A Unified Model for Longitudinal Multi-Modal Multi-View Prediction with Missingness
Mar 22, 2024Boqi Chen, Junier Oliva, Marc Niethammer
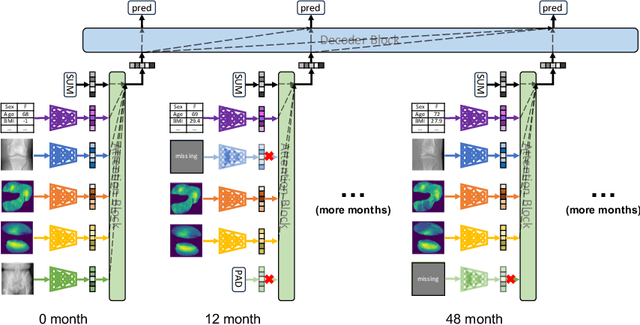
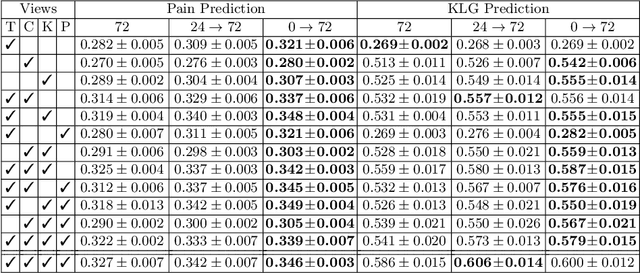
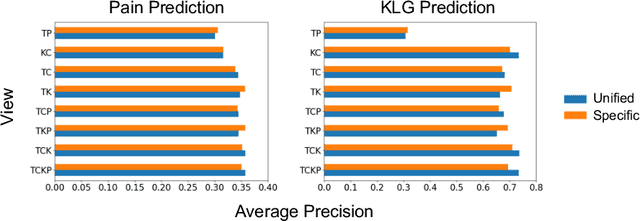
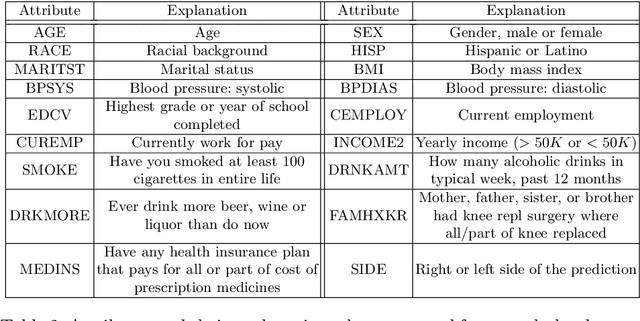
Medical records often consist of different modalities, such as images, text, and tabular information. Integrating all modalities offers a holistic view of a patient's condition, while analyzing them longitudinally provides a better understanding of disease progression. However, real-world longitudinal medical records present challenges: 1) patients may lack some or all of the data for a specific timepoint, and 2) certain modalities or views might be absent for all patients during a particular period. In this work, we introduce a unified model for longitudinal multi-modal multi-view prediction with missingness. Our method allows as many timepoints as desired for input, and aims to leverage all available data, regardless of their availability. We conduct extensive experiments on the knee osteoarthritis dataset from the Osteoarthritis Initiative for pain and Kellgren-Lawrence grade prediction at a future timepoint. We demonstrate the effectiveness of our method by comparing results from our unified model to specific models that use the same modality and view combinations during training and evaluation. We also show the benefit of having extended temporal data and provide post-hoc analysis for a deeper understanding of each modality/view's importance for different tasks.
Anomaly Detection via Gumbel Noise Score Matching
Apr 06, 2023Ahsan Mahmood, Junier Oliva, Martin Styner
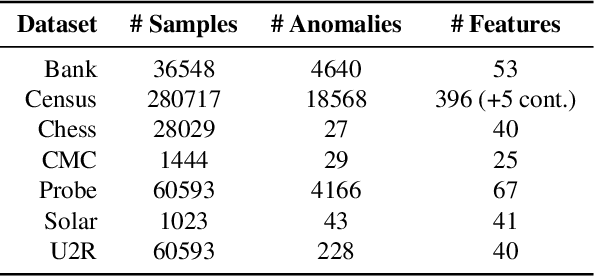
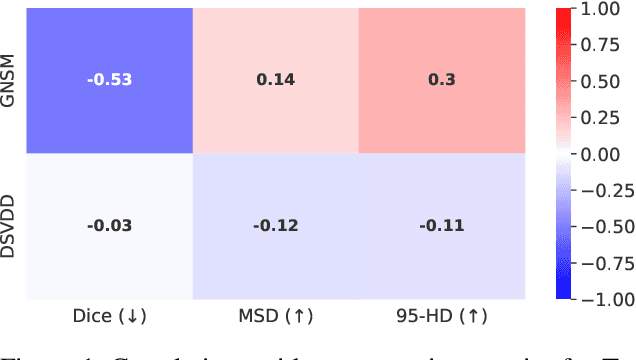
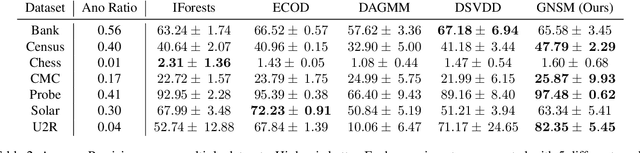

We propose Gumbel Noise Score Matching (GNSM), a novel unsupervised method to detect anomalies in categorical data. GNSM accomplishes this by estimating the scores, i.e. the gradients of log likelihoods w.r.t.~inputs, of continuously relaxed categorical distributions. We test our method on a suite of anomaly detection tabular datasets. GNSM achieves a consistently high performance across all experiments. We further demonstrate the flexibility of GNSM by applying it to image data where the model is tasked to detect poor segmentation predictions. Images ranked anomalous by GNSM show clear segmentation failures, with the outputs of GNSM strongly correlating with segmentation metrics computed on ground-truth. We outline the score matching training objective utilized by GNSM and provide an open-source implementation of our work.
Acquisition Conditioned Oracle for Nongreedy Active Feature Acquisition
Feb 27, 2023Michael Valancius, Max Lennon, Junier Oliva
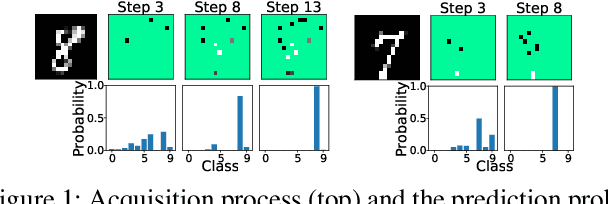
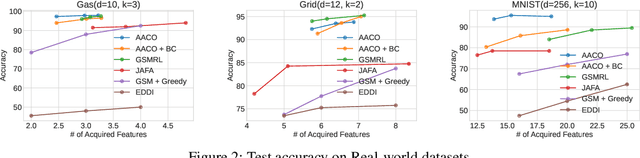
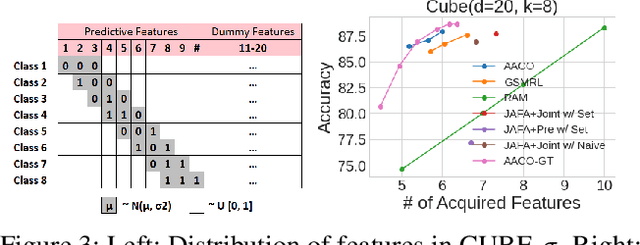
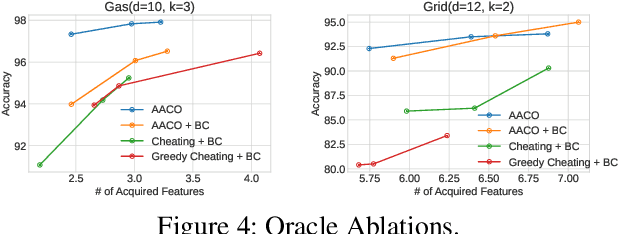
We develop novel methodology for active feature acquisition (AFA), the study of how to sequentially acquire a dynamic (on a per instance basis) subset of features that minimizes acquisition costs whilst still yielding accurate predictions. The AFA framework can be useful in a myriad of domains, including health care applications where the cost of acquiring additional features for a patient (in terms of time, money, risk, etc.) can be weighed against the expected improvement to diagnostic performance. Previous approaches for AFA have employed either: deep learning RL techniques, which have difficulty training policies in the AFA MDP due to sparse rewards and a complicated action space; deep learning surrogate generative models, which require modeling complicated multidimensional conditional distributions; or greedy policies, which fail to account for how joint feature acquisitions can be informative together for better predictions. In this work we show that we can bypass many of these challenges with a novel, nonparametric oracle based approach, which we coin the acquisition conditioned oracle (ACO). Extensive experiments show the superiority of the ACO to state-of-the-art AFA methods when acquiring features for both predictions and general decision-making.
Learning to Retrieve Videos by Asking Questions
May 13, 2022Avinash Madasu, Junier Oliva, Gedas Bertasius
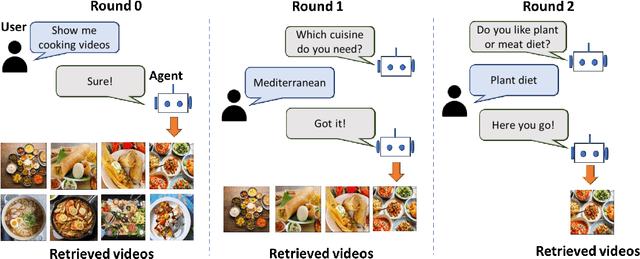

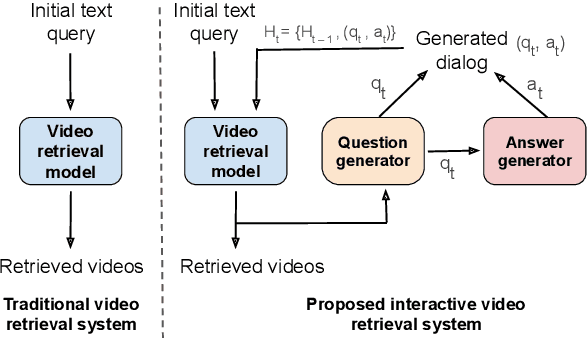

The majority of traditional text-to-video retrieval systems operate in static environments, i.e., there is no interaction between the user and the agent beyond the initial textual query provided by the user. This can be suboptimal if the initial query has ambiguities, which would lead to many falsely retrieved videos. To overcome this limitation, we propose a novel framework for Video Retrieval using Dialog (ViReD), which enables the user to interact with an AI agent via multiple rounds of dialog. The key contribution of our framework is a novel multimodal question generator that learns to ask questions that maximize the subsequent video retrieval performance. Our multimodal question generator uses (i) the video candidates retrieved during the last round of interaction with the user and (ii) the text-based dialog history documenting all previous interactions, to generate questions that incorporate both visual and linguistic cues relevant to video retrieval. Furthermore, to generate maximally informative questions, we propose an Information-Guided Supervision (IGS), which guides the question generator to ask questions that would boost subsequent video retrieval accuracy. We validate the effectiveness of our interactive ViReD framework on the AVSD dataset, showing that our interactive method performs significantly better than traditional non-interactive video retrieval systems. Furthermore, we also demonstrate that our proposed approach also generalizes to the real-world settings that involve interactions with real humans, thus, demonstrating the robustness and generality of our framework
Interpretable Single-Cell Set Classification with Kernel Mean Embeddings
Feb 10, 2022Siyuan Shan, Vishal Baskaran, Haidong Yi, Jolene Ranek, Natalie Stanley, Junier Oliva
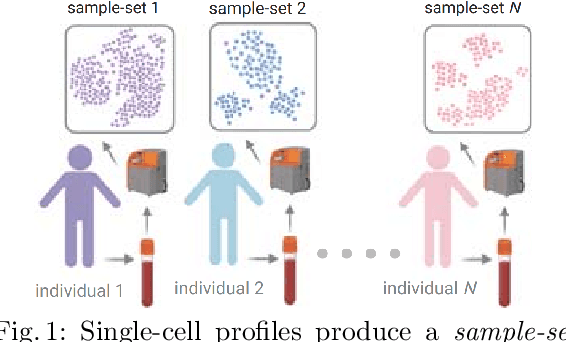
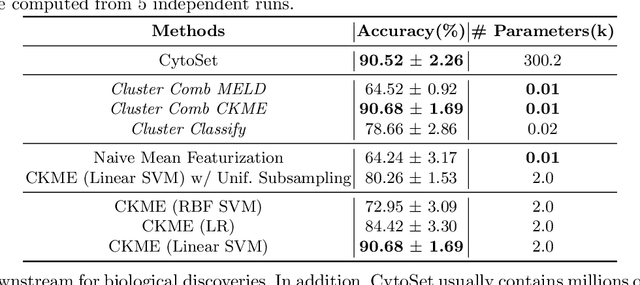
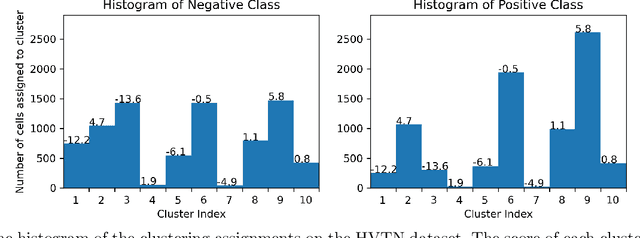
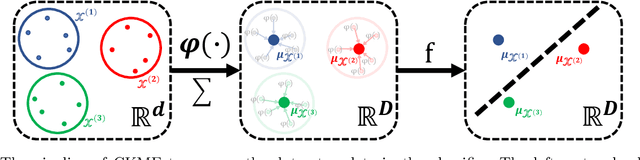
Modern single-cell flow and mass cytometry technologies measure the expression of several proteins of the individual cells within a blood or tissue sample. Each profiled biological sample is thus represented by a set of hundreds of thousands of multidimensional cell feature vectors, which incurs a high computational cost to predict each biological sample's associated phenotype with machine learning models. Such a large set cardinality also limits the interpretability of machine learning models due to the difficulty in tracking how each individual cell influences the ultimate prediction. Using Kernel Mean Embedding to encode the cellular landscape of each profiled biological sample, we can train a simple linear classifier and achieve state-of-the-art classification accuracy on 3 flow and mass cytometry datasets. Our model contains few parameters but still performs similarly to deep learning models with millions of parameters. In contrast with deep learning approaches, the linearity and sub-selection step of our model make it easy to interpret classification results. Clustering analysis further shows that our method admits rich biological interpretability for linking cellular heterogeneity to clinical phenotype.
Multiscale Score Matching for Out-of-Distribution Detection
Oct 27, 2020Ahsan Mahmood, Junier Oliva, Martin Styner
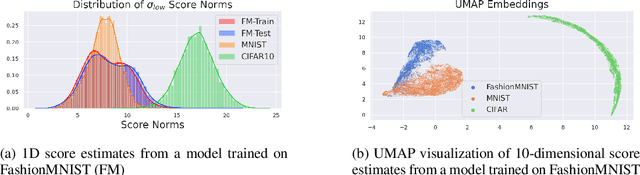
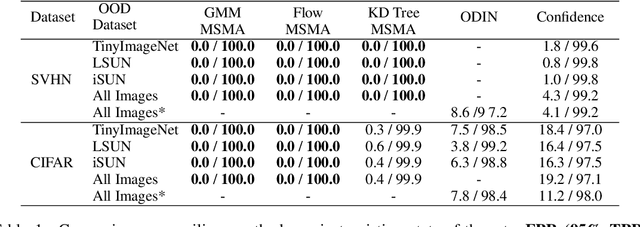
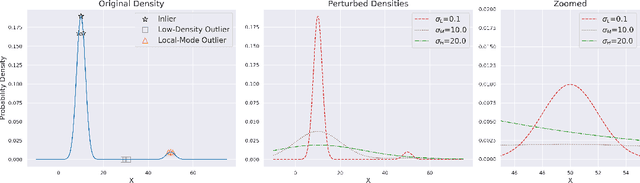
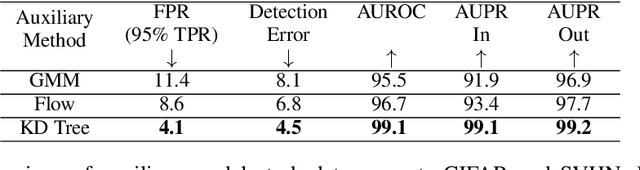
We present a new methodology for detecting out-of-distribution (OOD) images by utilizing norms of the score estimates at multiple noise scales. A score is defined to be the gradient of the log density with respect to the input data. Our methodology is completely unsupervised and follows a straight forward training scheme. First, we train a deep network to estimate scores for levels of noise. Once trained, we calculate the noisy score estimates for N in-distribution samples and take the L2-norms across the input dimensions (resulting in an NxL matrix). Then we train an auxiliary model (such as a Gaussian Mixture Model) to learn the in-distribution spatial regions in this L-dimensional space. This auxiliary model can now be used to identify points that reside outside the learned space. Despite its simplicity, our experiments show that this methodology significantly outperforms the state-of-the-art in detecting out-of-distribution images. For example, our method can effectively separate CIFAR-10 (inlier) and SVHN (OOD) images, a setting which has been previously shown to be difficult for deep likelihood models.
Deep Message Passing on Sets
Sep 21, 2019Yifeng Shi, Junier Oliva, Marc Niethammer
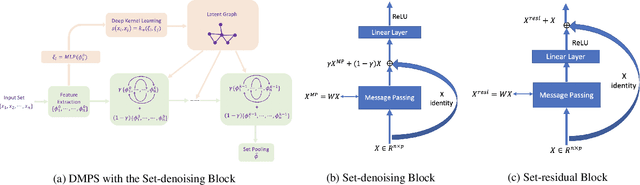
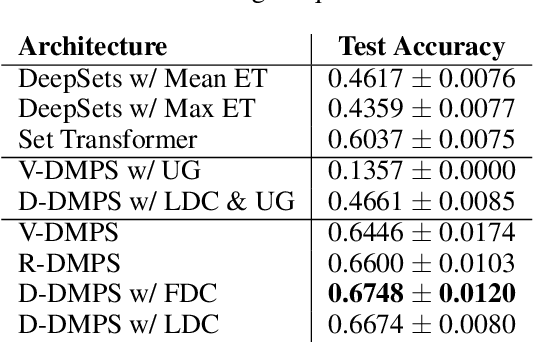
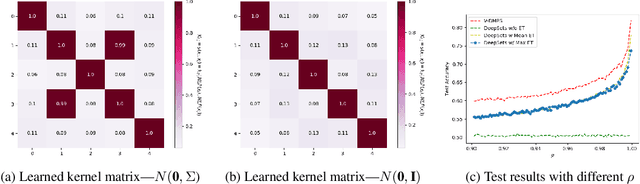
Modern methods for learning over graph input data have shown the fruitfulness of accounting for relationships among elements in a collection. However, most methods that learn over set input data use only rudimentary approaches to exploit intra-collection relationships. In this work we introduce Deep Message Passing on Sets (DMPS), a novel method that incorporates relational learning for sets. DMPS not only connects learning on graphs with learning on sets via deep kernel learning, but it also bridges message passing on sets and traditional diffusion dynamics commonly used in denoising models. Based on these connections, we develop two new blocks for relational learning on sets: the set-denoising block and the set-residual block. The former is motivated by the connection between message passing on general graphs and diffusion-based denoising models, whereas the latter is inspired by the well-known residual network. In addition to demonstrating the interpretability of our model by learning the true underlying relational structure experimentally, we also show the effectiveness of our approach on both synthetic and real-world datasets by achieving results that are competitive with or outperform the state-of-the-art.
Meta-Neighborhoods
Sep 18, 2019Siyuan Shan, Junier Oliva
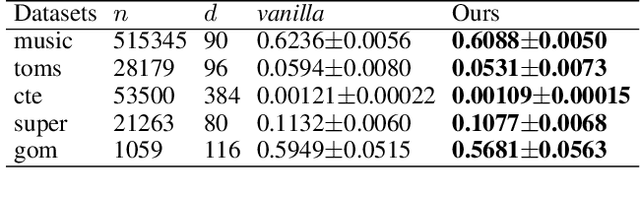
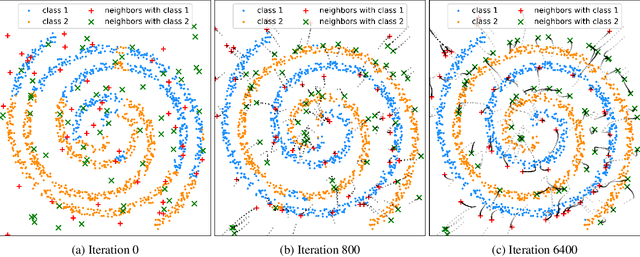
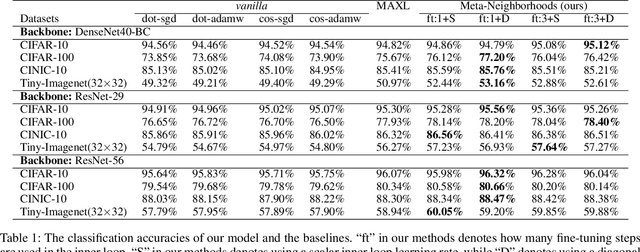
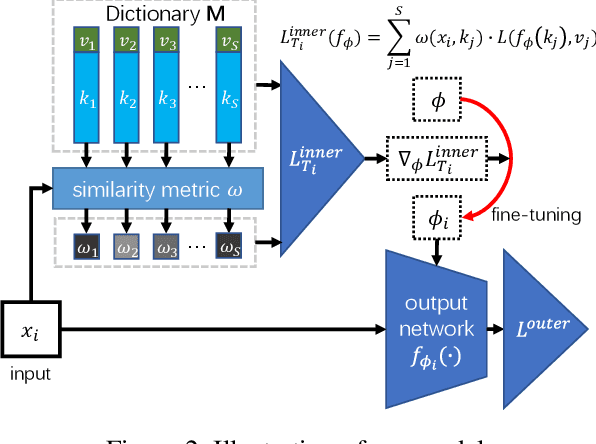

Traditional methods for training neural networks use training data just once, as it is discarded after training. Instead, in this work we also leverage the training data during testing to adjust the network and gain more expressivity. Our approach, named Meta-Neighborhoods, is developed under a multi-task learning framework and is a generalization of k-nearest neighbors methods. It can flexibly adapt network parameters w.r.t. different query data using their respective local neighborhood information. Local information is learned and stored in a dictionary of learnable neighbors rather than directly retrieved from the training set for greater flexibility and performance. The network parameters and the dictionary are optimized end-to-end via meta-learning. Extensive experiments demonstrate that Meta-Neighborhoods consistently improved classification and regression performance across various network architectures and datasets. We also observed superior improvements than other state-of-the-art meta-learning methods designed to improve supervised learning.
MolecularRNN: Generating realistic molecular graphs with optimized properties
May 31, 2019Mariya Popova, Mykhailo Shvets, Junier Oliva, Olexandr Isayev
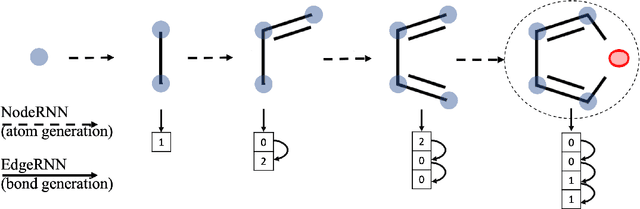
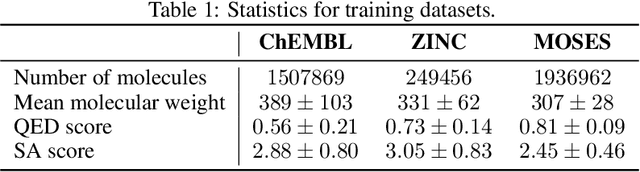
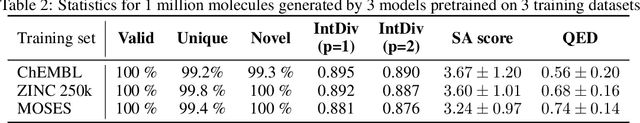
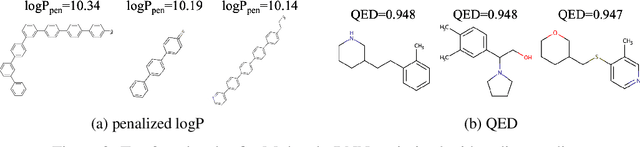
Designing new molecules with a set of predefined properties is a core problem in modern drug discovery and development. There is a growing need for de-novo design methods that would address this problem. We present MolecularRNN, the graph recurrent generative model for molecular structures. Our model generates diverse realistic molecular graphs after likelihood pretraining on a big database of molecules. We perform an analysis of our pretrained models on large-scale generated datasets of 1 million samples. Further, the model is tuned with policy gradient algorithm, provided a critic that estimates the reward for the property of interest. We show a significant distribution shift to the desired range for lipophilicity, drug-likeness, and melting point outperforming state-of-the-art works. With the use of rejection sampling based on valency constraints, our model yields 100% validity. Moreover, we show that invalid molecules provide a rich signal to the model through the use of structure penalty in our reinforcement learning pipeline.
Permutation Invariant Likelihoods and Equivariant Transformations
Feb 05, 2019Chris Bender, Juan Jose Garcia, Kevin O'Connor, Junier Oliva

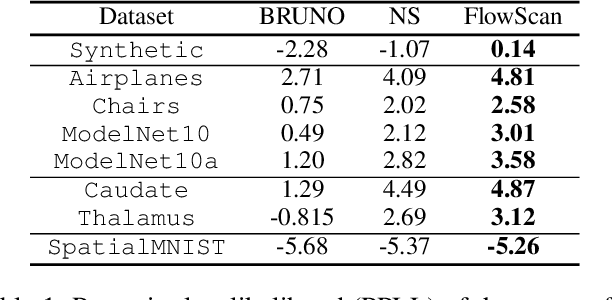
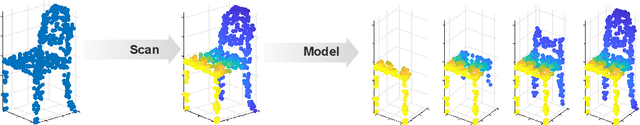
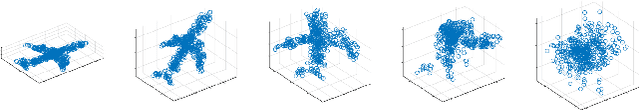
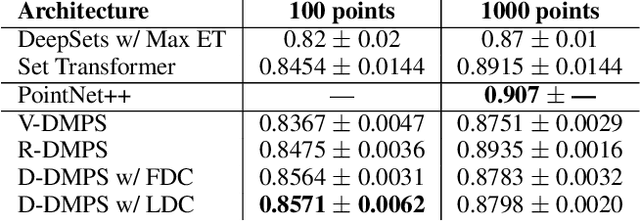
In this work, we fill a substantial void in machine learning and statistical methodology by developing extensive generative density estimation techniques for exchangeable non-iid data. We do so through the use of permutation invariant likelihoods and permutation equivariant transformations of variables. These methods exploit the intradependencies within sets in ways that are independent of ordering (for likelihoods) or order preserving (for transformations). The proposed techniques are able to directly model exchangeable data (such as sets) without the need to account for permutations or assume independence of elements. We consider applications to point clouds and provide several interesting experiments on both synthetic and real-world datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge