Duality of Bures and Shape Distances with Implications for Comparing Neural Representations
Nov 19, 2023Sarah E. Harvey, Brett W. Larsen, Alex H. Williams
A multitude of (dis)similarity measures between neural network representations have been proposed, resulting in a fragmented research landscape. Most of these measures fall into one of two categories. First, measures such as linear regression, canonical correlations analysis (CCA), and shape distances, all learn explicit mappings between neural units to quantify similarity while accounting for expected invariances. Second, measures such as representational similarity analysis (RSA), centered kernel alignment (CKA), and normalized Bures similarity (NBS) all quantify similarity in summary statistics, such as stimulus-by-stimulus kernel matrices, which are already invariant to expected symmetries. Here, we take steps towards unifying these two broad categories of methods by observing that the cosine of the Riemannian shape distance (from category 1) is equal to NBS (from category 2). We explore how this connection leads to new interpretations of shape distances and NBS, and draw contrasts of these measures with CKA, a popular similarity measure in the deep learning literature.
Soft Matching Distance: A metric on neural representations that captures single-neuron tuning
Nov 16, 2023Meenakshi Khosla, Alex H. Williams
Common measures of neural representational (dis)similarity are designed to be insensitive to rotations and reflections of the neural activation space. Motivated by the premise that the tuning of individual units may be important, there has been recent interest in developing stricter notions of representational (dis)similarity that require neurons to be individually matched across networks. When two networks have the same size (i.e. same number of neurons), a distance metric can be formulated by optimizing over neuron index permutations to maximize tuning curve alignment. However, it is not clear how to generalize this metric to measure distances between networks with different sizes. Here, we leverage a connection to optimal transport theory to derive a natural generalization based on "soft" permutations. The resulting metric is symmetric, satisfies the triangle inequality, and can be interpreted as a Wasserstein distance between two empirical distributions. Further, our proposed metric avoids counter-intuitive outcomes suffered by alternative approaches, and captures complementary geometric insights into neural representations that are entirely missed by rotation-invariant metrics.
Estimating Shape Distances on Neural Representations with Limited Samples
Oct 09, 2023Dean A. Pospisil, Brett W. Larsen, Sarah E. Harvey, Alex H. Williams
Measuring geometric similarity between high-dimensional network representations is a topic of longstanding interest to neuroscience and deep learning. Although many methods have been proposed, only a few works have rigorously analyzed their statistical efficiency or quantified estimator uncertainty in data-limited regimes. Here, we derive upper and lower bounds on the worst-case convergence of standard estimators of shape distance$\unicode{x2014}$a measure of representational dissimilarity proposed by Williams et al. (2021). These bounds reveal the challenging nature of the problem in high-dimensional feature spaces. To overcome these challenges, we introduce a new method-of-moments estimator with a tunable bias-variance tradeoff. We show that this estimator achieves superior performance to standard estimators in simulation and on neural data, particularly in high-dimensional settings. Thus, we lay the foundation for a rigorous statistical theory for high-dimensional shape analysis, and we contribute a new estimation method that is well-suited to practical scientific settings.
Representational dissimilarity metric spaces for stochastic neural networks
Nov 21, 2022Lyndon R. Duong, Jingyang Zhou, Josue Nassar, Jules Berman, Jeroen Olieslagers, Alex H. Williams

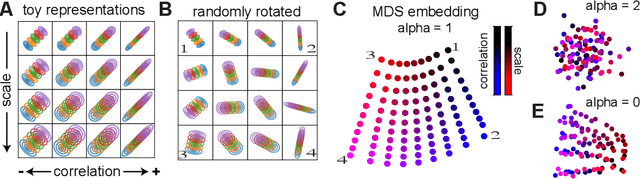

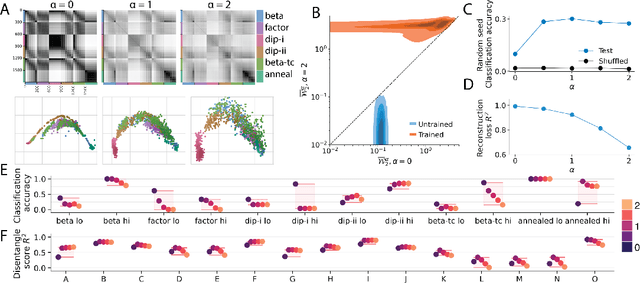
Quantifying similarity between neural representations -- e.g. hidden layer activation vectors -- is a perennial problem in deep learning and neuroscience research. Existing methods compare deterministic responses (e.g. artificial networks that lack stochastic layers) or averaged responses (e.g., trial-averaged firing rates in biological data). However, these measures of deterministic representational similarity ignore the scale and geometric structure of noise, both of which play important roles in neural computation. To rectify this, we generalize previously proposed shape metrics (Williams et al. 2021) to quantify differences in stochastic representations. These new distances satisfy the triangle inequality, and thus can be used as a rigorous basis for many supervised and unsupervised analyses. Leveraging this novel framework, we find that the stochastic geometries of neurobiological representations of oriented visual gratings and naturalistic scenes respectively resemble untrained and trained deep network representations. Further, we are able to more accurately predict certain network attributes (e.g. training hyperparameters) from its position in stochastic (versus deterministic) shape space.
Spatiotemporal Clustering with Neyman-Scott Processes via Connections to Bayesian Nonparametric Mixture Models
Jan 14, 2022Yixin Wang, Anthony Degleris, Alex H. Williams, Scott W. Linderman
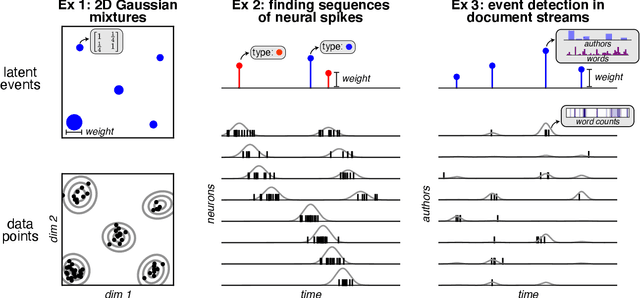

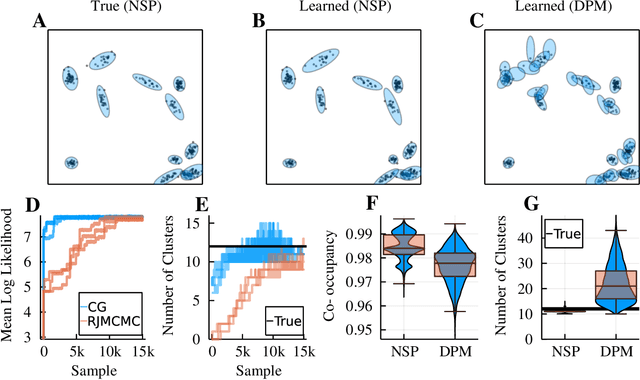
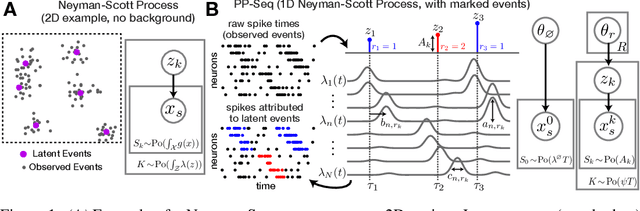
Neyman-Scott processes (NSPs) are point process models that generate clusters of points in time or space. They are natural models for a wide range of phenomena, ranging from neural spike trains to document streams. The clustering property is achieved via a doubly stochastic formulation: first, a set of latent events is drawn from a Poisson process; then, each latent event generates a set of observed data points according to another Poisson process. This construction is similar to Bayesian nonparametric mixture models like the Dirichlet process mixture model (DPMM) in that the number of latent events (i.e. clusters) is a random variable, but the point process formulation makes the NSP especially well suited to modeling spatiotemporal data. While many specialized algorithms have been developed for DPMMs, comparatively fewer works have focused on inference in NSPs. Here, we present novel connections between NSPs and DPMMs, with the key link being a third class of Bayesian mixture models called mixture of finite mixture models (MFMMs). Leveraging this connection, we adapt the standard collapsed Gibbs sampling algorithm for DPMMs to enable scalable Bayesian inference on NSP models. We demonstrate the potential of Neyman-Scott processes on a variety of applications including sequence detection in neural spike trains and event detection in document streams.
Generalized Shape Metrics on Neural Representations
Oct 27, 2021Alex H. Williams, Erin Kunz, Simon Kornblith, Scott W. Linderman
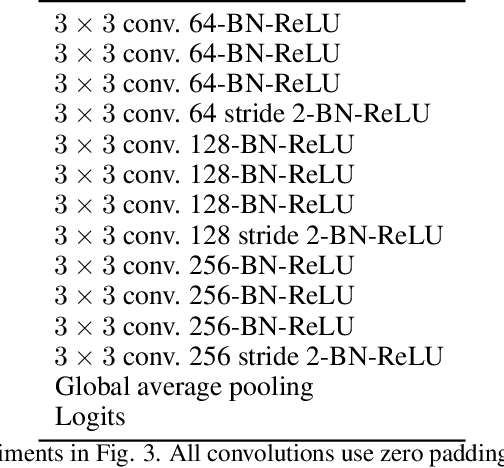
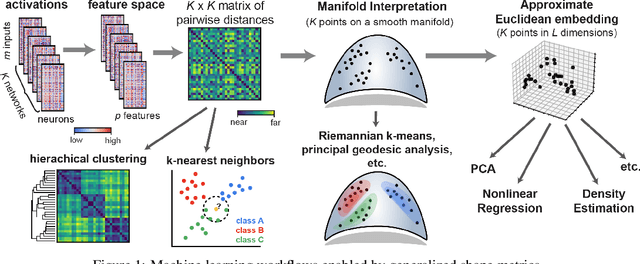


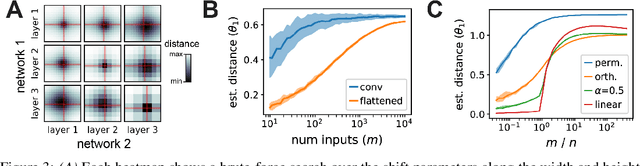
Understanding the operation of biological and artificial networks remains a difficult and important challenge. To identify general principles, researchers are increasingly interested in surveying large collections of networks that are trained on, or biologically adapted to, similar tasks. A standardized set of analysis tools is now needed to identify how network-level covariates -- such as architecture, anatomical brain region, and model organism -- impact neural representations (hidden layer activations). Here, we provide a rigorous foundation for these analyses by defining a broad family of metric spaces that quantify representational dissimilarity. Using this framework we modify existing representational similarity measures based on canonical correlation analysis to satisfy the triangle inequality, formulate a novel metric that respects the inductive biases in convolutional layers, and identify approximate Euclidean embeddings that enable network representations to be incorporated into essentially any off-the-shelf machine learning method. We demonstrate these methods on large-scale datasets from biology (Allen Institute Brain Observatory) and deep learning (NAS-Bench-101). In doing so, we identify relationships between neural representations that are interpretable in terms of anatomical features and model performance.
Point process models for sequence detection in high-dimensional neural spike trains
Oct 10, 2020Alex H. Williams, Anthony Degleris, Yixin Wang, Scott W. Linderman
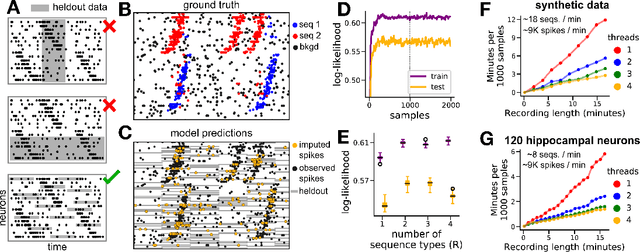
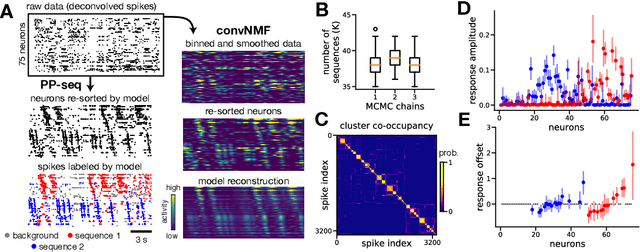
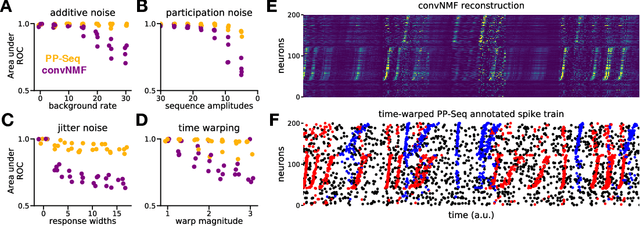
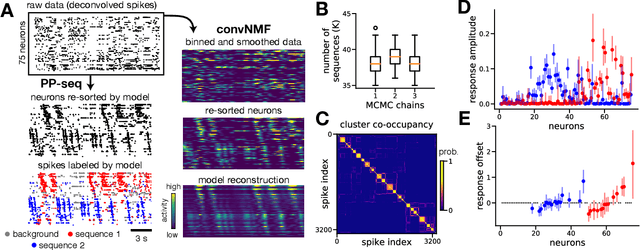
Sparse sequences of neural spikes are posited to underlie aspects of working memory, motor production, and learning. Discovering these sequences in an unsupervised manner is a longstanding problem in statistical neuroscience. Promising recent work utilized a convolutive nonnegative matrix factorization model to tackle this challenge. However, this model requires spike times to be discretized, utilizes a sub-optimal least-squares criterion, and does not provide uncertainty estimates for model predictions or estimated parameters. We address each of these shortcomings by developing a point process model that characterizes fine-scale sequences at the level of individual spikes and represents sequence occurrences as a small number of marked events in continuous time. This ultra-sparse representation of sequence events opens new possibilities for spike train modeling. For example, we introduce learnable time warping parameters to model sequences of varying duration, which have been experimentally observed in neural circuits. We demonstrate these advantages on experimental recordings from songbird higher vocal center and rodent hippocampus.
Universality and individuality in neural dynamics across large populations of recurrent networks
Jul 19, 2019Niru Maheswaranathan, Alex H. Williams, Matthew D. Golub, Surya Ganguli, David Sussillo
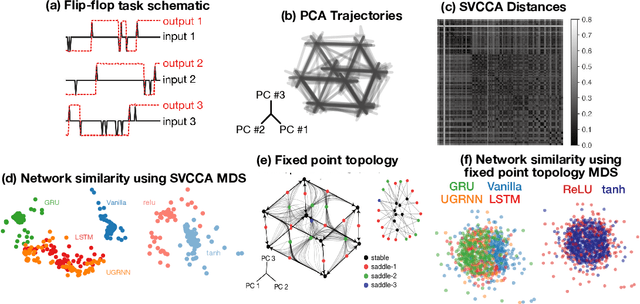
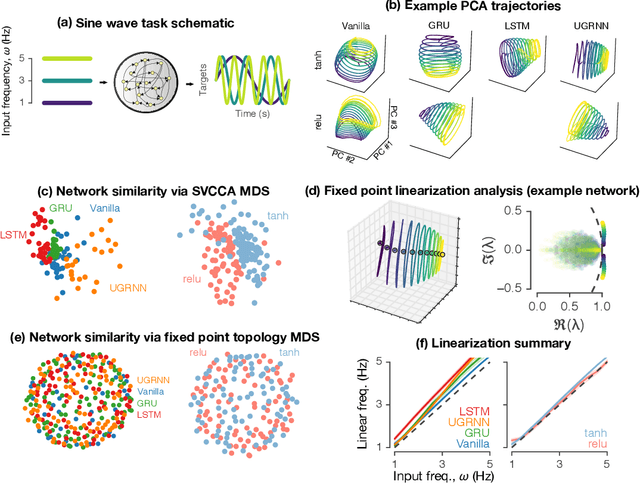
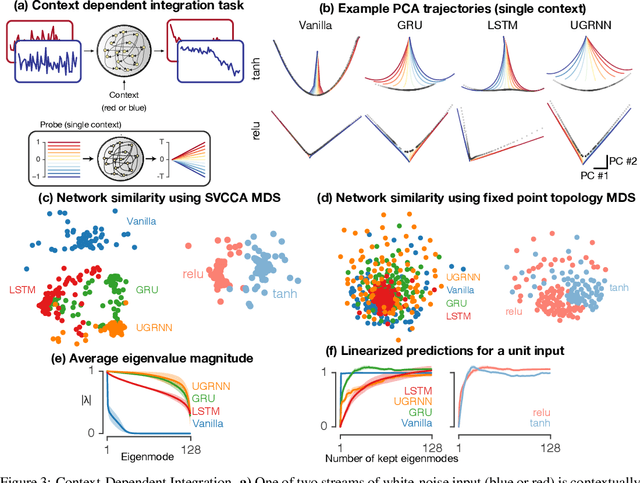
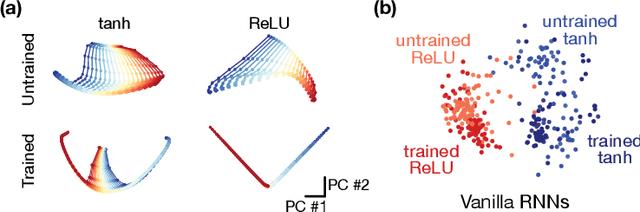
Task-based modeling with recurrent neural networks (RNNs) has emerged as a popular way to infer the computational function of different brain regions. These models are quantitatively assessed by comparing the low-dimensional neural representations of the model with the brain, for example using canonical correlation analysis (CCA). However, the nature of the detailed neurobiological inferences one can draw from such efforts remains elusive. For example, to what extent does training neural networks to solve common tasks uniquely determine the network dynamics, independent of modeling architectural choices? Or alternatively, are the learned dynamics highly sensitive to different model choices? Knowing the answer to these questions has strong implications for whether and how we should use task-based RNN modeling to understand brain dynamics. To address these foundational questions, we study populations of thousands of networks, with commonly used RNN architectures, trained to solve neuroscientifically motivated tasks and characterize their nonlinear dynamics. We find the geometry of the RNN representations can be highly sensitive to different network architectures, yielding a cautionary tale for measures of similarity that rely representational geometry, such as CCA. Moreover, we find that while the geometry of neural dynamics can vary greatly across architectures, the underlying computational scaffold---the topological structure of fixed points, transitions between them, limit cycles, and linearized dynamics---often appears universal across all architectures.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge