A Self-explaining Neural Architecture for Generalizable Concept Learning
May 01, 2024Sanchit Sinha, Guangzhi Xiong, Aidong Zhang
With the wide proliferation of Deep Neural Networks in high-stake applications, there is a growing demand for explainability behind their decision-making process. Concept learning models attempt to learn high-level 'concepts' - abstract entities that align with human understanding, and thus provide interpretability to DNN architectures. However, in this paper, we demonstrate that present SOTA concept learning approaches suffer from two major problems - lack of concept fidelity wherein the models fail to learn consistent concepts among similar classes and limited concept interoperability wherein the models fail to generalize learned concepts to new domains for the same task. Keeping these in mind, we propose a novel self-explaining architecture for concept learning across domains which - i) incorporates a new concept saliency network for representative concept selection, ii) utilizes contrastive learning to capture representative domain invariant concepts, and iii) uses a novel prototype-based concept grounding regularization to improve concept alignment across domains. We demonstrate the efficacy of our proposed approach over current SOTA concept learning approaches on four widely used real-world datasets. Empirical results show that our method improves both concept fidelity measured through concept overlap and concept interoperability measured through domain adaptation performance.
Benchmarking Retrieval-Augmented Generation for Medicine
Feb 23, 2024Guangzhi Xiong, Qiao Jin, Zhiyong Lu, Aidong Zhang
While large language models (LLMs) have achieved state-of-the-art performance on a wide range of medical question answering (QA) tasks, they still face challenges with hallucinations and outdated knowledge. Retrieval-augmented generation (RAG) is a promising solution and has been widely adopted. However, a RAG system can involve multiple flexible components, and there is a lack of best practices regarding the optimal RAG setting for various medical purposes. To systematically evaluate such systems, we propose the Medical Information Retrieval-Augmented Generation Evaluation (MIRAGE), a first-of-its-kind benchmark including 7,663 questions from five medical QA datasets. Using MIRAGE, we conducted large-scale experiments with over 1.8 trillion prompt tokens on 41 combinations of different corpora, retrievers, and backbone LLMs through the MedRAG toolkit introduced in this work. Overall, MedRAG improves the accuracy of six different LLMs by up to 18% over chain-of-thought prompting, elevating the performance of GPT-3.5 and Mixtral to GPT-4-level. Our results show that the combination of various medical corpora and retrievers achieves the best performance. In addition, we discovered a log-linear scaling property and the "lost-in-the-middle" effects in medical RAG. We believe our comprehensive evaluations can serve as practical guidelines for implementing RAG systems for medicine.
Biomedical Question Answering: A Comprehensive Review
Feb 10, 2021Qiao Jin, Zheng Yuan, Guangzhi Xiong, Qianlan Yu, Chuanqi Tan, Mosha Chen, Songfang Huang, Xiaozhong Liu, Sheng Yu



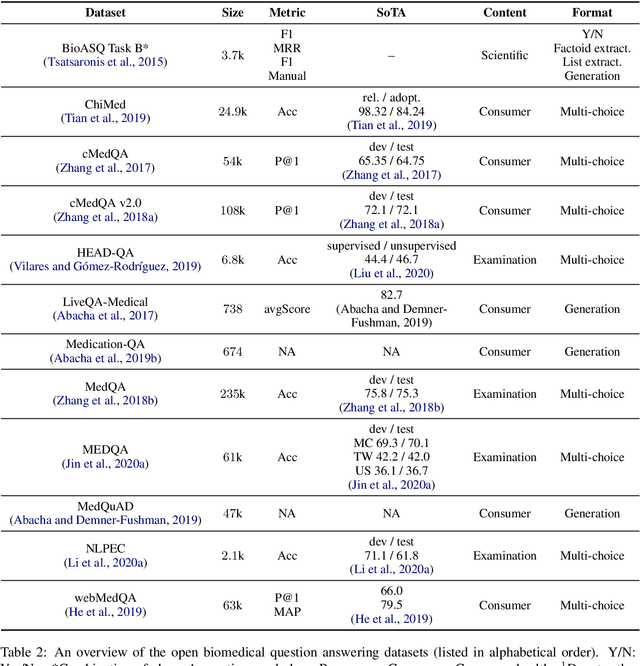
Question Answering (QA) is a benchmark Natural Language Processing (NLP) task where models predict the answer for a given question using related documents, images, knowledge bases and question-answer pairs. Automatic QA has been successfully applied in various domains like search engines and chatbots. However, for specific domains like biomedicine, QA systems are still rarely used in real-life settings. Biomedical QA (BQA), as an emerging QA task, enables innovative applications to effectively perceive, access and understand complex biomedical knowledge. In this work, we provide a critical review of recent efforts in BQA. We comprehensively investigate prior BQA approaches, which are classified into 6 major methodologies (open-domain, knowledge base, information retrieval, machine reading comprehension, question entailment and visual QA), 4 topics of contents (scientific, clinical, consumer health and examination) and 5 types of formats (yes/no, extraction, generation, multi-choice and retrieval). In the end, we highlight several key challenges of BQA and explore potential directions for future works.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge