Multi-Task Learning for Post-transplant Cause of Death Analysis: A Case Study on Liver Transplant
Mar 30, 2023Sirui Ding, Qiaoyu Tan, Chia-yuan Chang, Na Zou, Kai Zhang, Nathan R. Hoot, Xiaoqian Jiang, Xia Hu
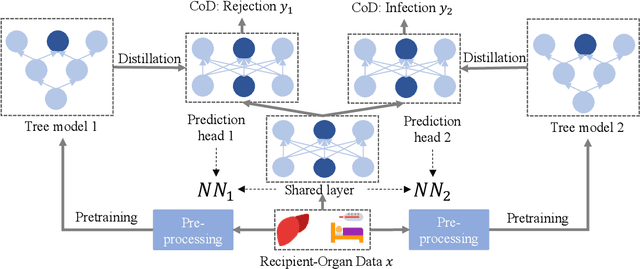
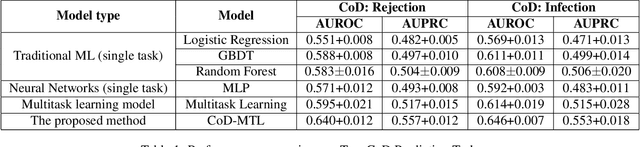
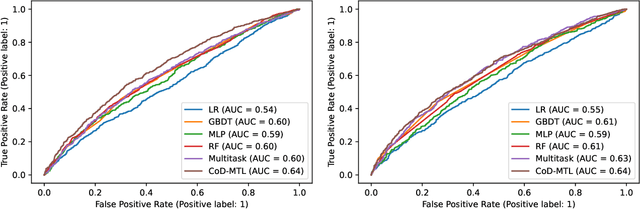
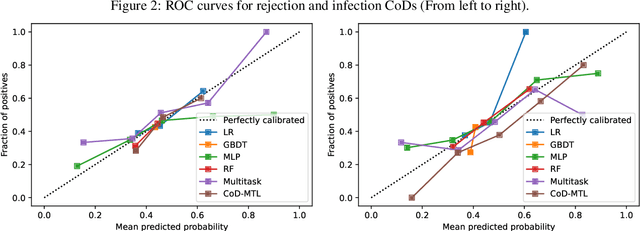
Organ transplant is the essential treatment method for some end-stage diseases, such as liver failure. Analyzing the post-transplant cause of death (CoD) after organ transplant provides a powerful tool for clinical decision making, including personalized treatment and organ allocation. However, traditional methods like Model for End-stage Liver Disease (MELD) score and conventional machine learning (ML) methods are limited in CoD analysis due to two major data and model-related challenges. To address this, we propose a novel framework called CoD-MTL leveraging multi-task learning to model the semantic relationships between various CoD prediction tasks jointly. Specifically, we develop a novel tree distillation strategy for multi-task learning, which combines the strength of both the tree model and multi-task learning. Experimental results are presented to show the precise and reliable CoD predictions of our framework. A case study is conducted to demonstrate the clinical importance of our method in the liver transplant.
Towards Fair Patient-Trial Matching via Patient-Criterion Level Fairness Constraint
Mar 24, 2023Chia-Yuan Chang, Jiayi Yuan, Sirui Ding, Qiaoyu Tan, Kai Zhang, Xiaoqian Jiang, Xia Hu, Na Zou

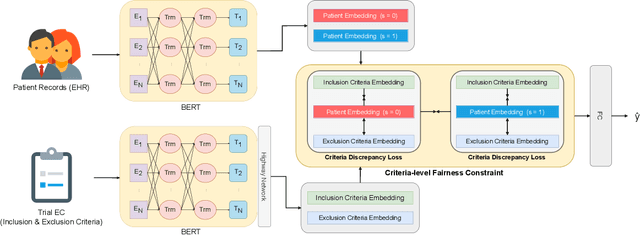
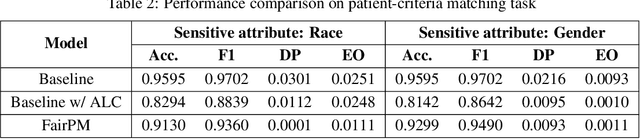
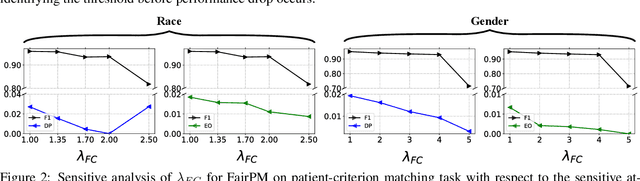
Clinical trials are indispensable in developing new treatments, but they face obstacles in patient recruitment and retention, hindering the enrollment of necessary participants. To tackle these challenges, deep learning frameworks have been created to match patients to trials. These frameworks calculate the similarity between patients and clinical trial eligibility criteria, considering the discrepancy between inclusion and exclusion criteria. Recent studies have shown that these frameworks outperform earlier approaches. However, deep learning models may raise fairness issues in patient-trial matching when certain sensitive groups of individuals are underrepresented in clinical trials, leading to incomplete or inaccurate data and potential harm. To tackle the issue of fairness, this work proposes a fair patient-trial matching framework by generating a patient-criterion level fairness constraint. The proposed framework considers the inconsistency between the embedding of inclusion and exclusion criteria among patients of different sensitive groups. The experimental results on real-world patient-trial and patient-criterion matching tasks demonstrate that the proposed framework can successfully alleviate the predictions that tend to be biased.
PheME: A deep ensemble framework for improving phenotype prediction from multi-modal data
Mar 19, 2023Shenghan Zhang, Haoxuan Li, Ruixiang Tang, Sirui Ding, Laila Rasmy, Degui Zhi, Na Zou, Xia Hu




Detailed phenotype information is fundamental to accurate diagnosis and risk estimation of diseases. As a rich source of phenotype information, electronic health records (EHRs) promise to empower diagnostic variant interpretation. However, how to accurately and efficiently extract phenotypes from the heterogeneous EHR data remains a challenge. In this work, we present PheME, an Ensemble framework using Multi-modality data of structured EHRs and unstructured clinical notes for accurate Phenotype prediction. Firstly, we employ multiple deep neural networks to learn reliable representations from the sparse structured EHR data and redundant clinical notes. A multi-modal model then aligns multi-modal features onto the same latent space to predict phenotypes. Secondly, we leverage ensemble learning to combine outputs from single-modal models and multi-modal models to improve phenotype predictions. We choose seven diseases to evaluate the phenotyping performance of the proposed framework. Experimental results show that using multi-modal data significantly improves phenotype prediction in all diseases, the proposed ensemble learning framework can further boost the performance.
Fairly Predicting Graft Failure in Liver Transplant for Organ Assigning
Feb 18, 2023Sirui Ding, Ruixiang Tang, Daochen Zha, Na Zou, Kai Zhang, Xiaoqian Jiang, Xia Hu




Liver transplant is an essential therapy performed for severe liver diseases. The fact of scarce liver resources makes the organ assigning crucial. Model for End-stage Liver Disease (MELD) score is a widely adopted criterion when making organ distribution decisions. However, it ignores post-transplant outcomes and organ/donor features. These limitations motivate the emergence of machine learning (ML) models. Unfortunately, ML models could be unfair and trigger bias against certain groups of people. To tackle this problem, this work proposes a fair machine learning framework targeting graft failure prediction in liver transplant. Specifically, knowledge distillation is employed to handle dense and sparse features by combining the advantages of tree models and neural networks. A two-step debiasing method is tailored for this framework to enhance fairness. Experiments are conducted to analyze unfairness issues in existing models and demonstrate the superiority of our method in both prediction and fairness performance.
Towards Automated Imbalanced Learning with Deep Hierarchical Reinforcement Learning
Aug 26, 2022Daochen Zha, Kwei-Herng Lai, Qiaoyu Tan, Sirui Ding, Na Zou, Xia Hu




Imbalanced learning is a fundamental challenge in data mining, where there is a disproportionate ratio of training samples in each class. Over-sampling is an effective technique to tackle imbalanced learning through generating synthetic samples for the minority class. While numerous over-sampling algorithms have been proposed, they heavily rely on heuristics, which could be sub-optimal since we may need different sampling strategies for different datasets and base classifiers, and they cannot directly optimize the performance metric. Motivated by this, we investigate developing a learning-based over-sampling algorithm to optimize the classification performance, which is a challenging task because of the huge and hierarchical decision space. At the high level, we need to decide how many synthetic samples to generate. At the low level, we need to determine where the synthetic samples should be located, which depends on the high-level decision since the optimal locations of the samples may differ for different numbers of samples. To address the challenges, we propose AutoSMOTE, an automated over-sampling algorithm that can jointly optimize different levels of decisions. Motivated by the success of SMOTE~\cite{chawla2002smote} and its extensions, we formulate the generation process as a Markov decision process (MDP) consisting of three levels of policies to generate synthetic samples within the SMOTE search space. Then we leverage deep hierarchical reinforcement learning to optimize the performance metric on the validation data. Extensive experiments on six real-world datasets demonstrate that AutoSMOTE significantly outperforms the state-of-the-art resampling algorithms. The code is at https://github.com/daochenzha/autosmote
AutoVideo: An Automated Video Action Recognition System
Aug 10, 2021Daochen Zha, Zaid Pervaiz Bhat, Yi-Wei Chen, Yicheng Wang, Sirui Ding, Anmoll Kumar Jain, Mohammad Qazim Bhat, Kwei-Herng Lai, Jiaben Chen, Na Zou, Xia Hu
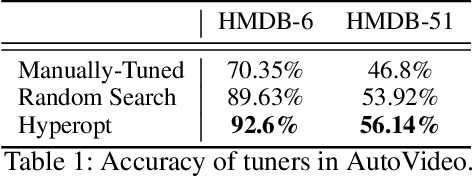
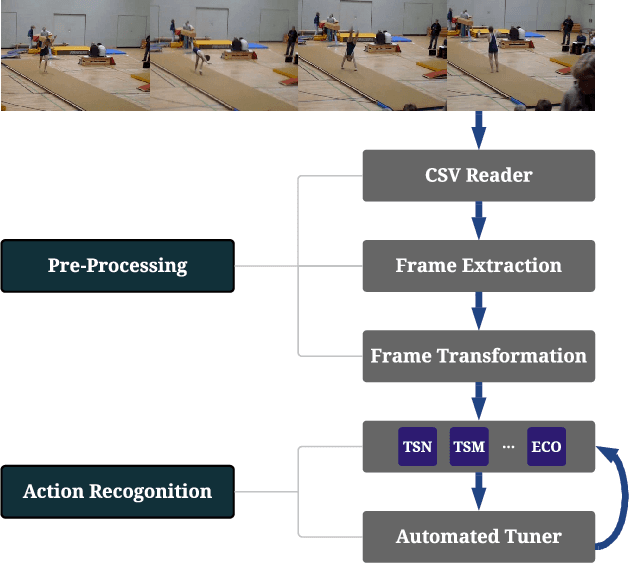

Action recognition is a crucial task for video understanding. In this paper, we present AutoVideo, a Python system for automated video action recognition. It currently supports seven action recognition algorithms and various pre-processing modules. Unlike the existing libraries that only provide model zoos, AutoVideo is built with the standard pipeline language. The basic building block is primitive, which wraps a pre-processing module or an algorithm with some hyperparameters. AutoVideo is highly modular and extendable. It can be easily combined with AutoML searchers. The pipeline language is quite general so that we can easily enrich AutoVideo with algorithms for various other video-related tasks in the future. AutoVideo is released under MIT license at https://github.com/datamllab/autovideo
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge