REL: An Entity Linker Standing on the Shoulders of Giants
Jun 02, 2020Johannes M. van Hulst, Faegheh Hasibi, Koen Dercksen, Krisztian Balog, Arjen P. de Vries

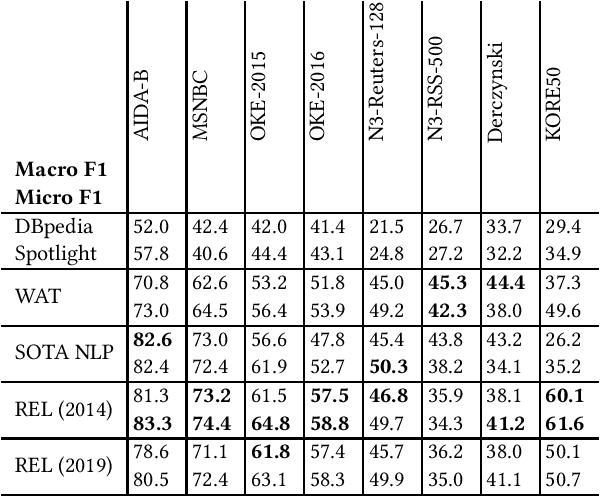
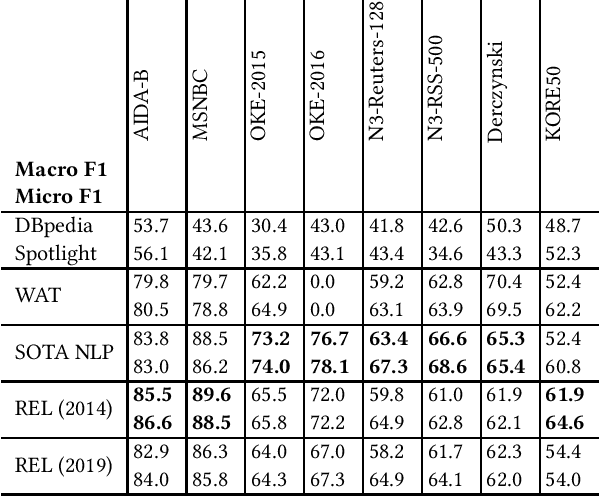
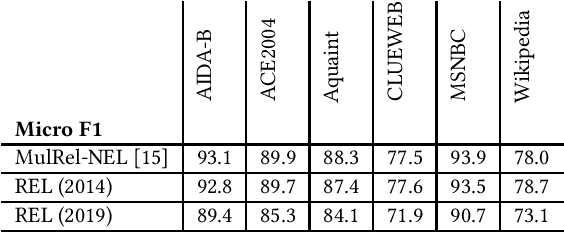
Entity linking is a standard component in modern retrieval system that is often performed by third-party toolkits. Despite the plethora of open source options, it is difficult to find a single system that has a modular architecture where certain components may be replaced, does not depend on external sources, can easily be updated to newer Wikipedia versions, and, most important of all, has state-of-the-art performance. The REL system presented in this paper aims to fill that gap. Building on state-of-the-art neural components from natural language processing research, it is provided as a Python package as well as a web API. We also report on an experimental comparison against both well-established systems and the current state-of-the-art on standard entity linking benchmarks.
Dealing with Label Scarcity in Computational Pathology: A Use Case in Prostate Cancer Classification
May 16, 2019Koen Dercksen, Wouter Bulten, Geert Litjens
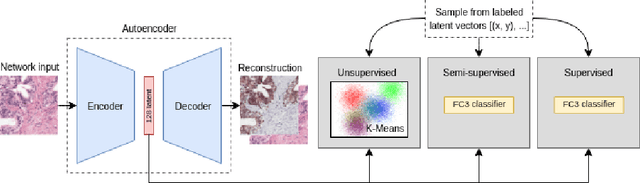
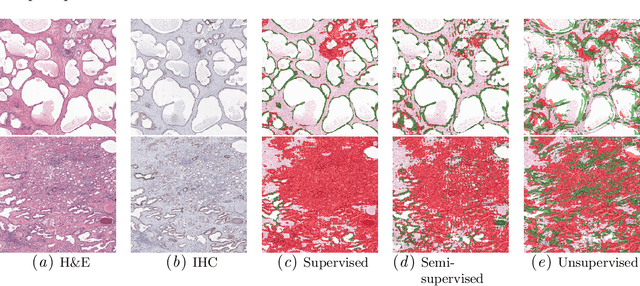
Large amounts of unlabelled data are commonplace for many applications in computational pathology, whereas labelled data is often expensive, both in time and cost, to acquire. We investigate the performance of unsupervised and supervised deep learning methods when few labelled data are available. Three methods are compared: clustering autoencoder latent vectors (unsupervised), a single layer classifier combined with a pre-trained autoencoder (semi-supervised), and a supervised CNN. We apply these methods on hematoxylin and eosin (H&E) stained prostatectomy images to classify tumour versus non-tumour tissue. Results show that semi-/unsupervised methods have an advantage over supervised learning when few labels are available. Additionally, we show that incorporating immunohistochemistry (IHC) stained data provides an increase in performance over only using H&E.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge