Nemotron-4 15B Technical Report
Feb 27, 2024Jupinder Parmar, Shrimai Prabhumoye, Joseph Jennings, Mostofa Patwary, Sandeep Subramanian, Dan Su, Chen Zhu, Deepak Narayanan, Aastha Jhunjhunwala, Ayush Dattagupta, Vibhu Jawa, Jiwei Liu, Ameya Mahabaleshwarkar, Osvald Nitski, Annika Brundyn, James Maki, Miguel Martinez, Jiaxuan You, John Kamalu, Patrick LeGresley, Denys Fridman, Jared Casper, Ashwath Aithal, Oleksii Kuchaiev, Mohammad Shoeybi, Jonathan Cohen, Bryan Catanzaro
We introduce Nemotron-4 15B, a 15-billion-parameter large multilingual language model trained on 8 trillion text tokens. Nemotron-4 15B demonstrates strong performance when assessed on English, multilingual, and coding tasks: it outperforms all existing similarly-sized open models on 4 out of 7 downstream evaluation areas and achieves competitive performance to the leading open models in the remaining ones. Specifically, Nemotron-4 15B exhibits the best multilingual capabilities of all similarly-sized models, even outperforming models over four times larger and those explicitly specialized for multilingual tasks.
The Importance of Background Information for Out of Distribution Generalization
Jun 17, 2022Jupinder Parmar, Khaled Saab, Brian Pogatchnik, Daniel Rubin, Christopher Ré
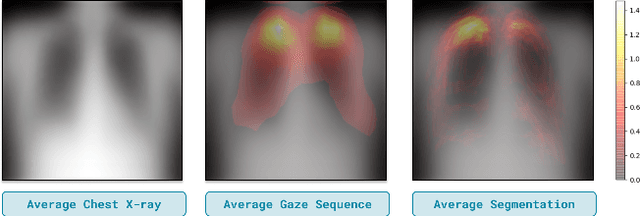
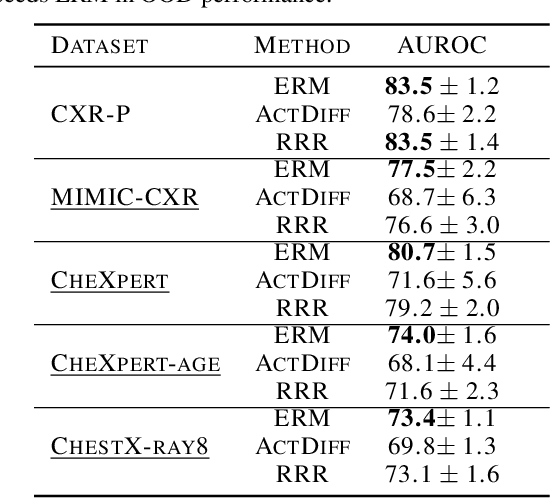


Domain generalization in medical image classification is an important problem for trustworthy machine learning to be deployed in healthcare. We find that existing approaches for domain generalization which utilize ground-truth abnormality segmentations to control feature attributions have poor out-of-distribution (OOD) performance relative to the standard baseline of empirical risk minimization (ERM). We investigate what regions of an image are important for medical image classification and show that parts of the background, that which is not contained in the abnormality segmentation, provides helpful signal. We then develop a new task-specific mask which covers all relevant regions. Utilizing this new segmentation mask significantly improves the performance of the existing methods on the OOD test sets. To obtain better generalization results than ERM, we find it necessary to scale up the training data size in addition to the usage of these task-specific masks.
Biomedical Information Extraction for Disease Gene Prioritization
Nov 12, 2020Jupinder Parmar, William Koehler, Martin Bringmann, Katharina Sophia Volz, Berk Kapicioglu



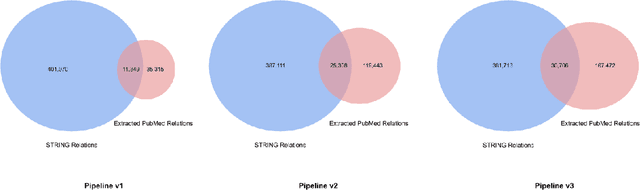
We introduce a biomedical information extraction (IE) pipeline that extracts biological relationships from text and demonstrate that its components, such as named entity recognition (NER) and relation extraction (RE), outperform state-of-the-art in BioNLP. We apply it to tens of millions of PubMed abstracts to extract protein-protein interactions (PPIs) and augment these extractions to a biomedical knowledge graph that already contains PPIs extracted from STRING, the leading structured PPI database. We show that, despite already containing PPIs from an established structured source, augmenting our own IE-based extractions to the graph allows us to predict novel disease-gene associations with a 20% relative increase in hit@30, an important step towards developing drug targets for uncured diseases.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge