Rapid hyperspectral photothermal mid-infrared spectroscopic imaging from sparse data for gynecologic cancer tissue subtyping
Feb 28, 2024Reza Reihanisaransari, Chalapathi Charan Gajjela, Xinyu Wu, Ragib Ishrak, Sara Corvigno, Yanping Zhong, Jinsong Liu, Anil K. Sood, David Mayerich, Sebastian Berisha, Rohith Reddy
Ovarian cancer detection has traditionally relied on a multi-step process that includes biopsy, tissue staining, and morphological analysis by experienced pathologists. While widely practiced, this conventional approach suffers from several drawbacks: it is qualitative, time-intensive, and heavily dependent on the quality of staining. Mid-infrared (MIR) hyperspectral photothermal imaging is a label-free, biochemically quantitative technology that, when combined with machine learning algorithms, can eliminate the need for staining and provide quantitative results comparable to traditional histology. However, this technology is slow. This work presents a novel approach to MIR photothermal imaging that enhances its speed by an order of magnitude. Our method significantly accelerates data collection by capturing a combination of high-resolution and interleaved, lower-resolution infrared band images and applying computational techniques for data interpolation. We effectively minimize data collection requirements by leveraging sparse data acquisition and employing curvelet-based reconstruction algorithms. This method enables the reconstruction of high-quality, high-resolution images from undersampled datasets and achieving a 10X improvement in data acquisition time. We assessed the performance of our sparse imaging methodology using a variety of quantitative metrics, including mean squared error (MSE), structural similarity index (SSIM), and tissue subtype classification accuracies, employing both random forest and convolutional neural network (CNN) models, accompanied by ROC curves. Our statistically robust analysis, based on data from 100 ovarian cancer patient samples and over 65 million data points, demonstrates the method's capability to produce superior image quality and accurately distinguish between different gynecological tissue types with segmentation accuracy exceeding 95%.
Stochastic Dimension-reduced Second-order Methods for Policy Optimization
Jan 28, 2023Jinsong Liu, Chenghan Xie, Qi Deng, Dongdong Ge, Yinyu Ye

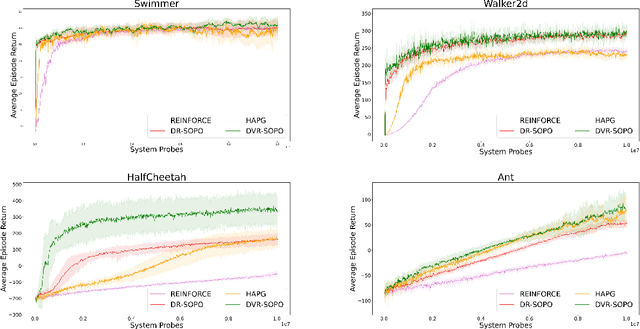
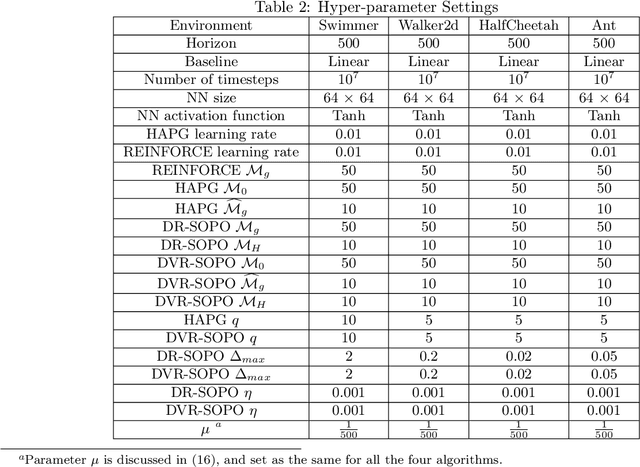
In this paper, we propose several new stochastic second-order algorithms for policy optimization that only require gradient and Hessian-vector product in each iteration, making them computationally efficient and comparable to policy gradient methods. Specifically, we propose a dimension-reduced second-order method (DR-SOPO) which repeatedly solves a projected two-dimensional trust region subproblem. We show that DR-SOPO obtains an $\mathcal{O}(\epsilon^{-3.5})$ complexity for reaching approximate first-order stationary condition and certain subspace second-order stationary condition. In addition, we present an enhanced algorithm (DVR-SOPO) which further improves the complexity to $\mathcal{O}(\epsilon^{-3})$ based on the variance reduction technique. Preliminary experiments show that our proposed algorithms perform favorably compared with stochastic and variance-reduced policy gradient methods.
Leveraging high-resolution spatial features in mid-infrared spectroscopic imaging to classify tissue subtypes in ovarian cancer
May 19, 2022Chalapathi Charan Gajjela, Matthew Brun, Rupali Mankar, Noah Kennedy, Sara Corvigno, Yanping Zhong, Jinsong Liu, Anil K. Sood, David Mayerich, Sebastian Berisha, Rohith Reddy




Mid-infrared spectroscopic imaging (MIRSI) is an emerging class of label-free techniques being leveraged for digital histopathology. Optical photothermal infrared (O-PTIR) is based on vibrational absorbance imaging using a pump-probe architecture capable of a 10x enhancement in spatial resolution relative to FTIR imaging. This allows truly sub-cellular spectroscopic investigation of tissue at biochemically important fingerprint wavelengths. Modern histopathologic identification of ovarian cancer involves tissue staining followed by morphological pattern recognition. This process is time-consuming, subjective, and requires extensive expertise. In this paper, we present the first label-free automated histological classification of ovarian tissue sub-types using MIRSI. We demonstrate that enhanced resolution of sub-cellular features, combined with spectroscopic information, enables reliable classification (0.98 AUC) of ovarian cell sub-types. Moreover, we present statistically robust validation from 74 patient samples with over 60 million data points. This demonstrates that sub-cellular resolution from five wavenumbers is sufficient to outperform state-of-the-art diffraction-limited techniques from up to 374 different wavenumbers. O-PTIR also performs measurements in back-reflection geometry, opening the door to future in vivo studies on glass slides.
Patch-based Fake Fingerprint Detection Using a Fully Convolutional Neural Network with a Small Number of Parameters and an Optimal Threshold
Mar 21, 2018Eunsoo Park, Xuenan Cui, Weonjin Kim, Jinsong Liu, Hakil Kim




Fingerprint authentication is widely used in biometrics due to its simple process, but it is vulnerable to fake fingerprints. This study proposes a patch-based fake fingerprint detection method using a fully convolutional neural network with a small number of parameters and an optimal threshold to solve the above-mentioned problem. Unlike the existing methods that classify a fingerprint as live or fake, the proposed method classifies fingerprints as fake, live, or background, so preprocessing methods such as segmentation are not needed. The proposed convolutional neural network (CNN) structure applies the Fire module of SqueezeNet, and the fewer parameters used require only 2.0 MB of memory. The network that has completed training is applied to the training data in a fully convolutional way, and the optimal threshold to distinguish fake fingerprints is determined, which is used in the final test. As a result of this study experiment, the proposed method showed an average classification error of 1.35%, demonstrating a fake fingerprint detection method using a high-performance CNN with a small number of parameters.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge