Sub2Full: split spectrum to boost OCT despeckling without clean data
Jan 18, 2024Lingyun Wang, Jose A Sahel, Shaohua Pi
Optical coherence tomography (OCT) suffers from speckle noise, causing the deterioration of image quality, especially in high-resolution modalities like visible light OCT (vis-OCT). The potential of conventional supervised deep learning denoising methods is limited by the difficulty of obtaining clean data. Here, we proposed an innovative self-supervised strategy called Sub2Full (S2F) for OCT despeckling without clean data. This approach works by acquiring two repeated B-scans, splitting the spectrum of the first repeat as a low-resolution input, and utilizing the full spectrum of the second repeat as the high-resolution target. The proposed method was validated on vis-OCT retinal images visualizing sublaminar structures in outer retina and demonstrated superior performance over conventional Noise2Noise and Noise2Void schemes. The code is available at https://github.com/PittOCT/Sub2Full-OCT-Denoising.
AG-CRC: Anatomy-Guided Colorectal Cancer Segmentation in CT with Imperfect Anatomical Knowledge
Oct 07, 2023Rongzhao Zhang, Zhian Bai, Ruoying Yu, Wenrao Pang, Lingyun Wang, Lifeng Zhu, Xiaofan Zhang, Huan Zhang, Weiguo Hu
When delineating lesions from medical images, a human expert can always keep in mind the anatomical structure behind the voxels. However, although high-quality (though not perfect) anatomical information can be retrieved from computed tomography (CT) scans with modern deep learning algorithms, it is still an open problem how these automatically generated organ masks can assist in addressing challenging lesion segmentation tasks, such as the segmentation of colorectal cancer (CRC). In this paper, we develop a novel Anatomy-Guided segmentation framework to exploit the auto-generated organ masks to aid CRC segmentation from CT, namely AG-CRC. First, we obtain multi-organ segmentation (MOS) masks with existing MOS models (e.g., TotalSegmentor) and further derive a more robust organ of interest (OOI) mask that may cover most of the colon-rectum and CRC voxels. Then, we propose an anatomy-guided training patch sampling strategy by optimizing a heuristic gain function that considers both the proximity of important regions (e.g., the tumor or organs of interest) and sample diversity. Third, we design a novel self-supervised learning scheme inspired by the topology of tubular organs like the colon to boost the model performance further. Finally, we employ a masked loss scheme to guide the model to focus solely on the essential learning region. We extensively evaluate the proposed method on two CRC segmentation datasets, where substantial performance improvement (5% to 9% in Dice) is achieved over current state-of-the-art medical image segmentation models, and the ablation studies further evidence the efficacy of every proposed component.
LFZip: Lossy compression of multivariate floating-point time series data via improved prediction
Nov 01, 2019Shubham Chandak, Kedar Tatwawadi, Chengtao Wen, Lingyun Wang, Juan Aparicio, Tsachy Weissman
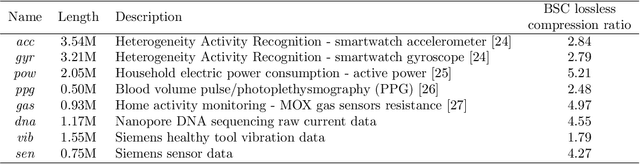
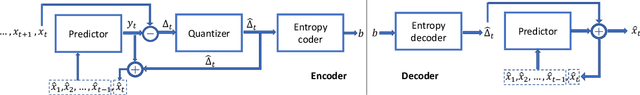

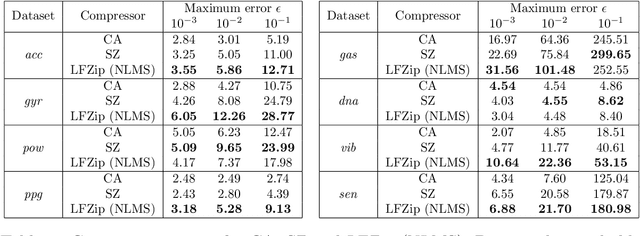
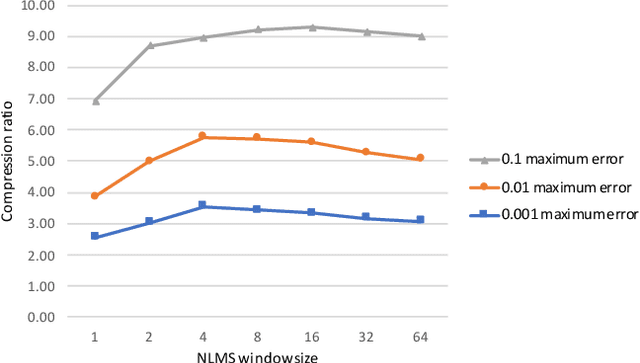
Time series data compression is emerging as an important problem with the growth in IoT devices and sensors. Due to the presence of noise in these datasets, lossy compression can often provide significant compression gains without impacting the performance of downstream applications. In this work, we propose an error-bounded lossy compressor, LFZip, for multivariate floating-point time series data that provides guaranteed reconstruction up to user-specified maximum absolute error. The compressor is based on the prediction-quantization-entropy coder framework and benefits from improved prediction using linear models and neural networks. We evaluate the compressor on several time series datasets where it outperforms the existing state-of-the-art error-bounded lossy compressors. The code and data are available at https://github.com/shubhamchandak94/LFZip
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge