Tensor Decompositions for Identifying Directed Graph Topologies and Tracking Dynamic Networks
Oct 26, 2016Yanning Shen, Brian Baingana, Georgios B. Giannakis

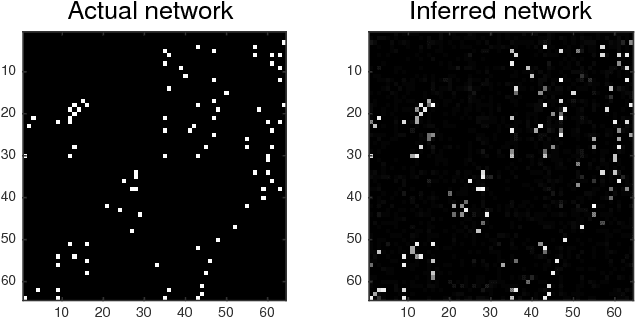
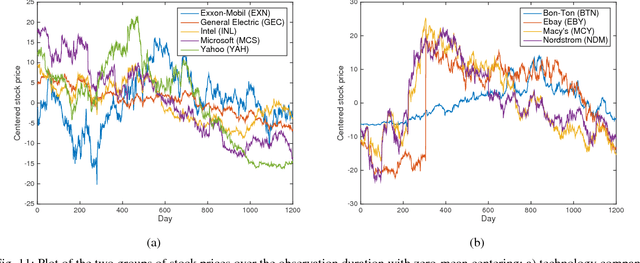

Directed networks are pervasive both in nature and engineered systems, often underlying the complex behavior observed in biological systems, microblogs and social interactions over the web, as well as global financial markets. Since their structures are often unobservable, in order to facilitate network analytics, one generally resorts to approaches capitalizing on measurable nodal processes to infer the unknown topology. Structural equation models (SEMs) are capable of incorporating exogenous inputs to resolve inherent directional ambiguities. However, conventional SEMs assume full knowledge of exogenous inputs, which may not be readily available in some practical settings. The present paper advocates a novel SEM-based topology inference approach that entails factorization of a three-way tensor, constructed from the observed nodal data, using the well-known parallel factor (PARAFAC) decomposition. It turns out that second-order piecewise stationary statistics of exogenous variables suffice to identify the hidden topology. Capitalizing on the uniqueness properties inherent to high-order tensor factorizations, it is shown that topology identification is possible under reasonably mild conditions. In addition, to facilitate real-time operation and inference of time-varying networks, an adaptive (PARAFAC) tensor decomposition scheme which tracks the topology-revealing tensor factors is developed. Extensive tests on simulated and real stock quote data demonstrate the merits of the novel tensor-based approach.
Nonlinear Structural Vector Autoregressive Models for Inferring Effective Brain Network Connectivity
Oct 20, 2016Yanning Shen, Brian Baingana, Georgios B. Giannakis
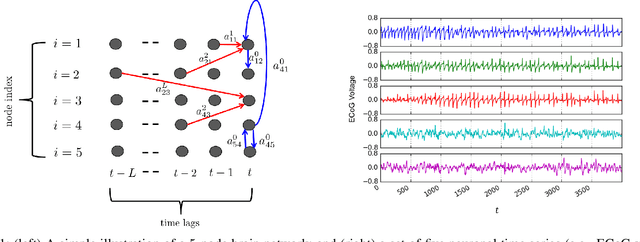



Structural equation models (SEMs) and vector autoregressive models (VARMs) are two broad families of approaches that have been shown useful in effective brain connectivity studies. While VARMs postulate that a given region of interest in the brain is directionally connected to another one by virtue of time-lagged influences, SEMs assert that causal dependencies arise due to contemporaneous effects, and may even be adopted when nodal measurements are not necessarily multivariate time series. To unify these complementary perspectives, linear structural vector autoregressive models (SVARMs) that leverage both contemporaneous and time-lagged nodal data have recently been put forth. Albeit simple and tractable, linear SVARMs are quite limited since they are incapable of modeling nonlinear dependencies between neuronal time series. To this end, the overarching goal of the present paper is to considerably broaden the span of linear SVARMs by capturing nonlinearities through kernels, which have recently emerged as a powerful nonlinear modeling framework in canonical machine learning tasks, e.g., regression, classification, and dimensionality reduction. The merits of kernel-based methods are extended here to the task of learning the effective brain connectivity, and an efficient regularized estimator is put forth to leverage the edge sparsity inherent to real-world complex networks. Judicious kernel choice from a preselected dictionary of kernels is also addressed using a data-driven approach. Extensive numerical tests on ECoG data captured through a study on epileptic seizures demonstrate that it is possible to unveil previously unknown causal links between brain regions of interest.
Tracking Switched Dynamic Network Topologies from Information Cascades
Jun 28, 2016Brian Baingana, Georgios B. Giannakis
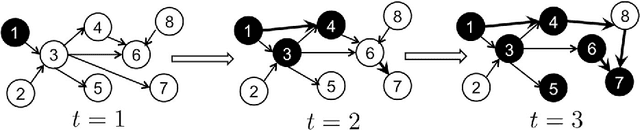
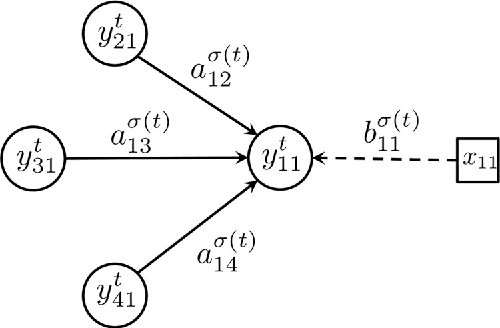
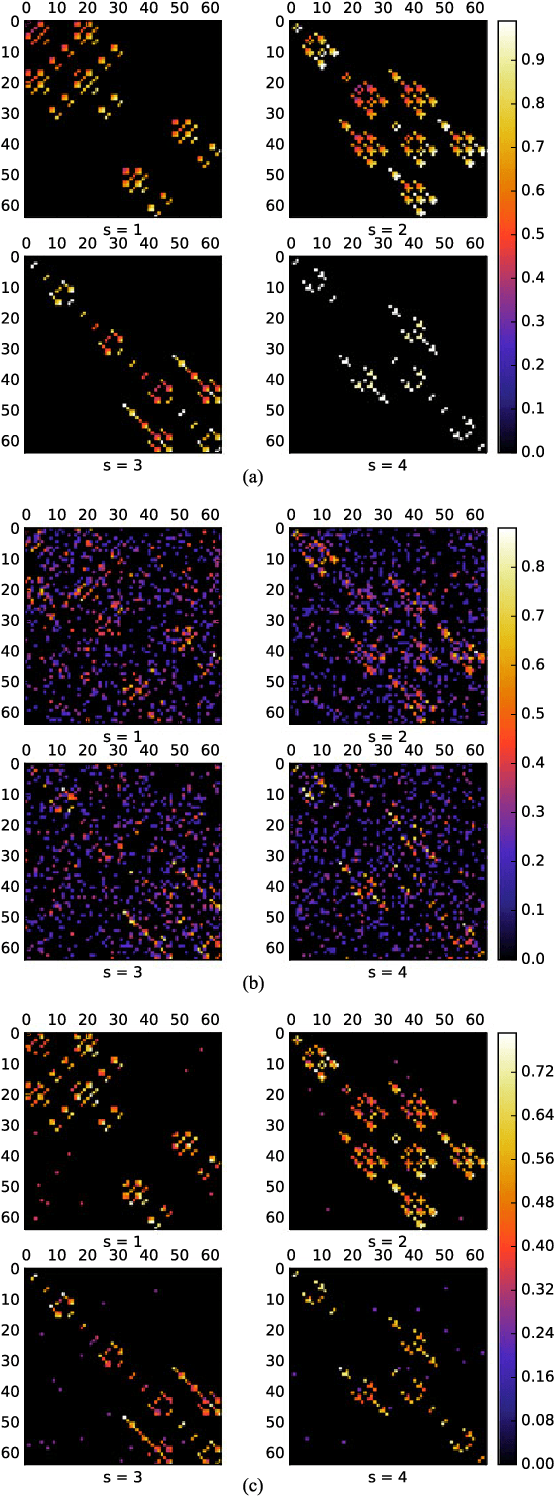
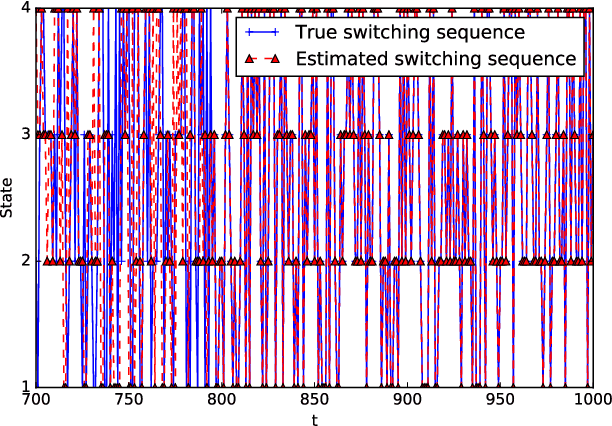
Contagions such as the spread of popular news stories, or infectious diseases, propagate in cascades over dynamic networks with unobservable topologies. However, "social signals" such as product purchase time, or blog entry timestamps are measurable, and implicitly depend on the underlying topology, making it possible to track it over time. Interestingly, network topologies often "jump" between discrete states that may account for sudden changes in the observed signals. The present paper advocates a switched dynamic structural equation model to capture the topology-dependent cascade evolution, as well as the discrete states driving the underlying topologies. Conditions under which the proposed switched model is identifiable are established. Leveraging the edge sparsity inherent to social networks, a recursive $\ell_1$-norm regularized least-squares estimator is put forth to jointly track the states and network topologies. An efficient first-order proximal-gradient algorithm is developed to solve the resulting optimization problem. Numerical experiments on both synthetic data and real cascades measured over the span of one year are conducted, and test results corroborate the efficacy of the advocated approach.
Kernel-Based Structural Equation Models for Topology Identification of Directed Networks
May 10, 2016Yanning Shen, Brian Baingana, Georgios B. Giannakis



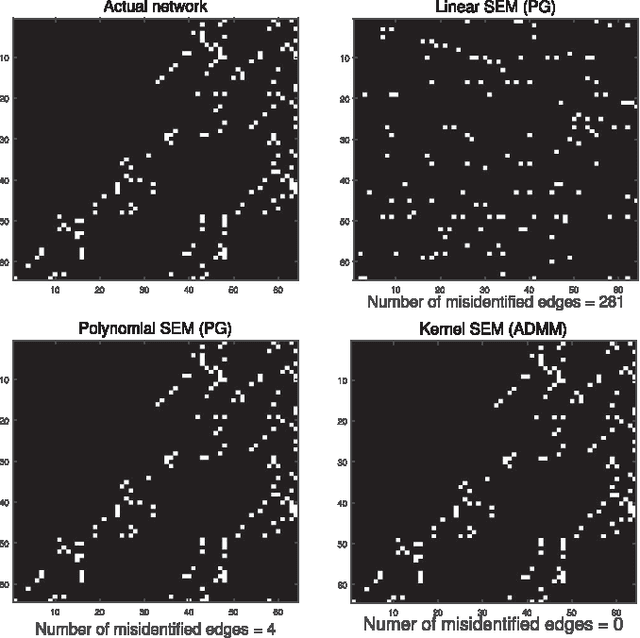
Structural equation models (SEMs) have been widely adopted for inference of causal interactions in complex networks. Recent examples include unveiling topologies of hidden causal networks over which processes such as spreading diseases, or rumors propagate. The appeal of SEMs in these settings stems from their simplicity and tractability, since they typically assume linear dependencies among observable variables. Acknowledging the limitations inherent to adopting linear models, the present paper advocates nonlinear SEMs, which account for (possible) nonlinear dependencies among network nodes. The advocated approach leverages kernels as a powerful encompassing framework for nonlinear modeling, and an efficient estimator with affordable tradeoffs is put forth. Interestingly, pursuit of the novel kernel-based approach yields a convex regularized estimator that promotes edge sparsity, and is amenable to proximal-splitting optimization methods. To this end, solvers with complementary merits are developed by leveraging the alternating direction method of multipliers, and proximal gradient iterations. Experiments conducted on simulated data demonstrate that the novel approach outperforms linear SEMs with respect to edge detection errors. Furthermore, tests on a real gene expression dataset unveil interesting new edges that were not revealed by linear SEMs, which could shed more light on regulatory behavior of human genes.
Joint community and anomaly tracking in dynamic networks
Jun 25, 2015Brian Baingana, Georgios B. Giannakis

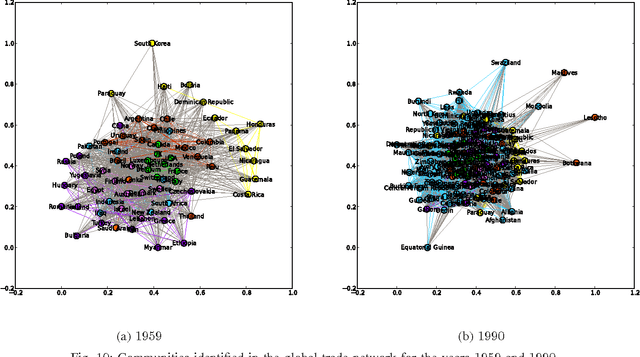
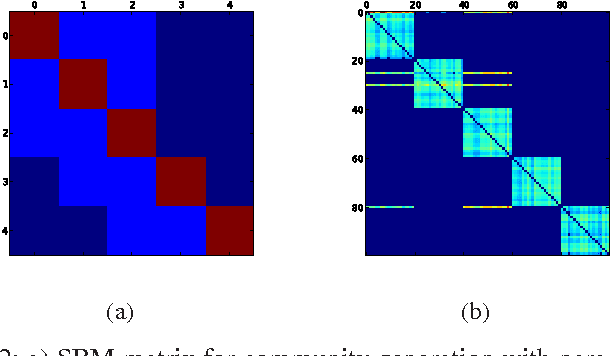

Most real-world networks exhibit community structure, a phenomenon characterized by existence of node clusters whose intra-edge connectivity is stronger than edge connectivities between nodes belonging to different clusters. In addition to facilitating a better understanding of network behavior, community detection finds many practical applications in diverse settings. Communities in online social networks are indicative of shared functional roles, or affiliation to a common socio-economic status, the knowledge of which is vital for targeted advertisement. In buyer-seller networks, community detection facilitates better product recommendations. Unfortunately, reliability of community assignments is hindered by anomalous user behavior often observed as unfair self-promotion, or "fake" highly-connected accounts created to promote fraud. The present paper advocates a novel approach for jointly tracking communities while detecting such anomalous nodes in time-varying networks. By postulating edge creation as the result of mutual community participation by node pairs, a dynamic factor model with anomalous memberships captured through a sparse outlier matrix is put forth. Efficient tracking algorithms suitable for both online and decentralized operation are developed. Experiments conducted on both synthetic and real network time series successfully unveil underlying communities and anomalous nodes.
Embedding Graphs under Centrality Constraints for Network Visualization
Jan 17, 2014Brian Baingana, Georgios B. Giannakis

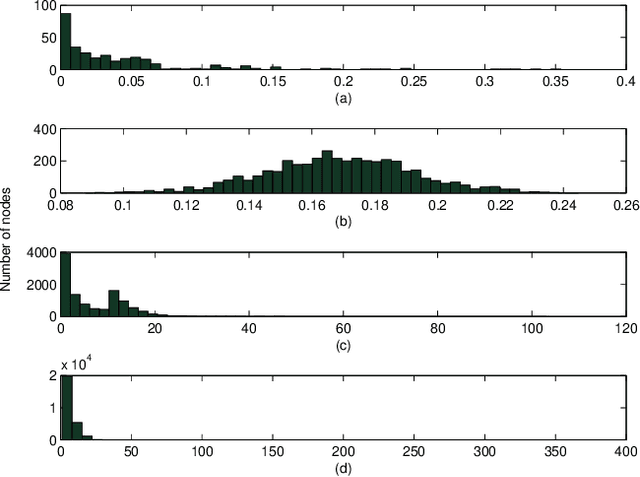
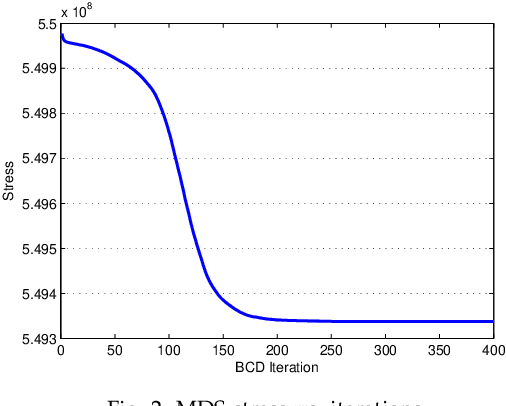

Visual rendering of graphs is a key task in the mapping of complex network data. Although most graph drawing algorithms emphasize aesthetic appeal, certain applications such as travel-time maps place more importance on visualization of structural network properties. The present paper advocates two graph embedding approaches with centrality considerations to comply with node hierarchy. The problem is formulated first as one of constrained multi-dimensional scaling (MDS), and it is solved via block coordinate descent iterations with successive approximations and guaranteed convergence to a KKT point. In addition, a regularization term enforcing graph smoothness is incorporated with the goal of reducing edge crossings. A second approach leverages the locally-linear embedding (LLE) algorithm which assumes that the graph encodes data sampled from a low-dimensional manifold. Closed-form solutions to the resulting centrality-constrained optimization problems are determined yielding meaningful embeddings. Experimental results demonstrate the efficacy of both approaches, especially for visualizing large networks on the order of thousands of nodes.
Centrality-constrained graph embedding
Feb 04, 2013Brian Baingana, Georgios B. Giannakis
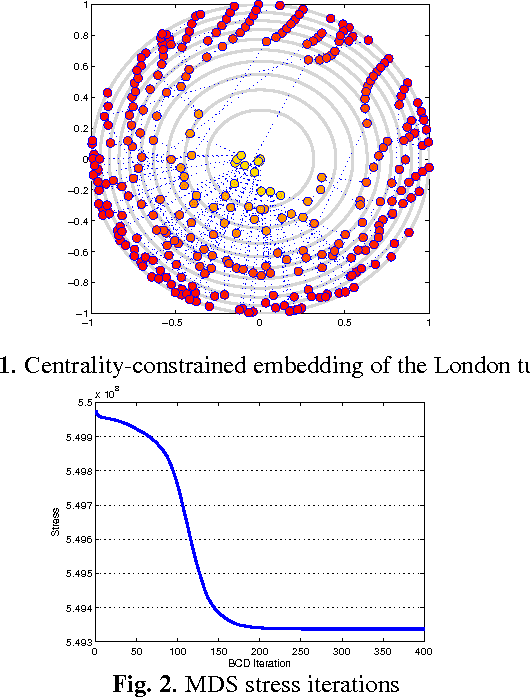
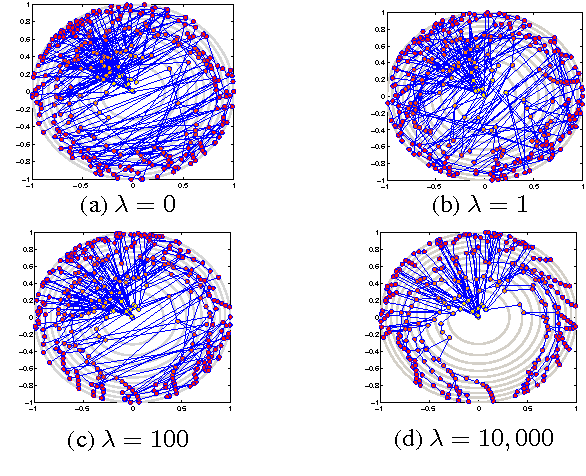
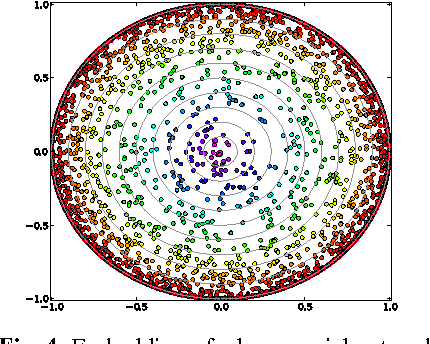
Visual rendering of graphs is a key task in the mapping of complex network data. Although most graph drawing algorithms emphasize aesthetic appeal, certain applications such as travel-time maps place more importance on visualization of structural network properties. The present paper advocates a graph embedding approach with centrality considerations to comply with node hierarchy. The problem is formulated as one of constrained multi-dimensional scaling (MDS), and it is solved via block coordinate descent iterations with successive approximations and guaranteed convergence to a KKT point. In addition, a regularization term enforcing graph smoothness is incorporated with the goal of reducing edge crossings. Experimental results demonstrate that the algorithm converges, and can be used to efficiently embed large graphs on the order of thousands of nodes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge