Computationally Efficient High-Dimensional Bayesian Optimization via Variable Selection
Sep 20, 2021Yihang Shen, Carl Kingsford




Bayesian Optimization (BO) is a method for globally optimizing black-box functions. While BO has been successfully applied to many scenarios, developing effective BO algorithms that scale to functions with high-dimensional domains is still a challenge. Optimizing such functions by vanilla BO is extremely time-consuming. Alternative strategies for high-dimensional BO that are based on the idea of embedding the high-dimensional space to the one with low dimension are sensitive to the choice of the embedding dimension, which needs to be pre-specified. We develop a new computationally efficient high-dimensional BO method that exploits variable selection. Our method is able to automatically learn axis-aligned sub-spaces, i.e. spaces containing selected variables, without the demand of any pre-specified hyperparameters. We theoretically analyze the computational complexity of our algorithm and derive the regret bound. We empirically show the efficacy of our method on several synthetic and real problems.
How much data is sufficient to learn high-performing algorithms?
Sep 09, 2019Maria-Florina Balcan, Dan DeBlasio, Travis Dick, Carl Kingsford, Tuomas Sandholm, Ellen Vitercik

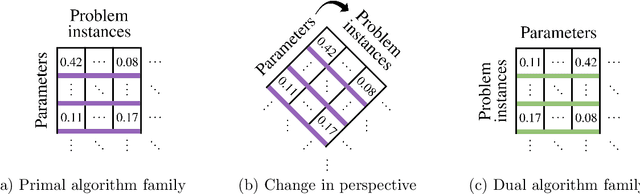
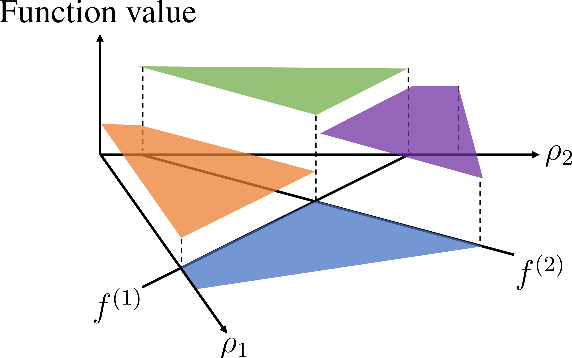
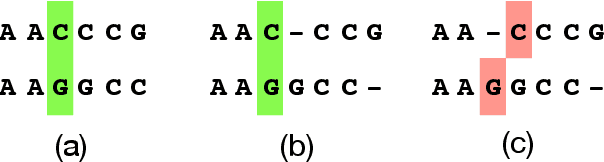
Algorithms for scientific analysis typically have tunable parameters that significantly influence computational efficiency and solution quality. If a parameter setting leads to strong algorithmic performance on average over a set of typical problem instances, that parameter setting---ideally---will perform well in the future. However, if the set of typical problem instances is small, average performance will not generalize to future performance. This raises the question: how large should this set be? We answer this question for any algorithm satisfying an easy-to-describe, ubiquitous property: its performance is a piecewise-structured function of its parameters. We are the first to provide a unified sample complexity framework for algorithm parameter configuration; prior research followed case-by-case analyses. We present applications from diverse domains, including biology, political science, and economics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge