Multilingual transformer and BERTopic for short text topic modeling: The case of Serbian
Feb 05, 2024Darija Medvecki, Bojana Bašaragin, Adela Ljajić, Nikola Milošević
This paper presents the results of the first application of BERTopic, a state-of-the-art topic modeling technique, to short text written in a morphologi-cally rich language. We applied BERTopic with three multilingual embed-ding models on two levels of text preprocessing (partial and full) to evalu-ate its performance on partially preprocessed short text in Serbian. We also compared it to LDA and NMF on fully preprocessed text. The experiments were conducted on a dataset of tweets expressing hesitancy toward COVID-19 vaccination. Our results show that with adequate parameter setting, BERTopic can yield informative topics even when applied to partially pre-processed short text. When the same parameters are applied in both prepro-cessing scenarios, the performance drop on partially preprocessed text is minimal. Compared to LDA and NMF, judging by the keywords, BERTopic offers more informative topics and gives novel insights when the number of topics is not limited. The findings of this paper can be significant for re-searchers working with other morphologically rich low-resource languages and short text.
A transformer-based method for zero and few-shot biomedical named entity recognition
May 12, 2023Miloš Košprdić, Nikola Prodanović, Adela Ljajić, Bojana Bašaragin, Nikola Milošević
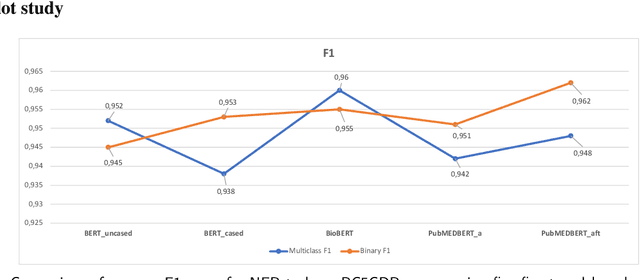
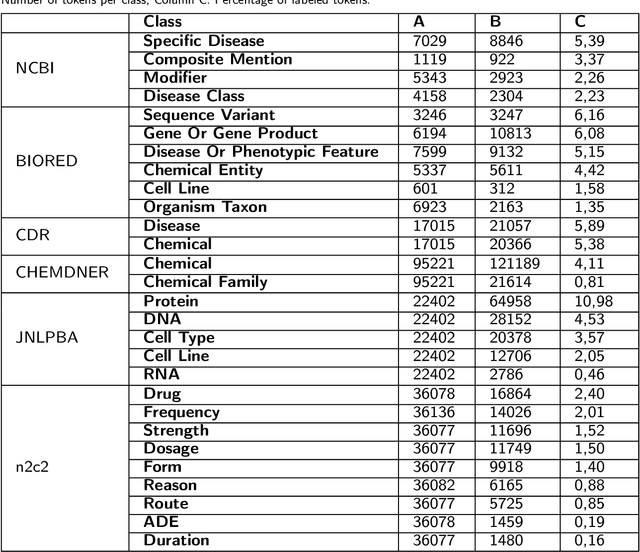

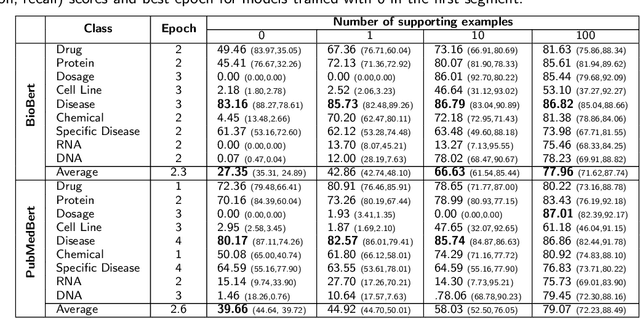
Supervised named entity recognition (NER) in the biomedical domain is dependent on large sets of annotated texts with the given named entities, whose creation can be time-consuming and expensive. Furthermore, the extraction of new entities often requires conducting additional annotation tasks and retraining the model. To address these challenges, this paper proposes a transformer-based method for zero- and few-shot NER in the biomedical domain. The method is based on transforming the task of multi-class token classification into binary token classification (token contains the searched entity or does not contain the searched entity) and pre-training on a larger amount of datasets and biomedical entities, from where the method can learn semantic relations between the given and potential classes. We have achieved average F1 scores of 35.44% for zero-shot NER, 50.10% for one-shot NER, 69.94% for 10-shot NER, and 79.51% for 100-shot NER on 9 diverse evaluated biomedical entities with PubMedBERT fine-tuned model. The results demonstrate the effectiveness of the proposed method for recognizing new entities with limited examples, with comparable or better results from the state-of-the-art zero- and few-shot NER methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge