Nonlinear Approximation and (Deep) ReLU Networks
May 05, 2019I. Daubechies, R. DeVore, S. Foucart, B. Hanin, G. Petrova
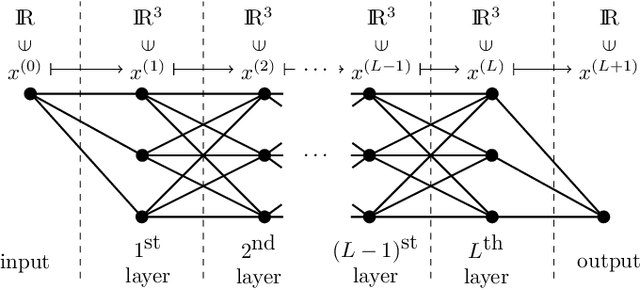
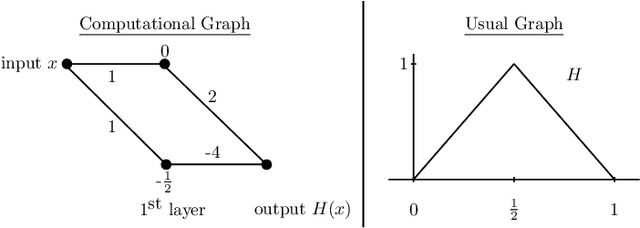
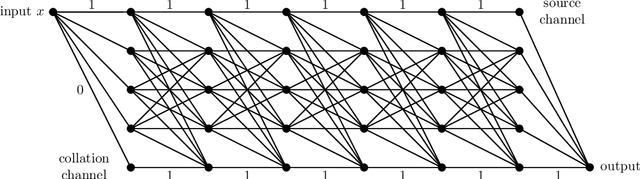
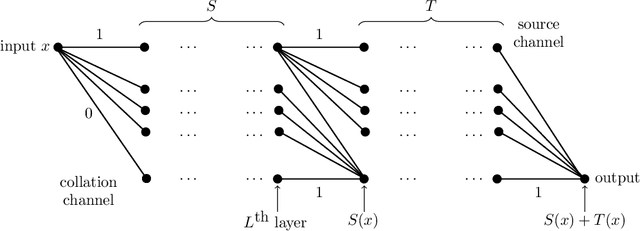
This article is concerned with the approximation and expressive powers of deep neural networks. This is an active research area currently producing many interesting papers. The results most commonly found in the literature prove that neural networks approximate functions with classical smoothness to the same accuracy as classical linear methods of approximation, e.g. approximation by polynomials or by piecewise polynomials on prescribed partitions. However, approximation by neural networks depending on n parameters is a form of nonlinear approximation and as such should be compared with other nonlinear methods such as variable knot splines or n-term approximation from dictionaries. The performance of neural networks in targeted applications such as machine learning indicate that they actually possess even greater approximation power than these traditional methods of nonlinear approximation. The main results of this article prove that this is indeed the case. This is done by exhibiting large classes of functions which can be efficiently captured by neural networks where classical nonlinear methods fall short of the task. The present article purposefully limits itself to studying the approximation of univariate functions by ReLU networks. Many generalizations to functions of several variables and other activation functions can be envisioned. However, even in this simplest of settings considered here, a theory that completely quantifies the approximation power of neural networks is still lacking.
Proceedings of the second "international Traveling Workshop on Interactions between Sparse models and Technology" (iTWIST'14)
Oct 09, 2014L. Jacques, C. De Vleeschouwer, Y. Boursier, P. Sudhakar, C. De Mol, A. Pizurica, S. Anthoine, P. Vandergheynst, P. Frossard, C. Bilen, S. Kitic, N. Bertin, R. Gribonval, N. Boumal, B. Mishra, P. -A. Absil, R. Sepulchre, S. Bundervoet, C. Schretter, A. Dooms, P. Schelkens, O. Chabiron, F. Malgouyres, J. -Y. Tourneret, N. Dobigeon, P. Chainais, C. Richard, B. Cornelis, I. Daubechies, D. Dunson, M. Dankova, P. Rajmic, K. Degraux, V. Cambareri, B. Geelen, G. Lafruit, G. Setti, J. -F. Determe, J. Louveaux, F. Horlin, A. Drémeau, P. Heas, C. Herzet, V. Duval, G. Peyré, A. Fawzi, M. Davies, N. Gillis, S. A. Vavasis, C. Soussen, L. Le Magoarou, J. Liang, J. Fadili, A. Liutkus, D. Martina, S. Gigan, L. Daudet, M. Maggioni, S. Minsker, N. Strawn, C. Mory, F. Ngole, J. -L. Starck, I. Loris, S. Vaiter, M. Golbabaee, D. Vukobratovic
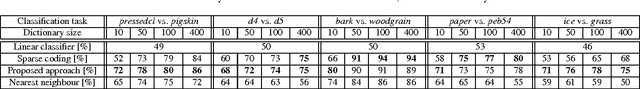

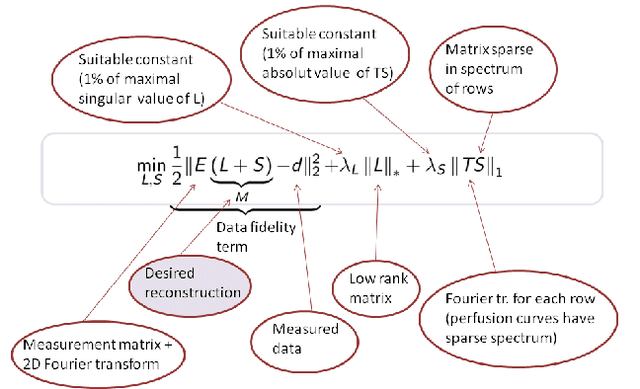
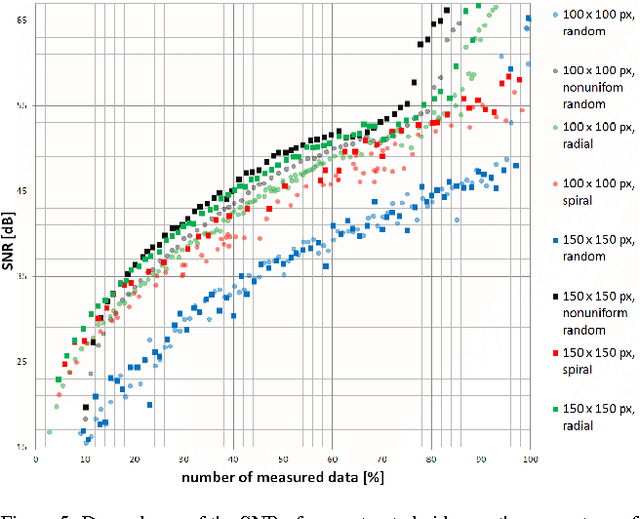
The implicit objective of the biennial "international - Traveling Workshop on Interactions between Sparse models and Technology" (iTWIST) is to foster collaboration between international scientific teams by disseminating ideas through both specific oral/poster presentations and free discussions. For its second edition, the iTWIST workshop took place in the medieval and picturesque town of Namur in Belgium, from Wednesday August 27th till Friday August 29th, 2014. The workshop was conveniently located in "The Arsenal" building within walking distance of both hotels and town center. iTWIST'14 has gathered about 70 international participants and has featured 9 invited talks, 10 oral presentations, and 14 posters on the following themes, all related to the theory, application and generalization of the "sparsity paradigm": Sparsity-driven data sensing and processing; Union of low dimensional subspaces; Beyond linear and convex inverse problem; Matrix/manifold/graph sensing/processing; Blind inverse problems and dictionary learning; Sparsity and computational neuroscience; Information theory, geometry and randomness; Complexity/accuracy tradeoffs in numerical methods; Sparsity? What's next?; Sparse machine learning and inference.
Algorithms to automatically quantify the geometric similarity of anatomical surfaces
Mar 15, 2012D. Boyer, Y. Lipman, E. St. Clair, J. Puente, T. Funkhouser, B. Patel, J. Jernvall, I. Daubechies
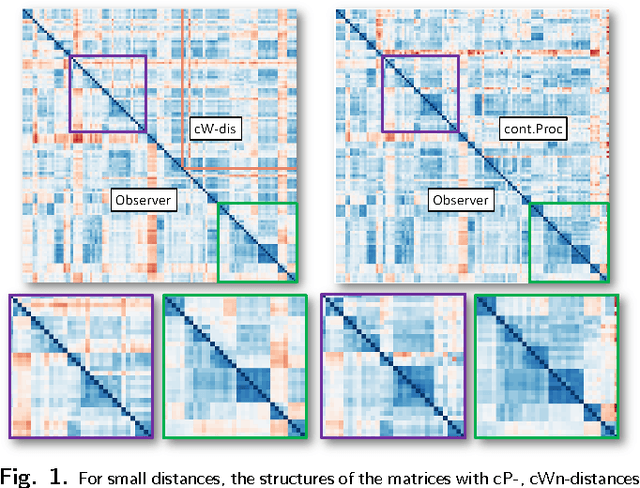
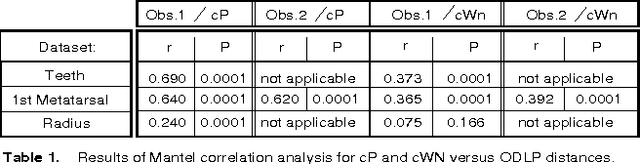


We describe new approaches for distances between pairs of 2-dimensional surfaces (embedded in 3-dimensional space) that use local structures and global information contained in inter-structure geometric relationships. We present algorithms to automatically determine these distances as well as geometric correspondences. This is motivated by the aspiration of students of natural science to understand the continuity of form that unites the diversity of life. At present, scientists using physical traits to study evolutionary relationships among living and extinct animals analyze data extracted from carefully defined anatomical correspondence points (landmarks). Identifying and recording these landmarks is time consuming and can be done accurately only by trained morphologists. This renders these studies inaccessible to non-morphologists, and causes phenomics to lag behind genomics in elucidating evolutionary patterns. Unlike other algorithms presented for morphological correspondences our approach does not require any preliminary marking of special features or landmarks by the user. It also differs from other seminal work in computational geometry in that our algorithms are polynomial in nature and thus faster, making pairwise comparisons feasible for significantly larger numbers of digitized surfaces. We illustrate our approach using three datasets representing teeth and different bones of primates and humans, and show that it leads to highly accurate results.
* Changes with respect to v1, v2: an Erratum was added, correcting the references for one of the three datasets. Note that the datasets and code for this paper can be obtained from the Data Conservancy (see Download column on v1, v2)
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge