Harnessing Intra-group Variations Via a Population-Level Context for Pathology Detection
Mar 04, 2024P. Bilha Githinji, Xi Yuan, Zhenglin Chen, Ijaz Gul, Dingqi Shang, Wen Liang, Jianming Deng, Dan Zeng, Dongmei yu, Chenggang Yan, Peiwu Qin
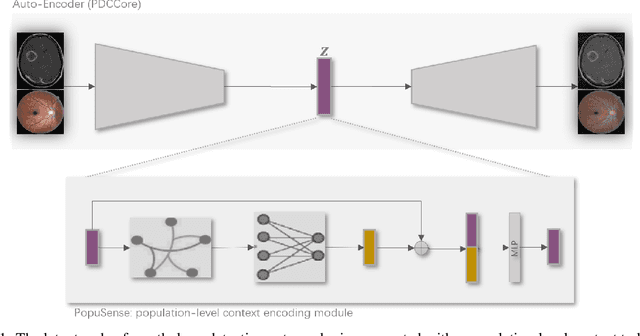
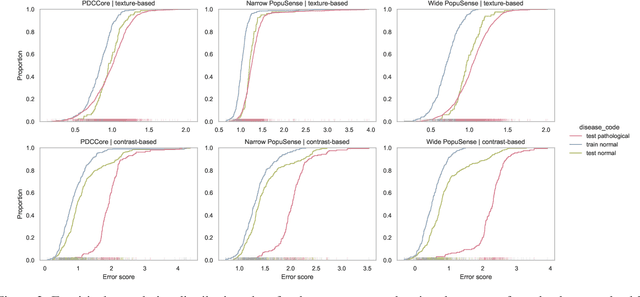
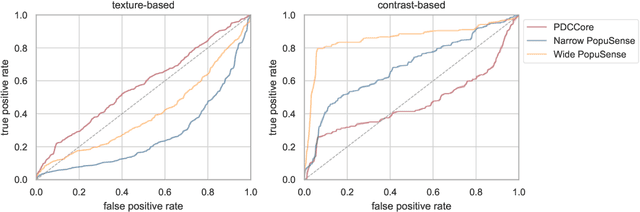
Realizing sufficient separability between the distributions of healthy and pathological samples is a critical obstacle for pathology detection convolutional models. Moreover, these models exhibit a bias for contrast-based images, with diminished performance on texture-based medical images. This study introduces the notion of a population-level context for pathology detection and employs a graph theoretic approach to model and incorporate it into the latent code of an autoencoder via a refinement module we term PopuSense. PopuSense seeks to capture additional intra-group variations inherent in biomedical data that a local or global context of the convolutional model might miss or smooth out. Experiments on contrast-based and texture-based images, with minimal adaptation, encounter the existing preference for intensity-based input. Nevertheless, PopuSense demonstrates improved separability in contrast-based images, presenting an additional avenue for refining representations learned by a model.
Neuro-Symbolic Learning: Principles and Applications in Ophthalmology
Jul 31, 2022Muhammad Hassan, Haifei Guan, Aikaterini Melliou, Yuqi Wang, Qianhui Sun, Sen Zeng, Wen Liang, Yiwei Zhang, Ziheng Zhang, Qiuyue Hu, Yang Liu, Shunkai Shi, Lin An, Shuyue Ma, Ijaz Gul, Muhammad Akmal Rahee, Zhou You, Canyang Zhang, Vijay Kumar Pandey, Yuxing Han, Yongbing Zhang, Ming Xu, Qiming Huang, Jiefu Tan, Qi Xing, Peiwu Qin, Dongmei Yu
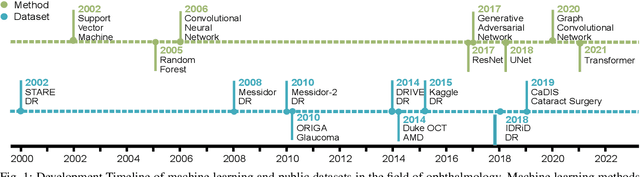
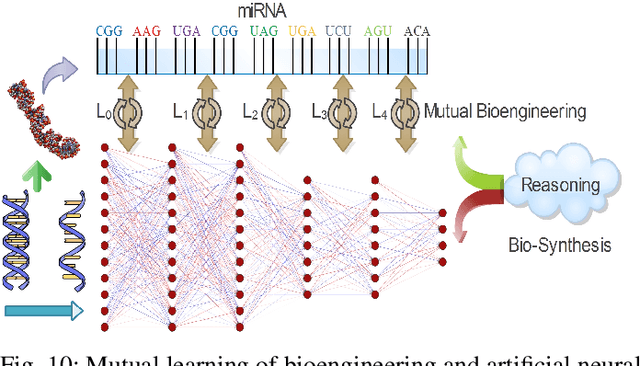
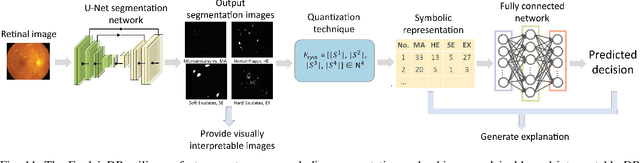
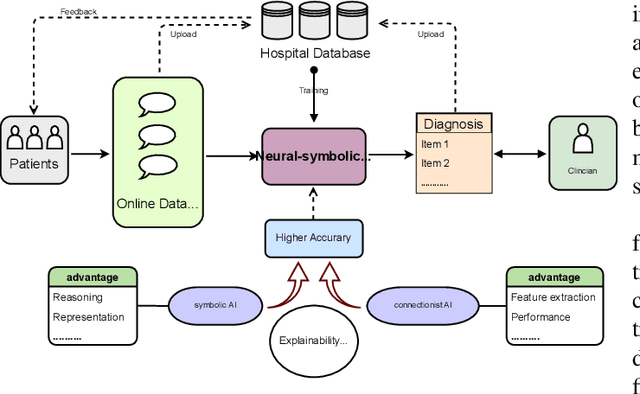
Neural networks have been rapidly expanding in recent years, with novel strategies and applications. However, challenges such as interpretability, explainability, robustness, safety, trust, and sensibility remain unsolved in neural network technologies, despite the fact that they will unavoidably be addressed for critical applications. Attempts have been made to overcome the challenges in neural network computing by representing and embedding domain knowledge in terms of symbolic representations. Thus, the neuro-symbolic learning (NeSyL) notion emerged, which incorporates aspects of symbolic representation and bringing common sense into neural networks (NeSyL). In domains where interpretability, reasoning, and explainability are crucial, such as video and image captioning, question-answering and reasoning, health informatics, and genomics, NeSyL has shown promising outcomes. This review presents a comprehensive survey on the state-of-the-art NeSyL approaches, their principles, advances in machine and deep learning algorithms, applications such as opthalmology, and most importantly, future perspectives of this emerging field.
RCMNet: A deep learning model assists CAR-T therapy for leukemia
May 06, 2022Ruitao Zhang, Xueying Han, Ijaz Gul, Shiyao Zhai, Ying Liu, Yongbing Zhang, Yuhan Dong, Lan Ma, Dongmei Yu, Jin Zhou, Peiwu Qin
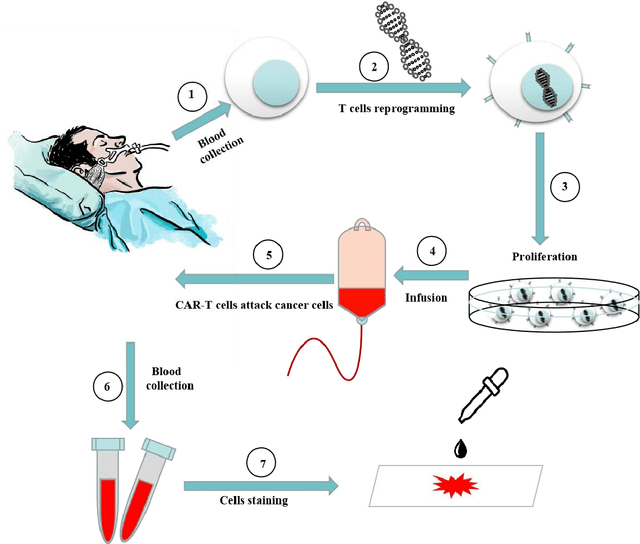
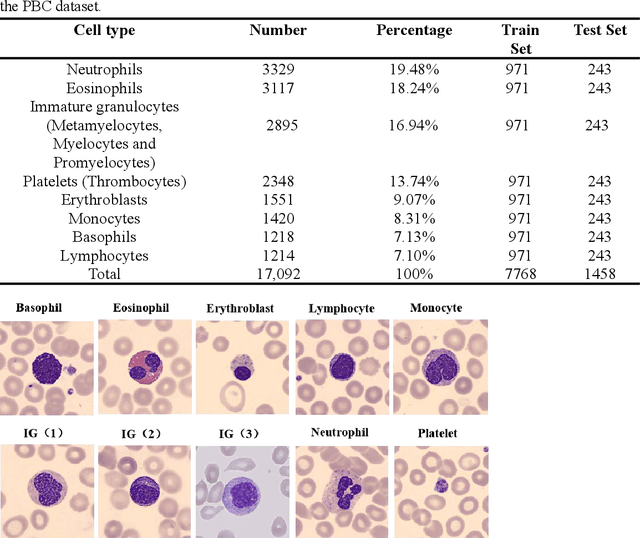
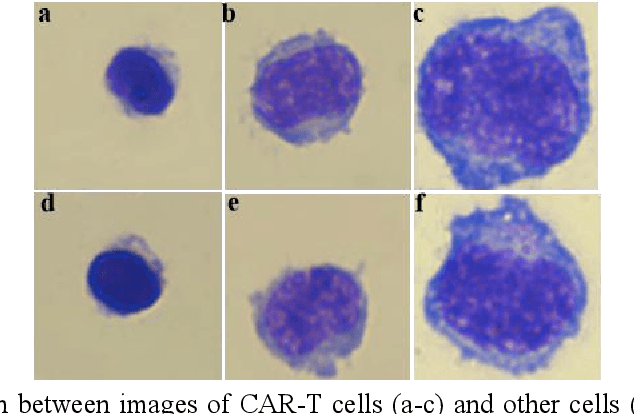

Acute leukemia is a type of blood cancer with a high mortality rate. Current therapeutic methods include bone marrow transplantation, supportive therapy, and chemotherapy. Although a satisfactory remission of the disease can be achieved, the risk of recurrence is still high. Therefore, novel treatments are demanding. Chimeric antigen receptor-T (CAR-T) therapy has emerged as a promising approach to treat and cure acute leukemia. To harness the therapeutic potential of CAR-T cell therapy for blood diseases, reliable cell morphological identification is crucial. Nevertheless, the identification of CAR-T cells is a big challenge posed by their phenotypic similarity with other blood cells. To address this substantial clinical challenge, herein we first construct a CAR-T dataset with 500 original microscopy images after staining. Following that, we create a novel integrated model called RCMNet (ResNet18 with CBAM and MHSA) that combines the convolutional neural network (CNN) and Transformer. The model shows 99.63% top-1 accuracy on the public dataset. Compared with previous reports, our model obtains satisfactory results for image classification. Although testing on the CAR-T cells dataset, a decent performance is observed, which is attributed to the limited size of the dataset. Transfer learning is adapted for RCMNet and a maximum of 83.36% accuracy has been achieved, which is higher than other SOTA models. The study evaluates the effectiveness of RCMNet on a big public dataset and translates it to a clinical dataset for diagnostic applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge