Retrieval-Based Reconstruction For Time-series Contrastive Learning
Nov 01, 2023Maxwell A. Xu, Alexander Moreno, Hui Wei, Benjamin M. Marlin, James M. Rehg
The success of self-supervised contrastive learning hinges on identifying positive data pairs that, when pushed together in embedding space, encode useful information for subsequent downstream tasks. However, in time-series, this is challenging because creating positive pairs via augmentations may break the original semantic meaning. We hypothesize that if we can retrieve information from one subsequence to successfully reconstruct another subsequence, then they should form a positive pair. Harnessing this intuition, we introduce our novel approach: REtrieval-BAsed Reconstruction (REBAR) contrastive learning. First, we utilize a convolutional cross-attention architecture to calculate the REBAR error between two different time-series. Then, through validation experiments, we show that the REBAR error is a predictor of mutual class membership, justifying its usage as a positive/negative labeler. Finally, once integrated into a contrastive learning framework, our REBAR method can learn an embedding that achieves state-of-the-art performance on downstream tasks across various modalities.
PulseImpute: A Novel Benchmark Task for Pulsative Physiological Signal Imputation
Dec 14, 2022Maxwell A. Xu, Alexander Moreno, Supriya Nagesh, V. Burak Aydemir, David W. Wetter, Santosh Kumar, James M. Rehg
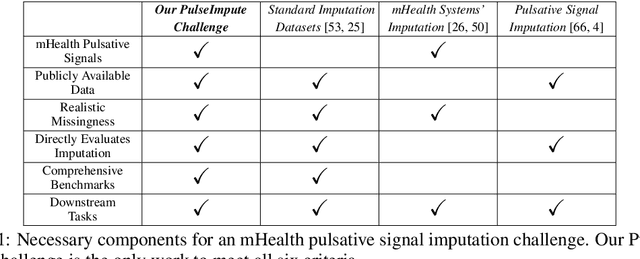
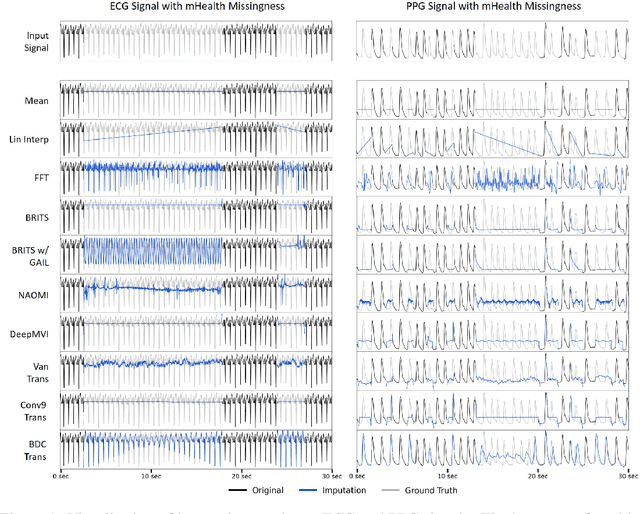
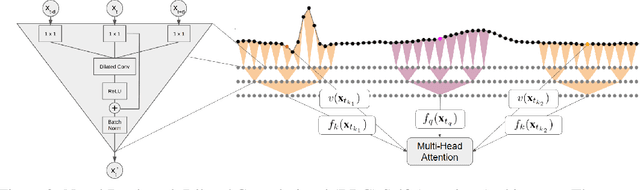

The promise of Mobile Health (mHealth) is the ability to use wearable sensors to monitor participant physiology at high frequencies during daily life to enable temporally-precise health interventions. However, a major challenge is frequent missing data. Despite a rich imputation literature, existing techniques are ineffective for the pulsative signals which comprise many mHealth applications, and a lack of available datasets has stymied progress. We address this gap with PulseImpute, the first large-scale pulsative signal imputation challenge which includes realistic mHealth missingness models, an extensive set of baselines, and clinically-relevant downstream tasks. Our baseline models include a novel transformer-based architecture designed to exploit the structure of pulsative signals. We hope that PulseImpute will enable the ML community to tackle this significant and challenging task.
Efficient Learning and Decoding of the Continuous-Time Hidden Markov Model for Disease Progression Modeling
Oct 26, 2021Yu-Ying Liu, Alexander Moreno, Maxwell A. Xu, Shuang Li, Jena C. McDaniel, Nancy C. Brady, Agata Rozga, Fuxin Li, Le Song, James M. Rehg
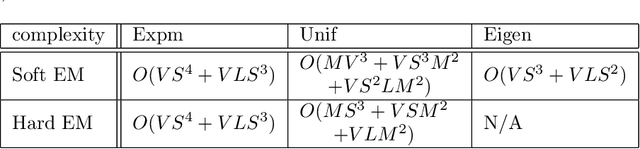
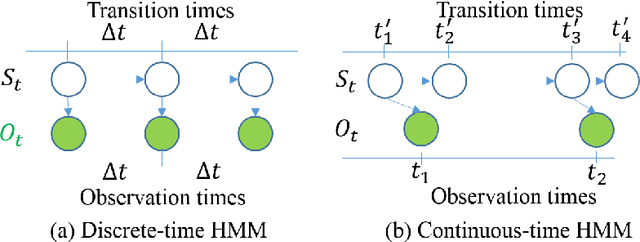

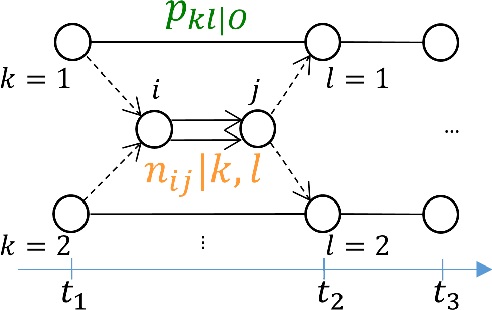
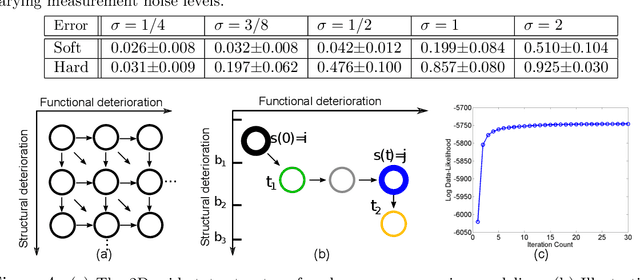
The Continuous-Time Hidden Markov Model (CT-HMM) is an attractive approach to modeling disease progression due to its ability to describe noisy observations arriving irregularly in time. However, the lack of an efficient parameter learning algorithm for CT-HMM restricts its use to very small models or requires unrealistic constraints on the state transitions. In this paper, we present the first complete characterization of efficient EM-based learning methods for CT-HMM models, as well as the first solution to decoding the optimal state transition sequence and the corresponding state dwelling time. We show that EM-based learning consists of two challenges: the estimation of posterior state probabilities and the computation of end-state conditioned statistics. We solve the first challenge by reformulating the estimation problem as an equivalent discrete time-inhomogeneous hidden Markov model. The second challenge is addressed by adapting three distinct approaches from the continuous time Markov chain (CTMC) literature to the CT-HMM domain. Additionally, we further improve the efficiency of the most efficient method by a factor of the number of states. Then, for decoding, we incorporate a state-of-the-art method from the (CTMC) literature, and extend the end-state conditioned optimal state sequence decoding to the CT-HMM case with the computation of the expected state dwelling time. We demonstrate the use of CT-HMMs with more than 100 states to visualize and predict disease progression using a glaucoma dataset and an Alzheimer's disease dataset, and to decode and visualize the most probable state transition trajectory for individuals on the glaucoma dataset, which helps to identify progressing phenotypes in a comprehensive way. Finally, we apply the CT-HMM modeling and decoding strategy to investigate the progression of language acquisition and development.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge