A Continuous Relaxation for Discrete Bayesian Optimization
Apr 26, 2024Richard Michael, Simon Bartels, Miguel González-Duque, Yevgen Zainchkovskyy, Jes Frellsen, Søren Hauberg, Wouter Boomsma
To optimize efficiently over discrete data and with only few available target observations is a challenge in Bayesian optimization. We propose a continuous relaxation of the objective function and show that inference and optimization can be computationally tractable. We consider in particular the optimization domain where very few observations and strict budgets exist; motivated by optimizing protein sequences for expensive to evaluate bio-chemical properties. The advantages of our approach are two-fold: the problem is treated in the continuous setting, and available prior knowledge over sequences can be incorporated directly. More specifically, we utilize available and learned distributions over the problem domain for a weighting of the Hellinger distance which yields a covariance function. We show that the resulting acquisition function can be optimized with both continuous or discrete optimization algorithms and empirically assess our method on two bio-chemical sequence optimization tasks.
Improving Adversarial Energy-Based Model via Diffusion Process
Mar 04, 2024Cong Geng, Tian Han, Peng-Tao Jiang, Hao Zhang, Jinwei Chen, Søren Hauberg, Bo Li
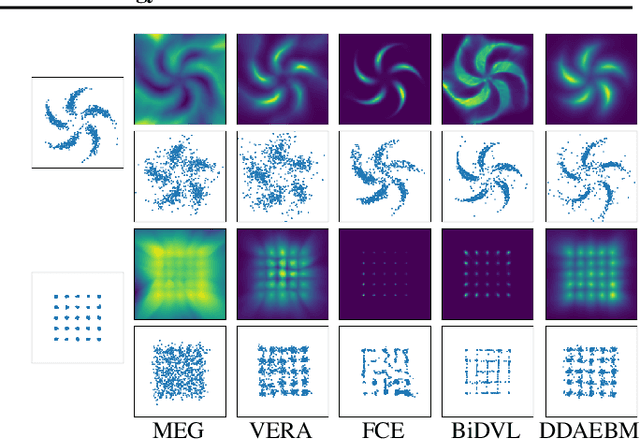

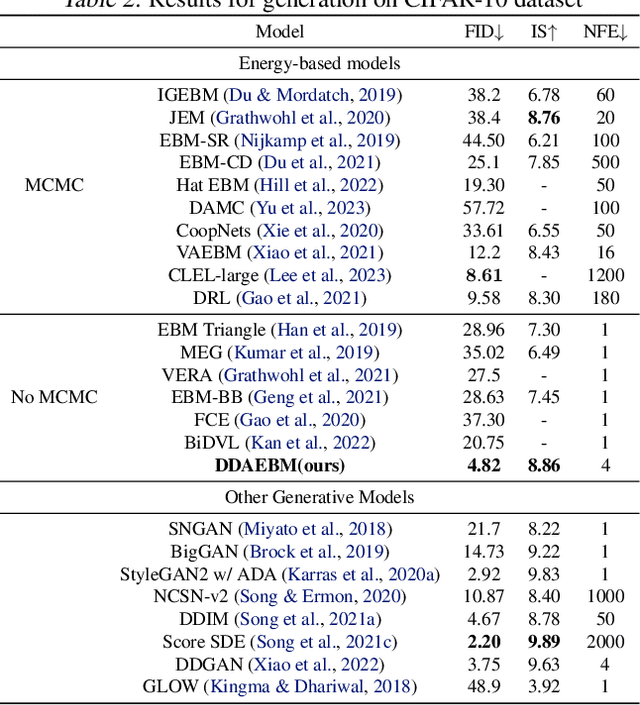

Generative models have shown strong generation ability while efficient likelihood estimation is less explored. Energy-based models~(EBMs) define a flexible energy function to parameterize unnormalized densities efficiently but are notorious for being difficult to train. Adversarial EBMs introduce a generator to form a minimax training game to avoid expensive MCMC sampling used in traditional EBMs, but a noticeable gap between adversarial EBMs and other strong generative models still exists. Inspired by diffusion-based models, we embedded EBMs into each denoising step to split a long-generated process into several smaller steps. Besides, we employ a symmetric Jeffrey divergence and introduce a variational posterior distribution for the generator's training to address the main challenges that exist in adversarial EBMs. Our experiments show significant improvement in generation compared to existing adversarial EBMs, while also providing a useful energy function for efficient density estimation.
Neural Contractive Dynamical Systems
Jan 17, 2024Hadi Beik-Mohammadi, Søren Hauberg, Georgios Arvanitidis, Nadia Figueroa, Gerhard Neumann, Leonel Rozo
Stability guarantees are crucial when ensuring a fully autonomous robot does not take undesirable or potentially harmful actions. Unfortunately, global stability guarantees are hard to provide in dynamical systems learned from data, especially when the learned dynamics are governed by neural networks. We propose a novel methodology to learn neural contractive dynamical systems, where our neural architecture ensures contraction, and hence, global stability. To efficiently scale the method to high-dimensional dynamical systems, we develop a variant of the variational autoencoder that learns dynamics in a low-dimensional latent representation space while retaining contractive stability after decoding. We further extend our approach to learning contractive systems on the Lie group of rotations to account for full-pose end-effector dynamic motions. The result is the first highly flexible learning architecture that provides contractive stability guarantees with capability to perform obstacle avoidance. Empirically, we demonstrate that our approach encodes the desired dynamics more accurately than the current state-of-the-art, which provides less strong stability guarantees.
Learning to Taste: A Multimodal Wine Dataset
Sep 07, 2023Thoranna Bender, Simon Moe Sørensen, Alireza Kashani, K. Eldjarn Hjorleifsson, Grethe Hyldig, Søren Hauberg, Serge Belongie, Frederik Warburg
We present WineSensed, a large multimodal wine dataset for studying the relations between visual perception, language, and flavor. The dataset encompasses 897k images of wine labels and 824k reviews of wines curated from the Vivino platform. It has over 350k unique vintages, annotated with year, region, rating, alcohol percentage, price, and grape composition. We obtained fine-grained flavor annotations on a subset by conducting a wine-tasting experiment with 256 participants who were asked to rank wines based on their similarity in flavor, resulting in more than 5k pairwise flavor distances. We propose a low-dimensional concept embedding algorithm that combines human experience with automatic machine similarity kernels. We demonstrate that this shared concept embedding space improves upon separate embedding spaces for coarse flavor classification (alcohol percentage, country, grape, price, rating) and aligns with the intricate human perception of flavor.
Variational Point Encoding Deformation for Dental Modeling
Jul 20, 2023Johan Ziruo Ye, Thomas Ørkild, Peter Lempel Søndergaard, Søren Hauberg




Digital dentistry has made significant advancements in recent years, yet numerous challenges remain to be addressed. In this study, we release a new extensive dataset of tooth meshes to encourage further research. Additionally, we propose Variational FoldingNet (VF-Net), which extends FoldingNet to enable probabilistic learning of point cloud representations. A key challenge in existing latent variable models for point clouds is the lack of a 1-to-1 mapping between input points and output points. Instead, they must rely on optimizing Chamfer distances, a metric that does not have a normalized distributional counterpart, preventing its usage in probabilistic models. We demonstrate that explicit minimization of Chamfer distances can be replaced by a suitable encoder, which allows us to increase computational efficiency while simplifying the probabilistic extension. Our experimental findings present empirical evidence demonstrating the superior performance of VF-Net over existing models in terms of dental scan reconstruction and extrapolation. Additionally, our investigation highlights the robustness of VF-Net's latent representations. These results underscore the promising prospects of VF-Net as an effective and reliable method for point cloud reconstruction and analysis.
Riemannian Laplace approximations for Bayesian neural networks
Jun 12, 2023Federico Bergamin, Pablo Moreno-Muñoz, Søren Hauberg, Georgios Arvanitidis




Bayesian neural networks often approximate the weight-posterior with a Gaussian distribution. However, practical posteriors are often, even locally, highly non-Gaussian, and empirical performance deteriorates. We propose a simple parametric approximate posterior that adapts to the shape of the true posterior through a Riemannian metric that is determined by the log-posterior gradient. We develop a Riemannian Laplace approximation where samples naturally fall into weight-regions with low negative log-posterior. We show that these samples can be drawn by solving a system of ordinary differential equations, which can be done efficiently by leveraging the structure of the Riemannian metric and automatic differentiation. Empirically, we demonstrate that our approach consistently improves over the conventional Laplace approximation across tasks. We further show that, unlike the conventional Laplace approximation, our method is not overly sensitive to the choice of prior, which alleviates a practical pitfall of current approaches.
On Masked Pre-training and the Marginal Likelihood
Jun 01, 2023Pablo Moreno-Muñoz, Pol G. Recasens, Søren Hauberg


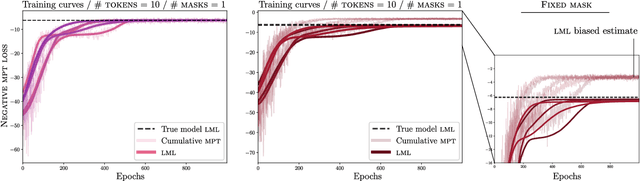

Masked pre-training removes random input dimensions and learns a model that can predict the missing values. Empirical results indicate that this intuitive form of self-supervised learning yields models that generalize very well to new domains. A theoretical understanding is, however, lacking. This paper shows that masked pre-training with a suitable cumulative scoring function corresponds to maximizing the model's marginal likelihood, which is de facto the Bayesian model selection measure of generalization. Beyond shedding light on the success of masked pre-training, this insight also suggests that Bayesian models can be trained with appropriately designed self-supervision. Empirically, we confirm the developed theory and explore the main learning principles of masked pre-training in large language models.
Laplacian Segmentation Networks: Improved Epistemic Uncertainty from Spatial Aleatoric Uncertainty
Mar 23, 2023Kilian Zepf, Selma Wanna, Marco Miani, Juston Moore, Jes Frellsen, Søren Hauberg, Aasa Feragen, Frederik Warburg




Out of distribution (OOD) medical images are frequently encountered, e.g. because of site- or scanner differences, or image corruption. OOD images come with a risk of incorrect image segmentation, potentially negatively affecting downstream diagnoses or treatment. To ensure robustness to such incorrect segmentations, we propose Laplacian Segmentation Networks (LSN) that jointly model epistemic (model) and aleatoric (data) uncertainty in image segmentation. We capture data uncertainty with a spatially correlated logit distribution. For model uncertainty, we propose the first Laplace approximation of the weight posterior that scales to large neural networks with skip connections that have high-dimensional outputs. Empirically, we demonstrate that modelling spatial pixel correlation allows the Laplacian Segmentation Network to successfully assign high epistemic uncertainty to out-of-distribution objects appearing within images.
Identifying latent distances with Finslerian geometry
Dec 20, 2022Alison Pouplin, David Eklund, Carl Henrik Ek, Søren Hauberg


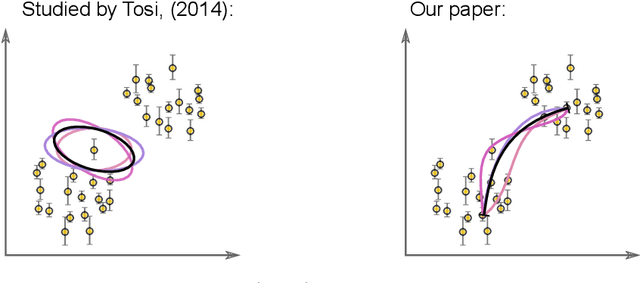

Riemannian geometry provides powerful tools to explore the latent space of generative models while preserving the inherent structure of the data manifold. Lengths, energies and volume measures can be derived from a pullback metric, defined through the immersion that maps the latent space to the data space. With this in mind, most generative models are stochastic, and so is the pullback metric. Manipulating stochastic objects is strenuous in practice. In order to perform operations such as interpolations, or measuring the distance between data points, we need a deterministic approximation of the pullback metric. In this work, we are defining a new metric as the expected length derived from the stochastic pullback metric. We show this metric is Finslerian, and we compare it with the expected pullback metric. In high dimensions, we show that the metrics converge to each other at a rate of $\mathcal{O}\left(\frac{1}{D}\right)$.
Probabilistic thermal stability prediction through sparsity promoting transformer representation
Nov 10, 2022Yevgen Zainchkovskyy, Jesper Ferkinghoff-Borg, Anja Bennett, Thomas Egebjerg, Nikolai Lorenzen, Per Jr. Greisen, Søren Hauberg, Carsten Stahlhut




Pre-trained protein language models have demonstrated significant applicability in different protein engineering task. A general usage of these pre-trained transformer models latent representation is to use a mean pool across residue positions to reduce the feature dimensions to further downstream tasks such as predicting bio-physics properties or other functional behaviours. In this paper we provide a two-fold contribution to machine learning (ML) driven drug design. Firstly, we demonstrate the power of sparsity by promoting penalization of pre-trained transformer models to secure more robust and accurate melting temperature (Tm) prediction of single-chain variable fragments with a mean absolute error of 0.23C. Secondly, we demonstrate the power of framing our prediction problem in a probabilistic framework. Specifically, we advocate for the need of adopting probabilistic frameworks especially in the context of ML driven drug design.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge