Artificial Neural Networks for Sensor Data Classification on Small Embedded Systems
Dec 15, 2020Marcus Venzke, Daniel Klisch, Philipp Kubik, Asad Ali, Jesper Dell Missier, Volker Turau




In this paper we investigate the usage of machine learning for interpreting measured sensor values in sensor modules. In particular we analyze the potential of artificial neural networks (ANNs) on low-cost micro-controllers with a few kilobytes of memory to semantically enrich data captured by sensors. The focus is on classifying temporal data series with a high level of reliability. Design and implementation of ANNs are analyzed considering Feed Forward Neural Networks (FFNNs) and Recurrent Neural Networks (RNNs). We validate the developed ANNs in a case study of optical hand gesture recognition on an 8-bit micro-controller. The best reliability was found for an FFNN with two layers and 1493 parameters requiring an execution time of 36 ms. We propose a workflow to develop ANNs for embedded devices.
HMIC: Hierarchical Medical Image Classification, A Deep Learning Approach
Jun 23, 2020Kamran Kowsari, Rasoul Sali, Lubaina Ehsan, William Adorno, Asad Ali, Sean Moore, Beatrice Amadi, Paul Kelly, Sana Syed, Donald Brown
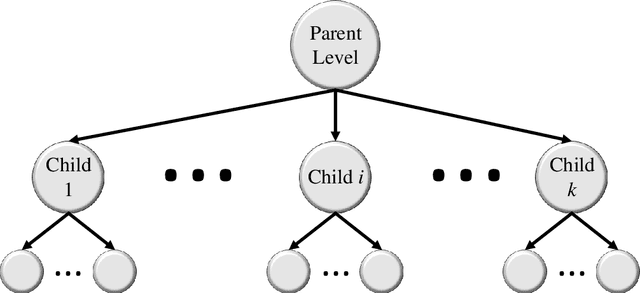
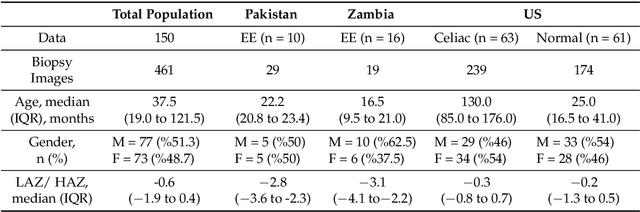
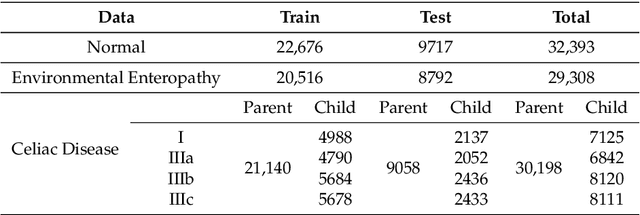
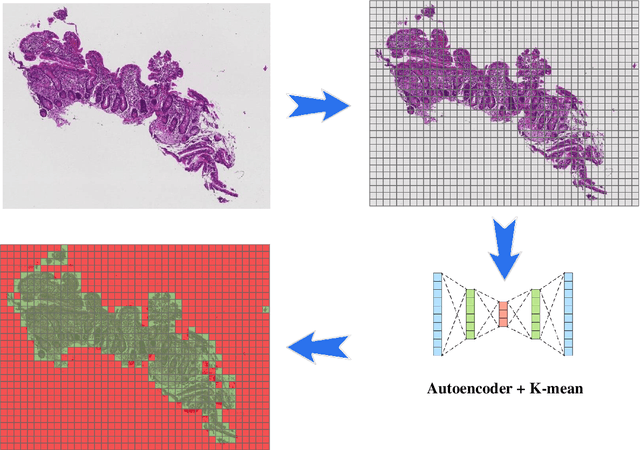
Image classification is central to the big data revolution in medicine. Improved information processing methods for diagnosis and classification of digital medical images have shown to be successful via deep learning approaches. As this field is explored, there are limitations to the performance of traditional supervised classifiers. This paper outlines an approach that is different from the current medical image classification tasks that view the issue as multi-class classification. We performed a hierarchical classification using our Hierarchical Medical Image classification (HMIC) approach. HMIC uses stacks of deep learning models to give particular comprehension at each level of the clinical picture hierarchy. For testing our performance, we use biopsy of the small bowel images that contain three categories in the parent level (Celiac Disease, Environmental Enteropathy, and histologically normal controls). For the child level, Celiac Disease Severity is classified into 4 classes (I, IIIa, IIIb, and IIIC).
Deep Learning for Visual Recognition of Environmental Enteropathy and Celiac Disease
Aug 08, 2019Aman Shrivastava, Karan Kant, Saurav Sengupta, Sung-Jun Kang, Marium Khan, Asad Ali, Sean R. Moore, Beatrice C. Amadi, Paul Kelly, Donald E. Brown, Sana Syed
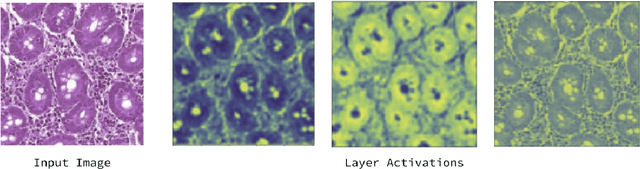
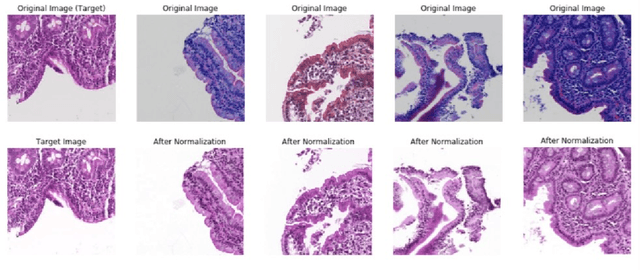
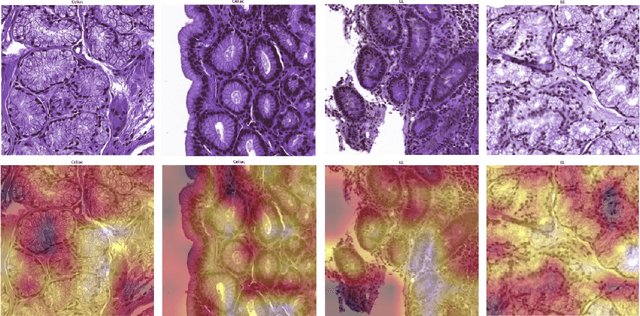
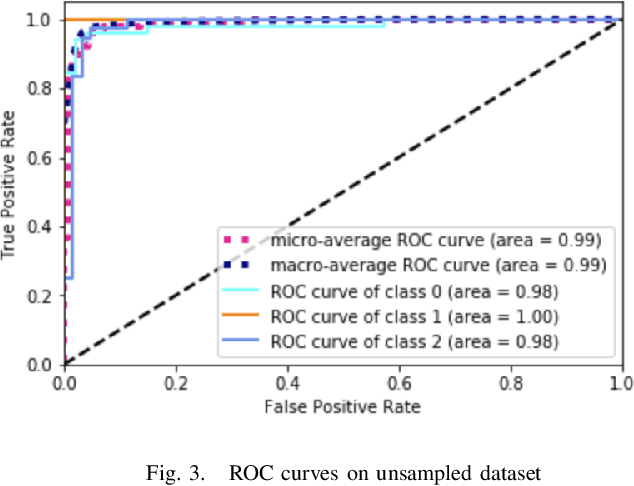

Physicians use biopsies to distinguish between different but histologically similar enteropathies. The range of syndromes and pathologies that could cause different gastrointestinal conditions makes this a difficult problem. Recently, deep learning has been used successfully in helping diagnose cancerous tissues in histopathological images. These successes motivated the research presented in this paper, which describes a deep learning approach that distinguishes between Celiac Disease (CD) and Environmental Enteropathy (EE) and normal tissue from digitized duodenal biopsies. Experimental results show accuracies of over 90% for this approach. We also look into interpreting the neural network model using Gradient-weighted Class Activation Mappings and filter activations on input images to understand the visual explanations for the decisions made by the model.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge