifiers, and utilizing LLMs with few-shot examples for augmentation of manually annotated data. Our comprehensive experiments demonstrate that employ-ing data augmentation using LLMs (GPT-4) with relatively small human-annotated data to train lightweight supervised classification models achieves superior results compared to training with human-annotated data alone. Supervised learners also outperform GPT-4 and GPT-3.5 in zero-shot settings. By leveraging this data augmentation strategy, we can harness the power of LLMs to develop smaller, more effective domain-specific NLP models. LLM-annotated data without human guidance for training light-weight supervised classification models is an ineffective strategy. However, LLM, as a zero-shot classifier, shows promise in excluding false negatives and potentially reducing the human effort required for data annotation. Future investigations are imperative to explore optimal training data sizes and the optimal amounts of augmented data.
Social Media as a Sensor: Analyzing Twitter Data for Breast Cancer Medication Effects Using Natural Language Processing
Feb 26, 2024Seibi Kobara, Alireza Rafiei, Masoud Nateghi, Selen Bozkurt, Rishikesan Kamaleswaran, Abeed Sarker
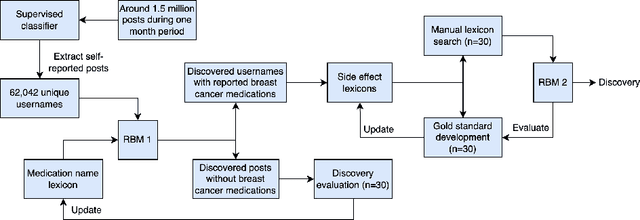
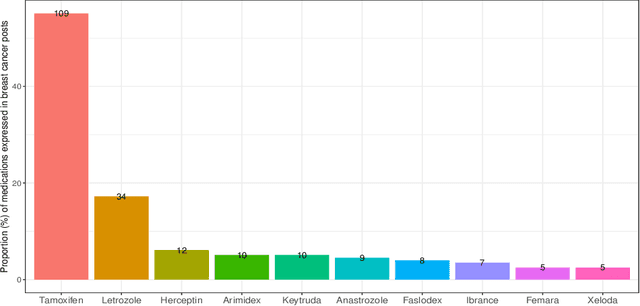
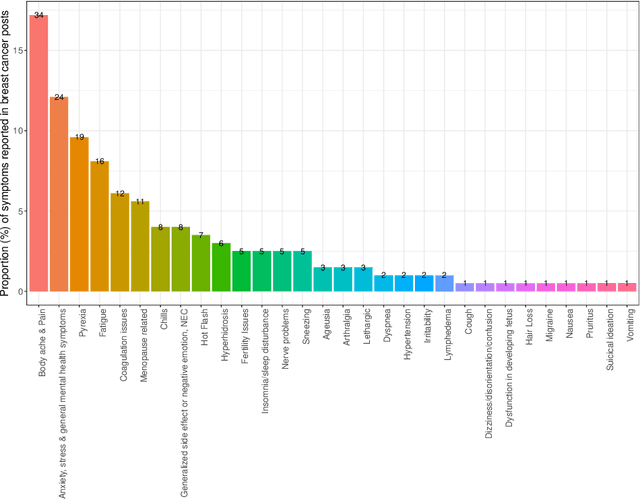
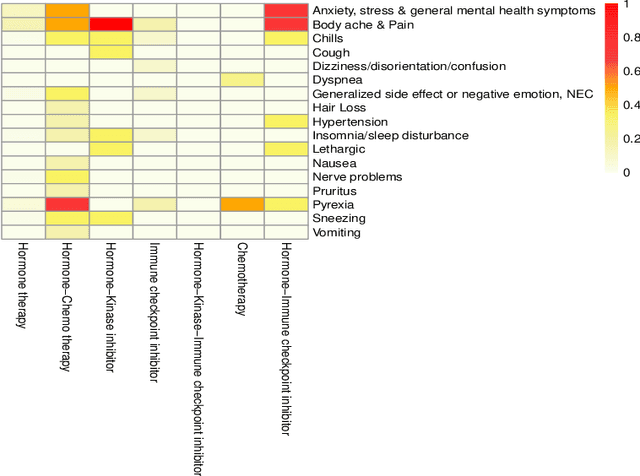
Breast cancer is a significant public health concern and is the leading cause of cancer-related deaths among women. Despite advances in breast cancer treatments, medication non-adherence remains a major problem. As electronic health records do not typically capture patient-reported outcomes that may reveal information about medication-related experiences, social media presents an attractive resource for enhancing our understanding of the patients' treatment experiences. In this paper, we developed natural language processing (NLP) based methodologies to study information posted by an automatically curated breast cancer cohort from social media. We employed a transformer-based classifier to identify breast cancer patients/survivors on X (Twitter) based on their self-reported information, and we collected longitudinal data from their profiles. We then designed a multi-layer rule-based model to develop a breast cancer therapy-associated side effect lexicon and detect patterns of medication usage and associated side effects among breast cancer patients. 1,454,637 posts were available from 583,962 unique users, of which 62,042 were detected as breast cancer members using our transformer-based model. 198 cohort members mentioned breast cancer medications with tamoxifen as the most common. Our side effect lexicon identified well-known side effects of hormone and chemotherapy. Furthermore, it discovered a subject feeling towards cancer and medications, which may suggest a pre-clinical phase of side effects or emotional distress. This analysis highlighted not only the utility of NLP techniques in unstructured social media data to identify self-reported breast cancer posts, medication usage patterns, and treatment side effects but also the richness of social data on such clinical questions.
Learning from Two Decades of Blood Pressure Data: Demography-Specific Patterns Across 75 Million Patient Encounters
Feb 05, 2024Seyedeh Somayyeh Mousavi, Yuting Guo, Abeed Sarker, Reza Sameni
Hypertension remains a global health concern with a rising prevalence, necessitating effective monitoring and understanding of blood pressure (BP) dynamics. This study delves into the wealth of information derived from BP measurement, a crucial approach in informing our understanding of hypertensive trends. Numerous studies have reported on the relationship between BP variation and various factors. In this research, we leveraged an extensive dataset comprising 75 million records spanning two decades, offering a unique opportunity to explore and analyze BP variations across demographic features such as age, race, and gender. Our findings revealed that gender-based BP variation was not statistically significant, challenging conventional assumptions. Interestingly, systolic blood pressure (SBP) consistently increased with age, while diastolic blood pressure (DBP) displayed a distinctive peak in the forties age group. Moreover, our analysis uncovered intriguing similarities in the distribution of BP among some of the racial groups. This comprehensive investigation contributes to the ongoing discourse on hypertension and underscores the importance of considering diverse demographic factors in understanding BP variations. Our results provide valuable insights that may inform personalized healthcare approaches tailored to specific demographic profiles.
Leveraging Large Language Models for Analyzing Blood Pressure Variations Across Biological Sex from Scientific Literature
Feb 02, 2024Yuting Guo, Seyedeh Somayyeh Mousavi, Reza Sameni, Abeed Sarker
Hypertension, defined as blood pressure (BP) that is above normal, holds paramount significance in the realm of public health, as it serves as a critical precursor to various cardiovascular diseases (CVDs) and significantly contributes to elevated mortality rates worldwide. However, many existing BP measurement technologies and standards might be biased because they do not consider clinical outcomes, comorbidities, or demographic factors, making them inconclusive for diagnostic purposes. There is limited data-driven research focused on studying the variance in BP measurements across these variables. In this work, we employed GPT-35-turbo, a large language model (LLM), to automatically extract the mean and standard deviation values of BP for both males and females from a dataset comprising 25 million abstracts sourced from PubMed. 993 article abstracts met our predefined inclusion criteria (i.e., presence of references to blood pressure, units of blood pressure such as mmHg, and mention of biological sex). Based on the automatically-extracted information from these articles, we conducted an analysis of the variations of BP values across biological sex. Our results showed the viability of utilizing LLMs to study the BP variations across different demographic factors.
Generalizable Natural Language Processing Framework for Migraine Reporting from Social Media
Dec 23, 2022Yuting Guo, Swati Rajwal, Sahithi Lakamana, Chia-Chun Chiang, Paul C. Menell, Adnan H. Shahid, Yi-Chieh Chen, Nikita Chhabra, Wan-Ju Chao, Chieh-Ju Chao, Todd J. Schwedt, Imon Banerjee, Abeed Sarker
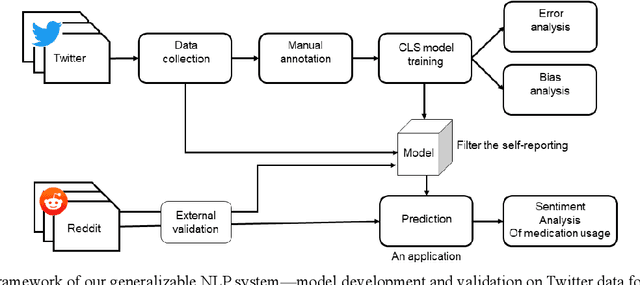
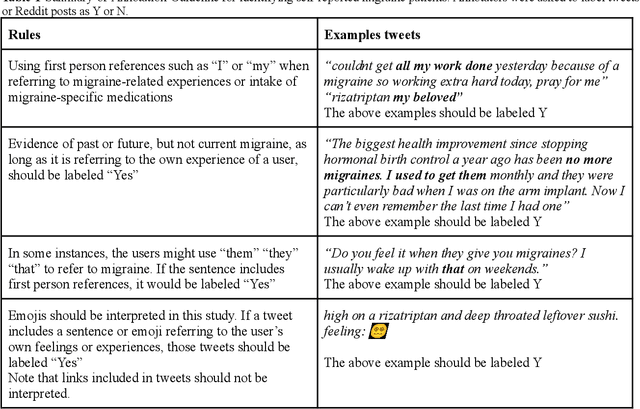
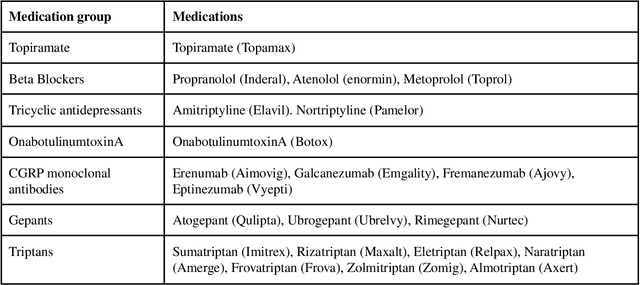
Migraine is a high-prevalence and disabling neurological disorder. However, information migraine management in real-world settings could be limited to traditional health information sources. In this paper, we (i) verify that there is substantial migraine-related chatter available on social media (Twitter and Reddit), self-reported by migraine sufferers; (ii) develop a platform-independent text classification system for automatically detecting self-reported migraine-related posts, and (iii) conduct analyses of the self-reported posts to assess the utility of social media for studying this problem. We manually annotated 5750 Twitter posts and 302 Reddit posts. Our system achieved an F1 score of 0.90 on Twitter and 0.93 on Reddit. Analysis of information posted by our 'migraine cohort' revealed the presence of a plethora of relevant information about migraine therapies and patient sentiments associated with them. Our study forms the foundation for conducting an in-depth analysis of migraine-related information using social media data.
Social media mining for toxicovigilance of prescription medications: End-to-end pipeline, challenges and future work
Nov 18, 2022Abeed Sarker
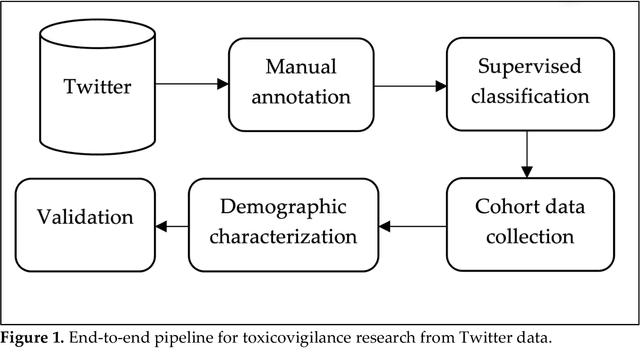
Substance use, substance use disorder, and overdoses related to substance use are major public health problems globally and in the United States. A key aspect of addressing these problems from a public health standpoint is improved surveillance. Traditional surveillance systems are laggy, and social media are potentially useful sources of timely data. However, mining knowledge from social media is challenging, and requires the development of advanced artificial intelligence, specifically natural language processing (NLP) and machine learning methods. We developed a sophisticated end-to-end pipeline for mining information about nonmedical prescription medication use from social media, namely Twitter and Reddit. Our pipeline employs supervised machine learning and NLP for filtering out noise and characterizing the chatter. In this paper, we describe our end-to-end pipeline developed over four years. In addition to describing our data mining infrastructure, we discuss existing challenges in social media mining for toxicovigilance, and possible future research directions.
Few-shot learning for medical text: A systematic review
Apr 21, 2022Yao Ge, Yuting Guo, Yuan-Chi Yang, Mohammed Ali Al-Garadi, Abeed Sarker
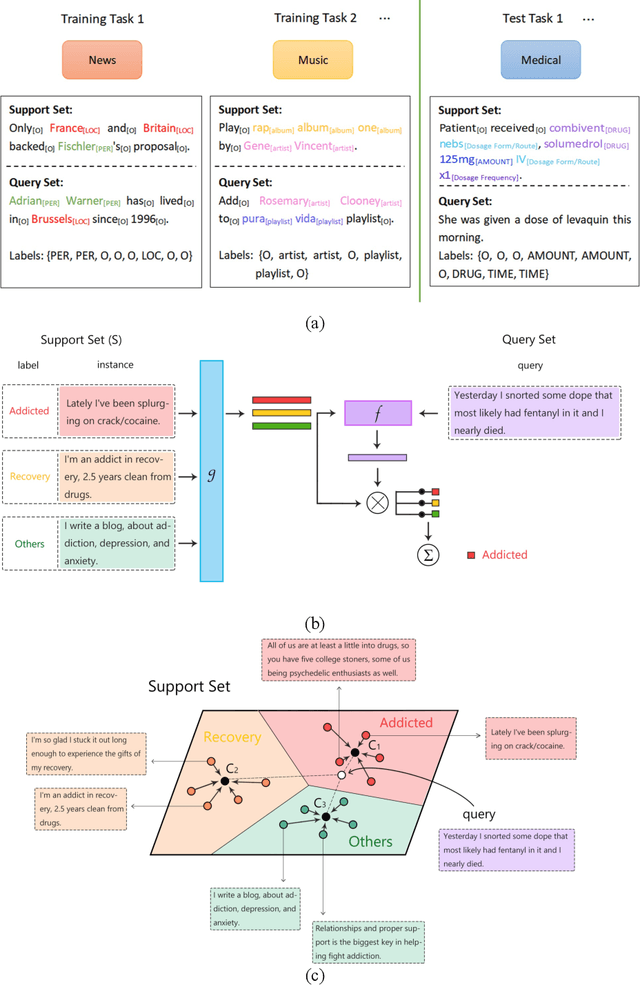
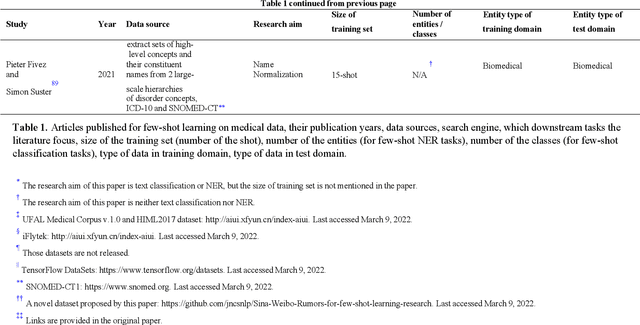
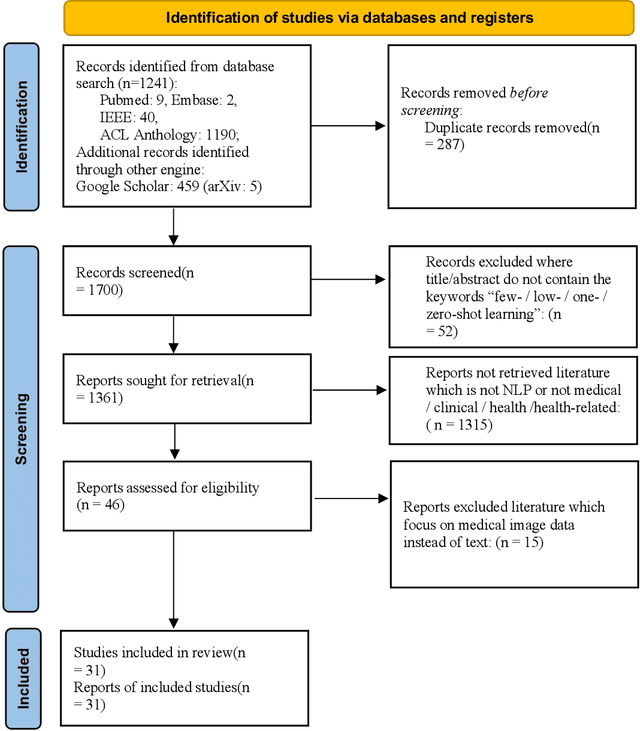
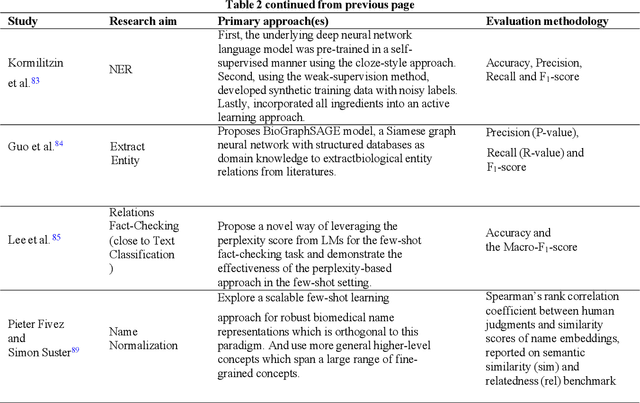
Objective: Few-shot learning (FSL) methods require small numbers of labeled instances for training. As many medical topics have limited annotated textual data in practical settings, FSL-based natural language processing (NLP) methods hold substantial promise. We aimed to conduct a systematic review to explore the state of FSL methods for medical NLP. Materials and Methods: We searched for articles published between January 2016 and August 2021 using PubMed/Medline, Embase, ACL Anthology, and IEEE Xplore Digital Library. To identify the latest relevant methods, we also searched other sources such as preprint servers (eg., medRxiv) via Google Scholar. We included all articles that involved FSL and any type of medical text. We abstracted articles based on data source(s), aim(s), training set size(s), primary method(s)/approach(es), and evaluation method(s). Results: 31 studies met our inclusion criteria-all published after 2018; 22 (71%) since 2020. Concept extraction/named entity recognition was the most frequently addressed task (13/31; 42%), followed by text classification (10/31; 32%). Twenty-one (68%) studies reconstructed existing datasets to create few-shot scenarios synthetically, and MIMIC-III was the most frequently used dataset (7/31; 23%). Common methods included FSL with attention mechanisms (12/31; 39%), prototypical networks (8/31; 26%), and meta-learning (6/31; 19%). Discussion: Despite the potential for FSL in biomedical NLP, progress has been limited compared to domain-independent FSL. This may be due to the paucity of standardized, public datasets, and the relative underperformance of FSL methods on biomedical topics. Creation and release of specialized datasets for biomedical FSL may aid method development by enabling comparative analyses.
Towards Automatic Bot Detection in Twitter for Health-related Tasks
Sep 29, 2019Anahita Davoudi, Ari Z. Klein, Abeed Sarker, Graciela Gonzalez-Hernandez
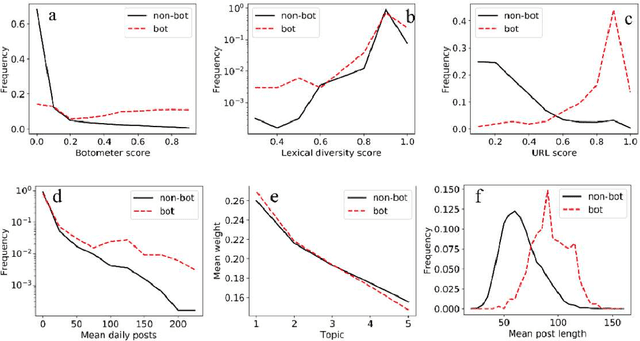

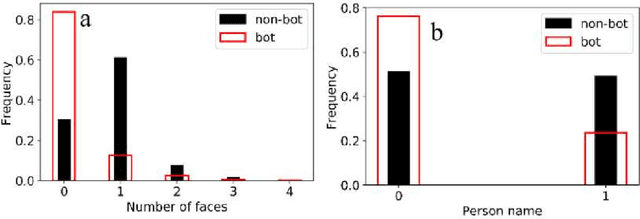
With the increasing use of social media data for health-related research, the credibility of the information from this source has been questioned as the posts may originate from automated accounts or "bots". While automatic bot detection approaches have been proposed, there are none that have been evaluated on users posting health-related information. In this paper, we extend an existing bot detection system and customize it for health-related research. Using a dataset of Twitter users, we first show that the system, which was designed for political bot detection, underperforms when applied to health-related Twitter users. We then incorporate additional features and a statistical machine learning classifier to significantly improve bot detection performance. Our approach obtains F_1 scores of 0.7 for the "bot" class, representing improvements of 0.339. Our approach is customizable and generalizable for bot detection in other health-related social media cohorts.
Deep Neural Networks Ensemble for Detecting Medication Mentions in Tweets
Apr 10, 2019Davy Weissenbacher, Abeed Sarker, Ari Klein, Karen O'Connor, Arjun Magge Ranganatha, Graciela Gonzalez-Hernandez
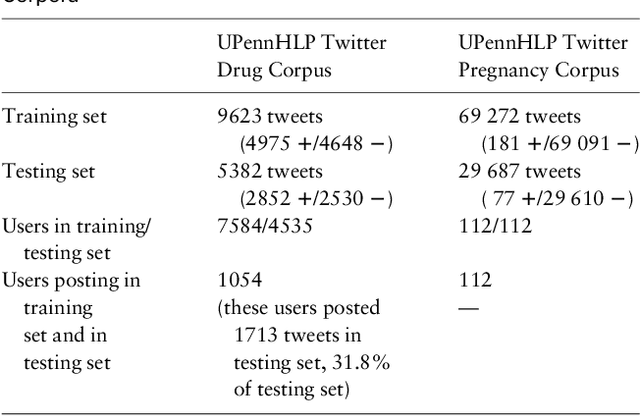
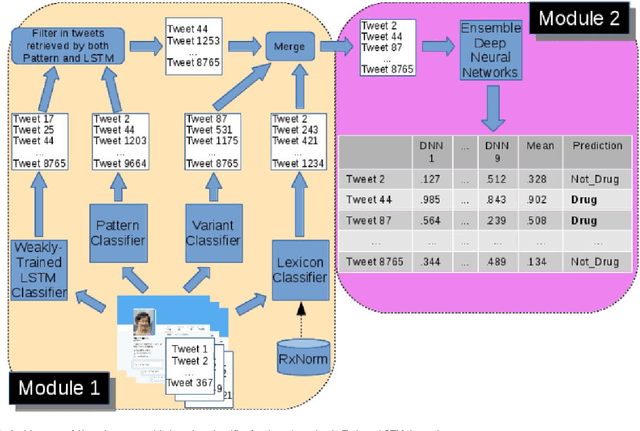
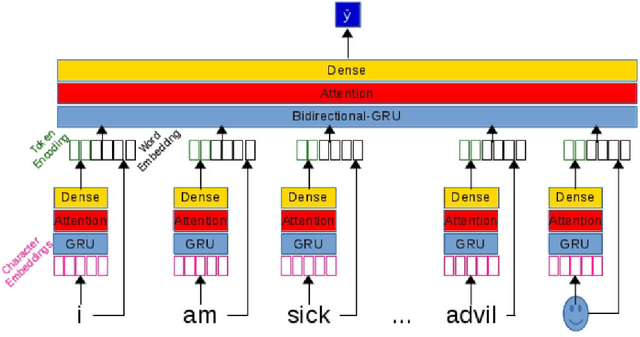
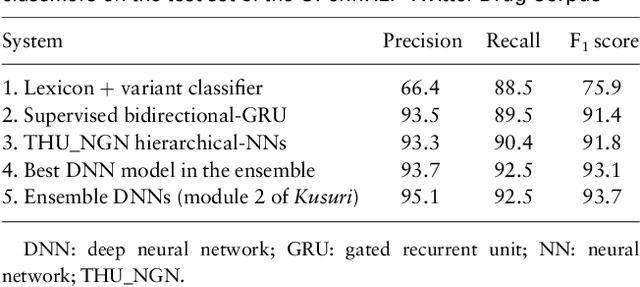
Objective: After years of research, Twitter posts are now recognized as an important source of patient-generated data, providing unique insights into population health. A fundamental step to incorporating Twitter data in pharmacoepidemiological research is to automatically recognize medication mentions in tweets. Given that lexical searches for medication names may fail due to misspellings or ambiguity with common words, we propose a more advanced method to recognize them. Methods: We present Kusuri, an Ensemble Learning classifier, able to identify tweets mentioning drug products and dietary supplements. Kusuri ("medication" in Japanese) is composed of two modules. First, four different classifiers (lexicon-based, spelling-variant-based, pattern-based and one based on a weakly-trained neural network) are applied in parallel to discover tweets potentially containing medication names. Second, an ensemble of deep neural networks encoding morphological, semantical and long-range dependencies of important words in the tweets discovered is used to make the final decision. Results: On a balanced (50-50) corpus of 15,005 tweets, Kusuri demonstrated performances close to human annotators with 93.7% F1-score, the best score achieved thus far on this corpus. On a corpus made of all tweets posted by 113 Twitter users (98,959 tweets, with only 0.26% mentioning medications), Kusuri obtained 76.3% F1-score. There is not a prior drug extraction system that compares running on such an extremely unbalanced dataset. Conclusion: The system identifies tweets mentioning drug names with performance high enough to ensure its usefulness and ready to be integrated in larger natural language processing systems.
Automatically Detecting Self-Reported Birth Defect Outcomes on Twitter for Large-scale Epidemiological Research
Oct 22, 2018Ari Z. Klein, Abeed Sarker, Davy Weissenbacher, Graciela Gonzalez-Hernandez
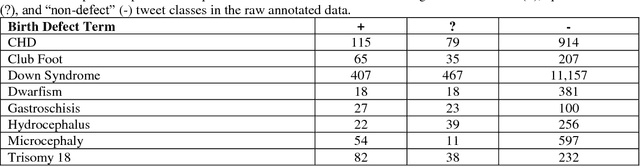
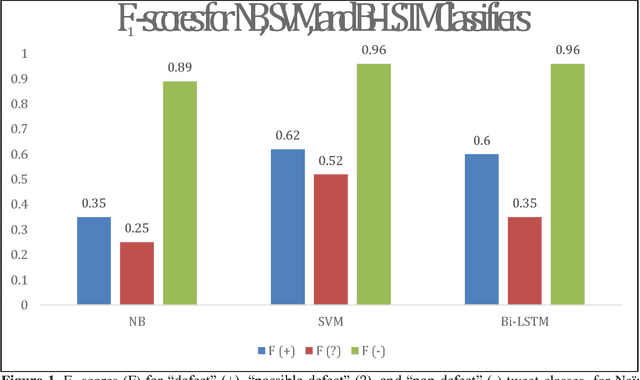
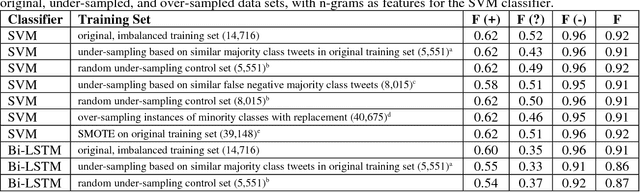

In recent work, we identified and studied a small cohort of Twitter users whose pregnancies with birth defect outcomes could be observed via their publicly available tweets. Exploiting social media's large-scale potential to complement the limited methods for studying birth defects, the leading cause of infant mortality, depends on the further development of automatic methods. The primary objective of this study was to take the first step towards scaling the use of social media for observing pregnancies with birth defect outcomes, namely, developing methods for automatically detecting tweets by users reporting their birth defect outcomes. We annotated and pre-processed approximately 23,000 tweets that mention birth defects in order to train and evaluate supervised machine learning algorithms, including feature-engineered and deep learning-based classifiers. We also experimented with various under-sampling and over-sampling approaches to address the class imbalance. A Support Vector Machine (SVM) classifier trained on the original, imbalanced data set, with n-grams, word clusters, and structural features, achieved the best baseline performance for the positive classes: an F1-score of 0.65 for the "defect" class and 0.51 for the "possible defect" class. Our contributions include (i) natural language processing (NLP) and supervised machine learning methods for automatically detecting tweets by users reporting their birth defect outcomes, (ii) a comparison of feature-engineered and deep learning-based classifiers trained on imbalanced, under-sampled, and over-sampled data, and (iii) an error analysis that could inform classification improvements using our publicly available corpus. Future work will focus on automating user-level analyses for cohort inclusion.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge