Deep Learning for Cancer Prognosis Prediction Using Portrait Photos by StyleGAN Embedding
Jun 28, 2023Amr Hagag, Ahmed Gomaa, Dominik Kornek, Andreas Maier, Rainer Fietkau, Christoph Bert, Florian Putz, Yixing Huang
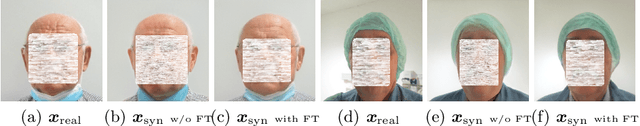

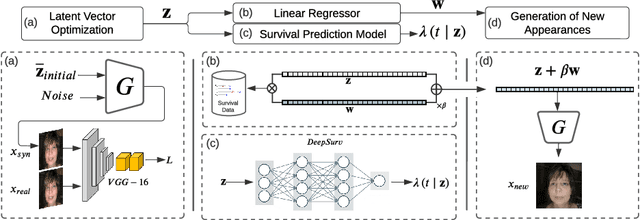

Survival prediction for cancer patients is critical for optimal treatment selection and patient management. Current patient survival prediction methods typically extract survival information from patients' clinical record data or biological and imaging data. In practice, experienced clinicians can have a preliminary assessment of patients' health status based on patients' observable physical appearances, which are mainly facial features. However, such assessment is highly subjective. In this work, the efficacy of objectively capturing and using prognostic information contained in conventional portrait photographs using deep learning for survival predication purposes is investigated for the first time. A pre-trained StyleGAN2 model is fine-tuned on a custom dataset of our cancer patients' photos to empower its generator with generative ability suitable for patients' photos. The StyleGAN2 is then used to embed the photographs to its highly expressive latent space. Utilizing the state-of-the-art survival analysis models and based on StyleGAN's latent space photo embeddings, this approach achieved a C-index of 0.677, which is notably higher than chance and evidencing the prognostic value embedded in simple 2D facial images. In addition, thanks to StyleGAN's interpretable latent space, our survival prediction model can be validated for relying on essential facial features, eliminating any biases from extraneous information like clothing or background. Moreover, a health attribute is obtained from regression coefficients, which has important potential value for patient care.
The Segment Anything foundation model achieves favorable brain tumor autosegmentation accuracy on MRI to support radiotherapy treatment planning
Apr 16, 2023Florian Putz, Johanna Grigo, Thomas Weissmann, Philipp Schubert, Daniel Hoefler, Ahmed Gomaa, Hassen Ben Tkhayat, Amr Hagag, Sebastian Lettmaier, Benjamin Frey, Udo S. Gaipl, Luitpold V. Distel, Sabine Semrau, Christoph Bert, Rainer Fietkau, Yixing Huang
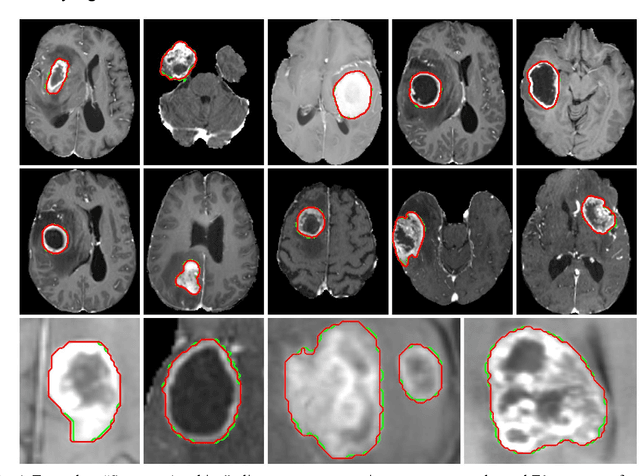
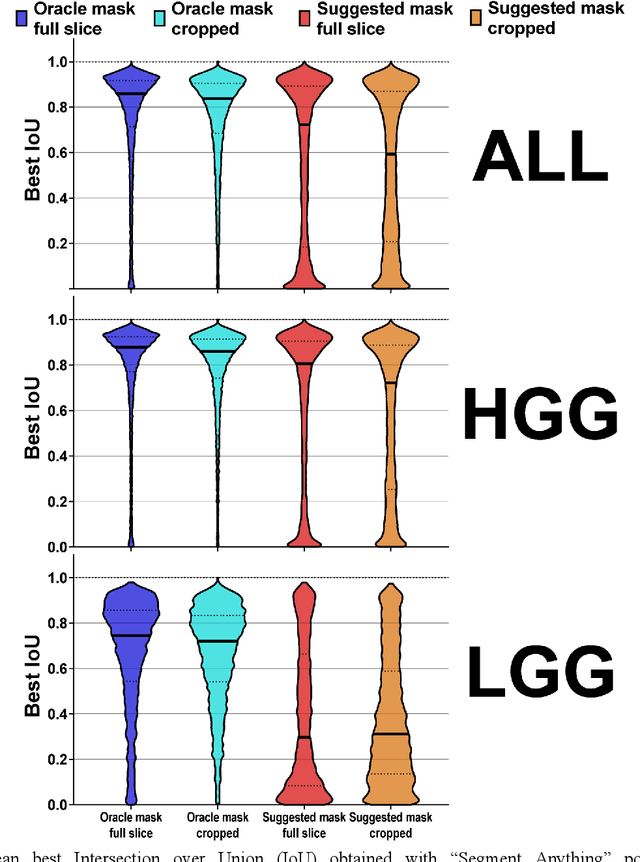
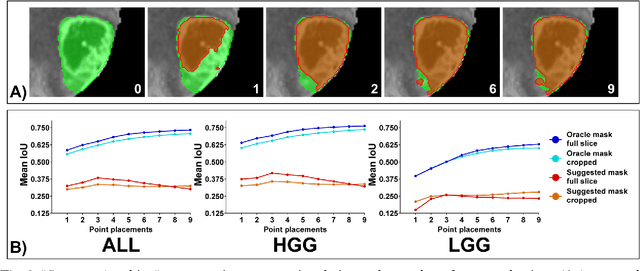
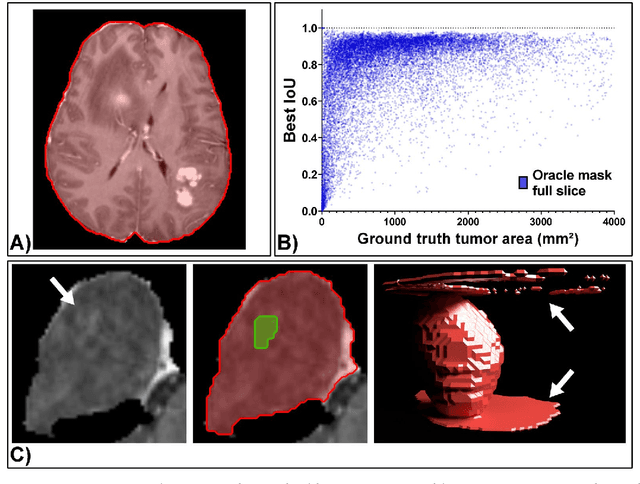
Background: Tumor segmentation in MRI is crucial in radiotherapy (RT) treatment planning for brain tumor patients. Segment anything (SA), a novel promptable foundation model for autosegmentation, has shown high accuracy for multiple segmentation tasks but was not evaluated on medical datasets yet. Methods: SA was evaluated in a point-to-mask task for glioma brain tumor autosegmentation on 16744 transversal slices from 369 MRI datasets (BraTS 2020). Up to 9 point prompts were placed per slice. Tumor core (enhancing tumor + necrotic core) was segmented on contrast-enhanced T1w sequences. Out of the 3 masks predicted by SA, accuracy was evaluated for the mask with the highest calculated IoU (oracle mask) and with highest model predicted IoU (suggested mask). In addition to assessing SA on whole MRI slices, SA was also evaluated on images cropped to the tumor (max. 3D extent + 2 cm). Results: Mean best IoU (mbIoU) using oracle mask on full MRI slices was 0.762 (IQR 0.713-0.917). Best 2D mask was achieved after a mean of 6.6 point prompts (IQR 5-9). Segmentation accuracy was significantly better for high- compared to low-grade glioma cases (mbIoU 0.789 vs. 0.668). Accuracy was worse using MRI slices cropped to the tumor (mbIoU 0.759) and was much worse using suggested mask (full slices 0.572). For all experiments, accuracy was low on peripheral slices with few tumor voxels (mbIoU, <300: 0.537 vs. >=300: 0.841). Stacking best oracle segmentations from full axial MRI slices, mean 3D DSC for tumor core was 0.872, which was improved to 0.919 by combining axial, sagittal and coronal masks. Conclusions: The Segment Anything foundation model, while trained on photos, can achieve high zero-shot accuracy for glioma brain tumor segmentation on MRI slices. The results suggest that Segment Anything can accelerate and facilitate RT treatment planning, when properly integrated in a clinical application.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge