Identifying Health Risks from Family History: A Survey of Natural Language Processing Techniques
Mar 15, 2024Xiang Dai, Sarvnaz Karimi, Nathan O'Callaghan



Electronic health records include information on patients' status and medical history, which could cover the history of diseases and disorders that could be hereditary. One important use of family history information is in precision health, where the goal is to keep the population healthy with preventative measures. Natural Language Processing (NLP) and machine learning techniques can assist with identifying information that could assist health professionals in identifying health risks before a condition is developed in their later years, saving lives and reducing healthcare costs. We survey the literature on the techniques from the NLP field that have been developed to utilise digital health records to identify risks of familial diseases. We highlight that rule-based methods are heavily investigated and are still actively used for family history extraction. Still, more recent efforts have been put into building neural models based on large-scale pre-trained language models. In addition to the areas where NLP has successfully been utilised, we also identify the areas where more research is needed to unlock the value of patients' records regarding data collection, task formulation and downstream applications.
Detecting Entities in the Astrophysics Literature: A Comparison of Word-based and Span-based Entity Recognition Methods
Nov 24, 2022Xiang Dai, Sarvnaz Karimi




Information Extraction from scientific literature can be challenging due to the highly specialised nature of such text. We describe our entity recognition methods developed as part of the DEAL (Detecting Entities in the Astrophysics Literature) shared task. The aim of the task is to build a system that can identify Named Entities in a dataset composed by scholarly articles from astrophysics literature. We planned our participation such that it enables us to conduct an empirical comparison between word-based tagging and span-based classification methods. When evaluated on two hidden test sets provided by the organizer, our best-performing submission achieved $F_1$ scores of 0.8307 (validation phase) and 0.7990 (testing phase).
Cost-effective Selection of Pretraining Data: A Case Study of Pretraining BERT on Social Media
Oct 02, 2020Xiang Dai, Sarvnaz Karimi, Ben Hachey, Cecile Paris




Recent studies on domain-specific BERT models show that effectiveness on downstream tasks can be improved when models are pretrained on in-domain data. Often, the pretraining data used in these models are selected based on their subject matter, e.g., biology or computer science. Given the range of applications using social media text, and its unique language variety, we pretrain two models on tweets and forum text respectively, and empirically demonstrate the effectiveness of these two resources. In addition, we investigate how similarity measures can be used to nominate in-domain pretraining data. We publicly release our pretrained models at https://bit.ly/35RpTf0.
Searching Scientific Literature for Answers on COVID-19 Questions
Jul 06, 2020Vincent Nguyen, Maciek Rybinski, Sarvnaz Karimi, Zhenchang Xing




Finding answers related to a pandemic of a novel disease raises new challenges for information seeking and retrieval, as the new information becomes available gradually. TREC COVID search track aims to assist in creating search tools to aid scientists, clinicians, policy makers and others with similar information needs in finding reliable answers from the scientific literature. We experiment with different ranking algorithms as part of our participation in this challenge. We propose a novel method for neural retrieval, and demonstrate its effectiveness on the TREC COVID search.
An Effective Transition-based Model for Discontinuous NER
Apr 28, 2020Xiang Dai, Sarvnaz Karimi, Ben Hachey, Cecile Paris




Unlike widely used Named Entity Recognition (NER) data sets in generic domains, biomedical NER data sets often contain mentions consisting of discontinuous spans. Conventional sequence tagging techniques encode Markov assumptions that are efficient but preclude recovery of these mentions. We propose a simple, effective transition-based model with generic neural encoding for discontinuous NER. Through extensive experiments on three biomedical data sets, we show that our model can effectively recognize discontinuous mentions without sacrificing the accuracy on continuous mentions.
Figurative Usage Detection of Symptom Words to Improve Personal Health Mention Detection
Jul 04, 2019Adith Iyer, Aditya Joshi, Sarvnaz Karimi, Ross Sparks, Cecile Paris
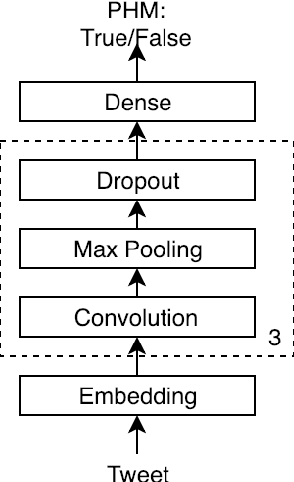
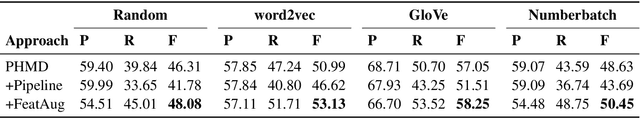
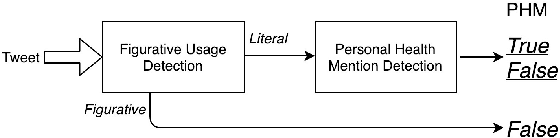
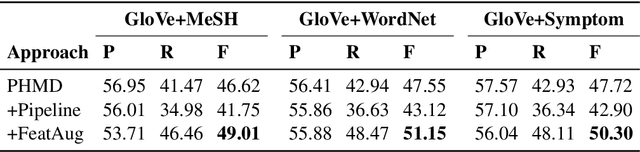
Personal health mention detection deals with predicting whether or not a given sentence is a report of a health condition. Past work mentions errors in this prediction when symptom words, i.e. names of symptoms of interest, are used in a figurative sense. Therefore, we combine a state-of-the-art figurative usage detection with CNN-based personal health mention detection. To do so, we present two methods: a pipeline-based approach and a feature augmentation-based approach. The introduction of figurative usage detection results in an average improvement of 2.21% F-score of personal health mention detection, in the case of the feature augmentation-based approach. This paper demonstrates the promise of using figurative usage detection to improve personal health mention detection.
A Comparison of Word-based and Context-based Representations for Classification Problems in Health Informatics
Jun 13, 2019Aditya Joshi, Sarvnaz Karimi, Ross Sparks, Cecile Paris, C Raina MacIntyre
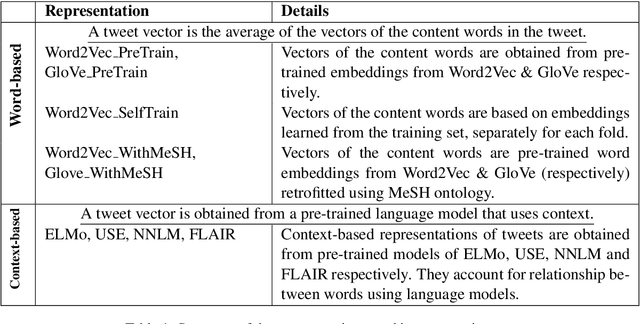
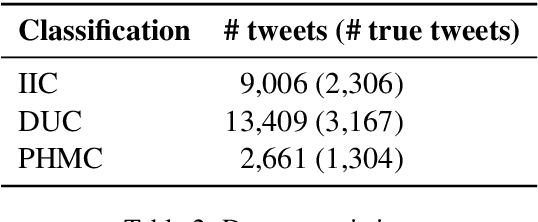
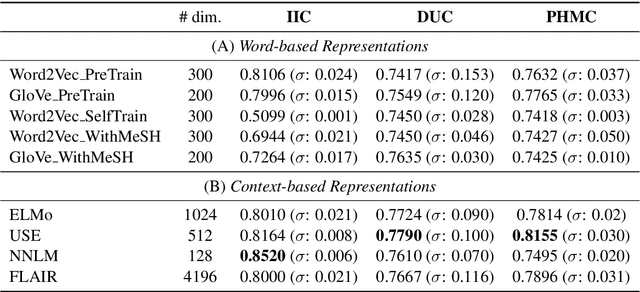
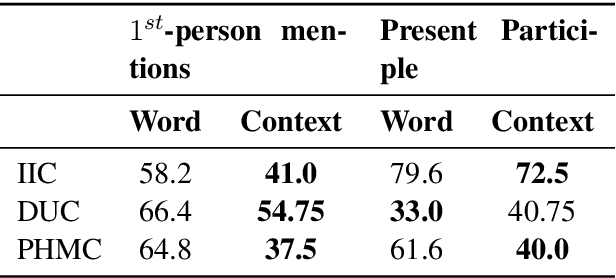
Distributed representations of text can be used as features when training a statistical classifier. These representations may be created as a composition of word vectors or as context-based sentence vectors. We compare the two kinds of representations (word versus context) for three classification problems: influenza infection classification, drug usage classification and personal health mention classification. For statistical classifiers trained for each of these problems, context-based representations based on ELMo, Universal Sentence Encoder, Neural-Net Language Model and FLAIR are better than Word2Vec, GloVe and the two adapted using the MESH ontology. There is an improvement of 2-4% in the accuracy when these context-based representations are used instead of word-based representations.
NNE: A Dataset for Nested Named Entity Recognition in English Newswire
Jun 04, 2019Nicky Ringland, Xiang Dai, Ben Hachey, Sarvnaz Karimi, Cecile Paris, James R. Curran




Named entity recognition (NER) is widely used in natural language processing applications and downstream tasks. However, most NER tools target flat annotation from popular datasets, eschewing the semantic information available in nested entity mentions. We describe NNE---a fine-grained, nested named entity dataset over the full Wall Street Journal portion of the Penn Treebank (PTB). Our annotation comprises 279,795 mentions of 114 entity types with up to 6 layers of nesting. We hope the public release of this large dataset for English newswire will encourage development of new techniques for nested NER.
Using Similarity Measures to Select Pretraining Data for NER
May 17, 2019Xiang Dai, Sarvnaz Karimi, Ben Hachey, Cecile Paris




Word vectors and Language Models (LMs) pretrained on a large amount of unlabelled data can dramatically improve various Natural Language Processing (NLP) tasks. However, the measure and impact of similarity between pretraining data and target task data are left to intuition. We propose three cost-effective measures to quantify different aspects of similarity between source pretraining and target task data. We demonstrate that these measures are good predictors of the usefulness of pretrained models for Named Entity Recognition (NER) over 30 data pairs. Results also suggest that pretrained LMs are more effective and more predictable than pretrained word vectors, but pretrained word vectors are better when pretraining data is dissimilar.
Survey of Text-based Epidemic Intelligence: A Computational Linguistic Perspective
Mar 14, 2019Aditya Joshi, Sarvnaz Karimi, Ross Sparks, Cecile Paris, C Raina MacIntyre
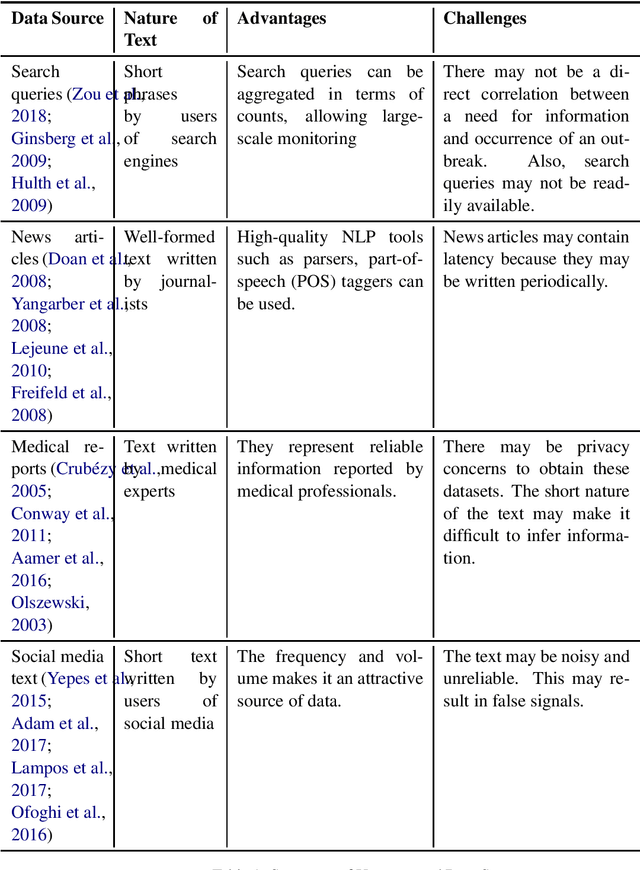
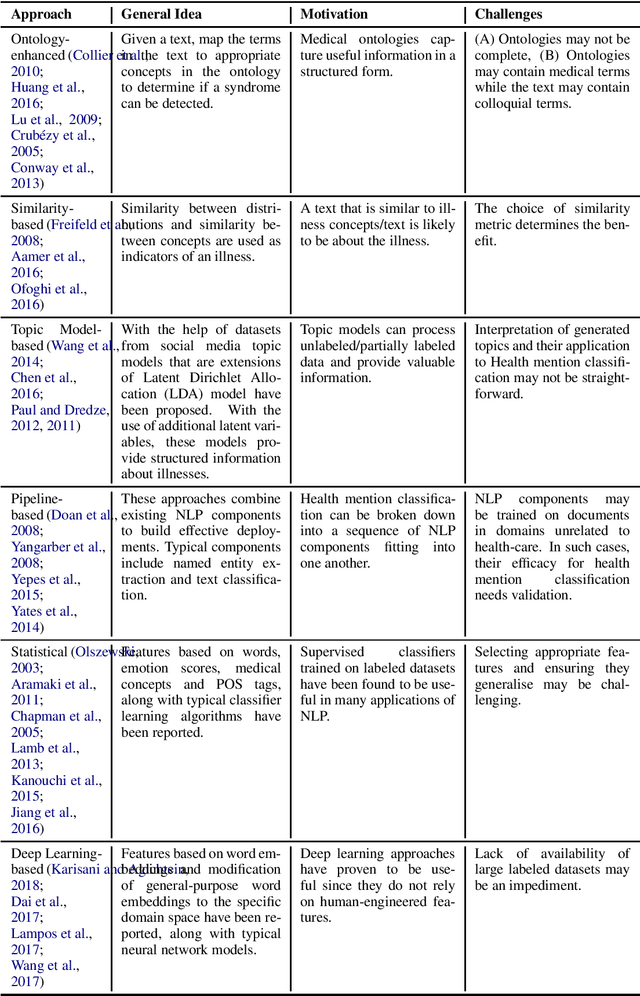
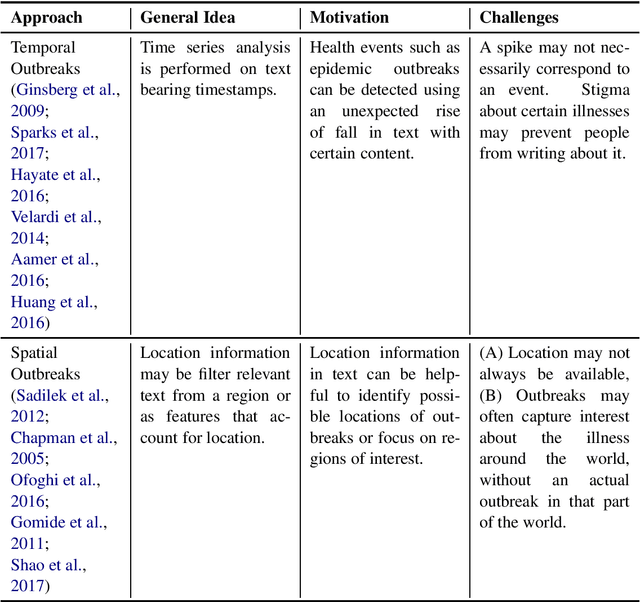
Epidemic intelligence deals with the detection of disease outbreaks using formal (such as hospital records) and informal sources (such as user-generated text on the web) of information. In this survey, we discuss approaches for epidemic intelligence that use textual datasets, referring to it as `text-based epidemic intelligence'. We view past work in terms of two broad categories: health mention classification (selecting relevant text from a large volume) and health event detection (predicting epidemic events from a collection of relevant text). The focus of our discussion is the underlying computational linguistic techniques in the two categories. The survey also provides details of the state-of-the-art in annotation techniques, resources and evaluation strategies for epidemic intelligence.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge