Defending Against Transfer Attacks From Public Models
Oct 26, 2023Chawin Sitawarin, Jaewon Chang, David Huang, Wesson Altoyan, David Wagner
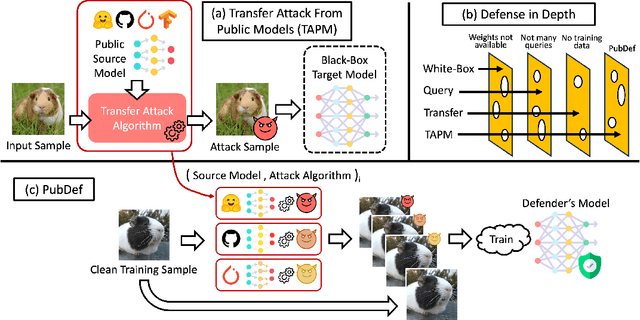
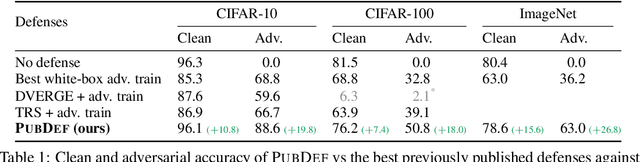
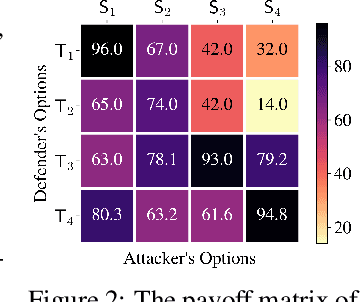
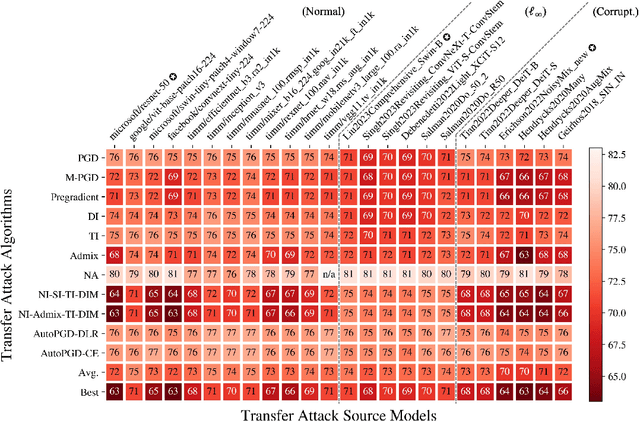
Adversarial attacks have been a looming and unaddressed threat in the industry. However, through a decade-long history of the robustness evaluation literature, we have learned that mounting a strong or optimal attack is challenging. It requires both machine learning and domain expertise. In other words, the white-box threat model, religiously assumed by a large majority of the past literature, is unrealistic. In this paper, we propose a new practical threat model where the adversary relies on transfer attacks through publicly available surrogate models. We argue that this setting will become the most prevalent for security-sensitive applications in the future. We evaluate the transfer attacks in this setting and propose a specialized defense method based on a game-theoretic perspective. The defenses are evaluated under 24 public models and 11 attack algorithms across three datasets (CIFAR-10, CIFAR-100, and ImageNet). Under this threat model, our defense, PubDef, outperforms the state-of-the-art white-box adversarial training by a large margin with almost no loss in the normal accuracy. For instance, on ImageNet, our defense achieves 62% accuracy under the strongest transfer attack vs only 36% of the best adversarially trained model. Its accuracy when not under attack is only 2% lower than that of an undefended model (78% vs 80%). We release our code at https://github.com/wagner-group/pubdef.
Interpretable Diabetic Retinopathy Diagnosis based on Biomarker Activation Map
Dec 13, 2022Pengxiao Zang, Tristan T. Hormel, Jie Wang, Yukun Guo, Steven T. Bailey, Christina J. Flaxel, David Huang, Thomas S. Hwang, Yali Jia




Deep learning classifiers provide the most accurate means of automatically diagnosing diabetic retinopathy (DR) based on optical coherence tomography (OCT) and its angiography (OCTA). The power of these models is attributable in part to the inclusion of hidden layers that provide the complexity required to achieve a desired task. However, hidden layers also render algorithm outputs difficult to interpret. Here we introduce a novel biomarker activation map (BAM) framework based on generative adversarial learning that allows clinicians to verify and understand classifiers decision-making. A data set including 456 macular scans were graded as non-referable or referable DR based on current clinical standards. A DR classifier that was used to evaluate our BAM was first trained based on this data set. The BAM generation framework was designed by combing two U-shaped generators to provide meaningful interpretability to this classifier. The main generator was trained to take referable scans as input and produce an output that would be classified by the classifier as non-referable. The BAM is then constructed as the difference image between the output and input of the main generator. To ensure that the BAM only highlights classifier-utilized biomarkers an assistant generator was trained to do the opposite, producing scans that would be classified as referable by the classifier from non-referable scans. The generated BAMs highlighted known pathologic features including nonperfusion area and retinal fluid. A fully interpretable classifier based on these highlights could help clinicians better utilize and verify automated DR diagnosis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge