A unified spectra analysis workflow for the assessment of microbial contamination of ready to eat green salads: Comparative study and application of non-invasive sensors
Mar 26, 2019Panagiotis Tsakanikas, Lemonia Christina Fengou, Evanthia Manthou, Alexandra Lianou, Efstathios Z. Panagou, George John E. Nychas
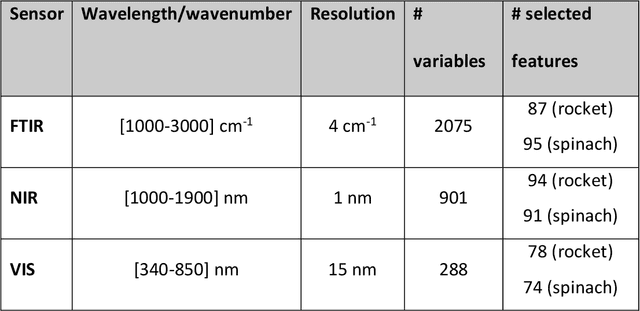
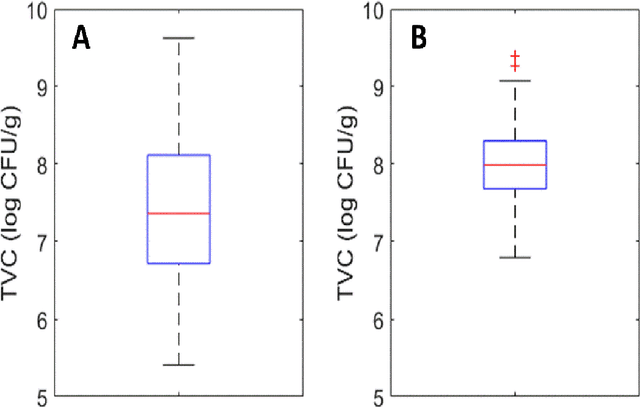
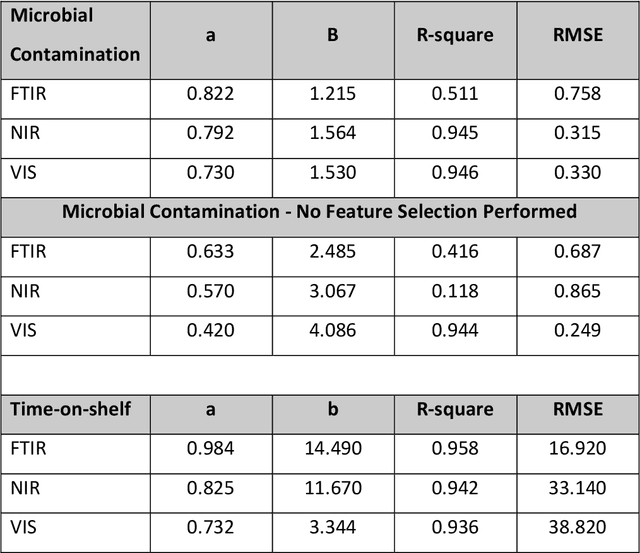
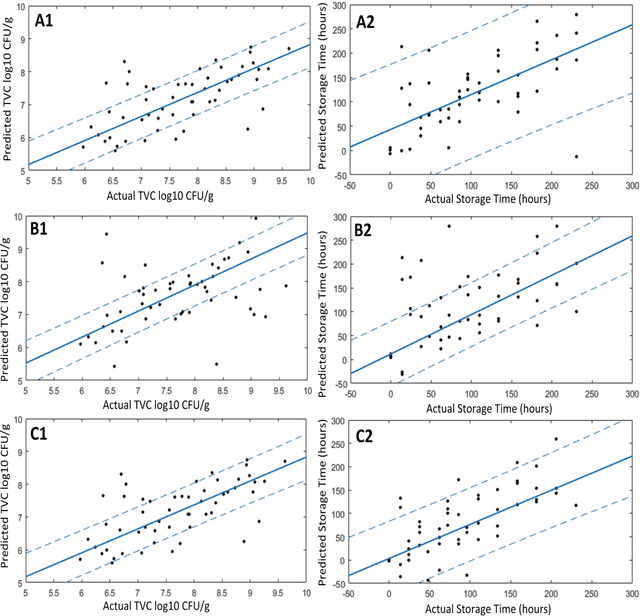
The present study provides a comparative assessment of non-invasive sensors as means of estimating the microbial contamination and time-on-shelf (i.e. storage time) of leafy green vegetables, using a novel unified spectra analysis workflow. Two fresh ready-to-eat green salads were used in the context of this study for the purpose of evaluating the efficiency and practical application of the presented workflow: rocket and baby spinach salads. The employed analysis workflow consisted of robust data normalization, powerful feature selection based on random forests regression, and selection of the number of partial least squares regression coefficients in the training process by estimating the knee-point on the explained variance plot. Training processes were based on microbiological and spectral data derived during storage of green salad samples at isothermal conditions (4, 8 and 12C), whereas testing was performed on data during storage under dynamic temperature conditions (simulating real-life temperature fluctuations in the food supply chain). Since an increasing interest in the use of non-invasive sensors in food quality assessment has been made evident in recent years, the unified spectra analysis workflow described herein, by being based on the creation/usage of limited sized featured sets, could be very useful in food-specific low-cost sensor development.
Tracking Single-Cells in Overcrowded Bacterial Colonies
Jun 22, 2017Athanasios D. Balomenos, Panagiotis Tsakanikas, Elias S. Manolakos
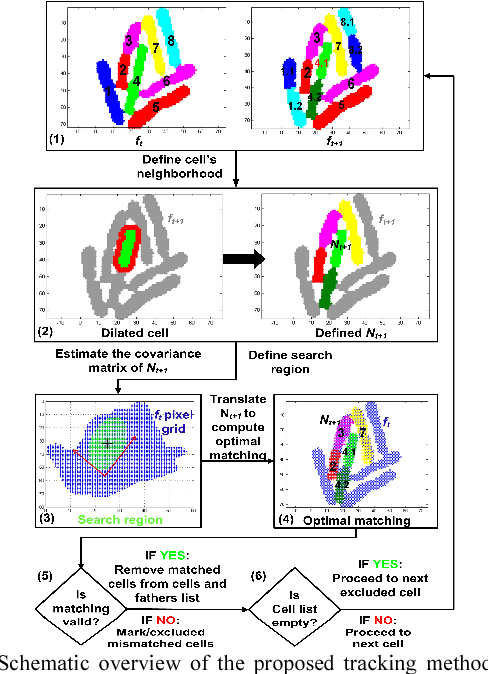

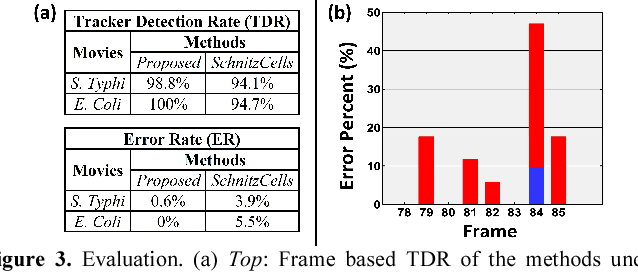
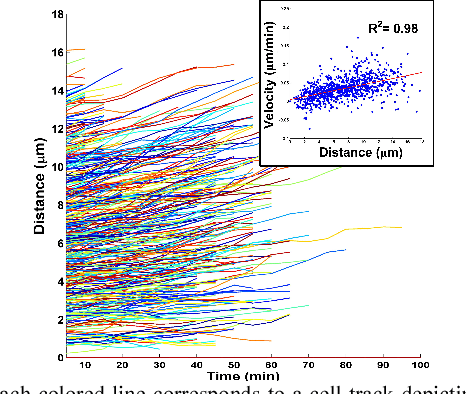
Cell tracking enables data extraction from time-lapse "cell movies" and promotes modeling biological processes at the single-cell level. We introduce a new fully automated computational strategy to track accurately cells across frames in time-lapse movies. Our method is based on a dynamic neighborhoods formation and matching approach, inspired by motion estimation algorithms for video compression. Moreover, it exploits "divide and conquer" opportunities to solve effectively the challenging cells tracking problem in overcrowded bacterial colonies. Using cell movies generated by different labs we demonstrate that the accuracy of the proposed method remains very high (exceeds 97%) even when analyzing large overcrowded microbial colonies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge