Tractography with T1-weighted MRI and associated anatomical constraints on clinical quality diffusion MRI
Mar 27, 2024Tian Yu, Yunhe Li, Michael E. Kim, Chenyu Gao, Qi Yang, Leon Y. Cai, Susane M. Resnick, Lori L. Beason-Held, Daniel C. Moyer, Kurt G. Schilling, Bennett A. Landman
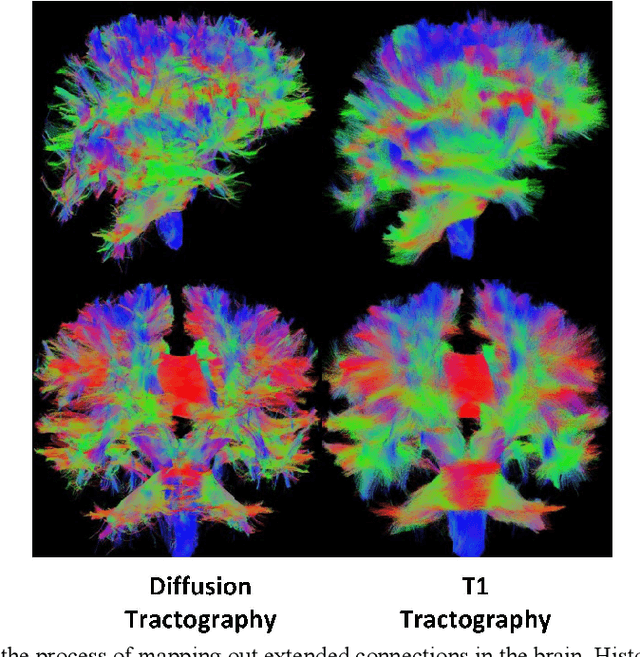
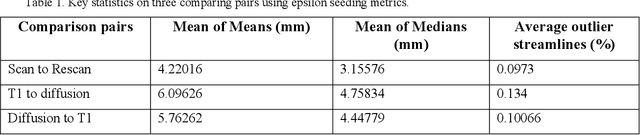
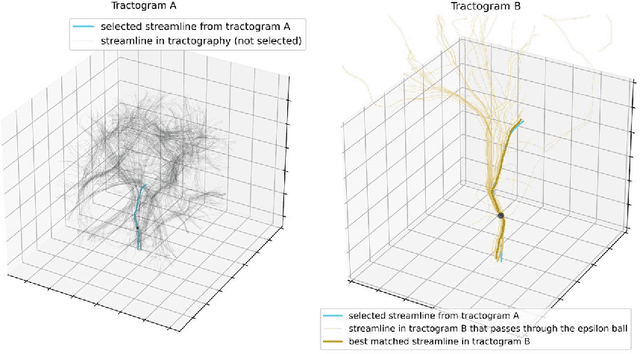
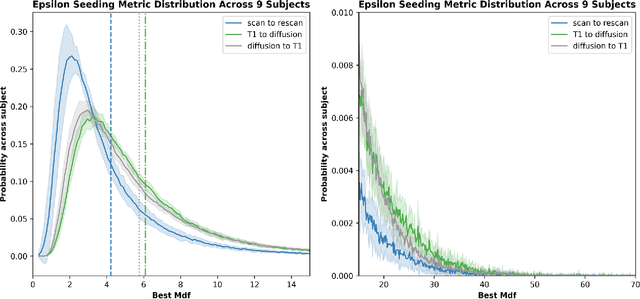
Diffusion MRI (dMRI) streamline tractography, the gold standard for in vivo estimation of brain white matter (WM) pathways, has long been considered indicative of macroscopic relationships with WM microstructure. However, recent advances in tractography demonstrated that convolutional recurrent neural networks (CoRNN) trained with a teacher-student framework have the ability to learn and propagate streamlines directly from T1 and anatomical contexts. Training for this network has previously relied on high-resolution dMRI. In this paper, we generalize the training mechanism to traditional clinical resolution data, which allows generalizability across sensitive and susceptible study populations. We train CoRNN on a small subset of the Baltimore Longitudinal Study of Aging (BLSA), which better resembles clinical protocols. Then, we define a metric, termed the epsilon ball seeding method, to compare T1 tractography and traditional diffusion tractography at the streamline level. Under this metric, T1 tractography generated by CoRNN reproduces diffusion tractography with approximately two millimeters of error.
Predicting Age from White Matter Diffusivity with Residual Learning
Nov 06, 2023Chenyu Gao, Michael E. Kim, Ho Hin Lee, Qi Yang, Nazirah Mohd Khairi, Praitayini Kanakaraj, Nancy R. Newlin, Derek B. Archer, Angela L. Jefferson, Warren D. Taylor, Brian D. Boyd, Lori L. Beason-Held, Susan M. Resnick, The BIOCARD Study Team, Yuankai Huo, Katherine D. Van Schaik, Kurt G. Schilling, Daniel Moyer, Ivana Išgum, Bennett A. Landman
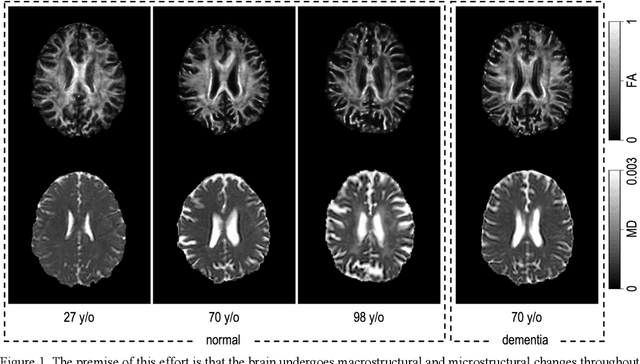
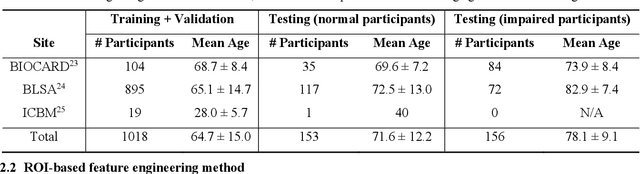
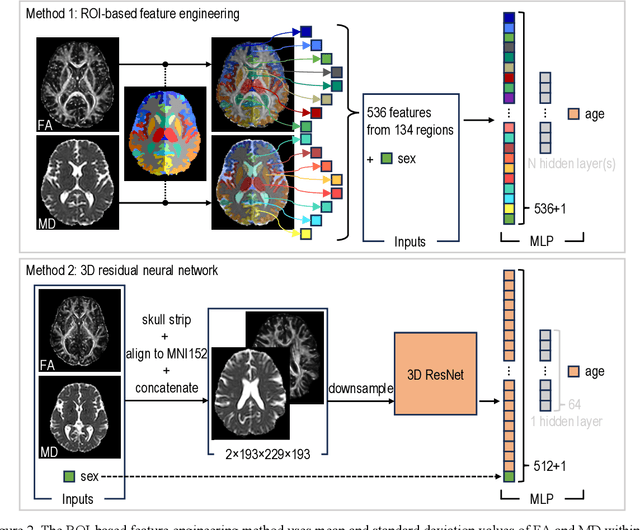
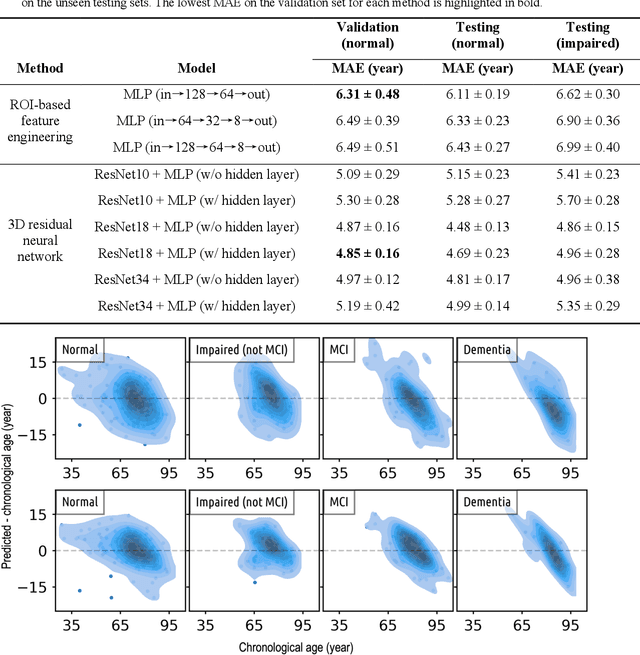
Imaging findings inconsistent with those expected at specific chronological age ranges may serve as early indicators of neurological disorders and increased mortality risk. Estimation of chronological age, and deviations from expected results, from structural MRI data has become an important task for developing biomarkers that are sensitive to such deviations. Complementary to structural analysis, diffusion tensor imaging (DTI) has proven effective in identifying age-related microstructural changes within the brain white matter, thereby presenting itself as a promising additional modality for brain age prediction. Although early studies have sought to harness DTI's advantages for age estimation, there is no evidence that the success of this prediction is owed to the unique microstructural and diffusivity features that DTI provides, rather than the macrostructural features that are also available in DTI data. Therefore, we seek to develop white-matter-specific age estimation to capture deviations from normal white matter aging. Specifically, we deliberately disregard the macrostructural information when predicting age from DTI scalar images, using two distinct methods. The first method relies on extracting only microstructural features from regions of interest. The second applies 3D residual neural networks (ResNets) to learn features directly from the images, which are non-linearly registered and warped to a template to minimize macrostructural variations. When tested on unseen data, the first method yields mean absolute error (MAE) of 6.11 years for cognitively normal participants and MAE of 6.62 years for cognitively impaired participants, while the second method achieves MAE of 4.69 years for cognitively normal participants and MAE of 4.96 years for cognitively impaired participants. We find that the ResNet model captures subtler, non-macrostructural features for brain age prediction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge