Speech emotion recognition from voice messages recorded in the wild
Mar 04, 2024Lucía Gómez-Zaragozá, Óscar Valls, Rocío del Amor, María José Castro-Bleda, Valery Naranjo, Mariano Alcañiz Raya, Javier Marín-Morales
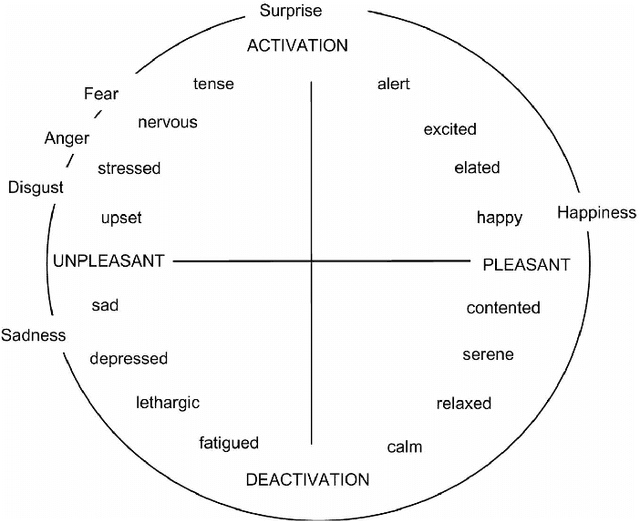
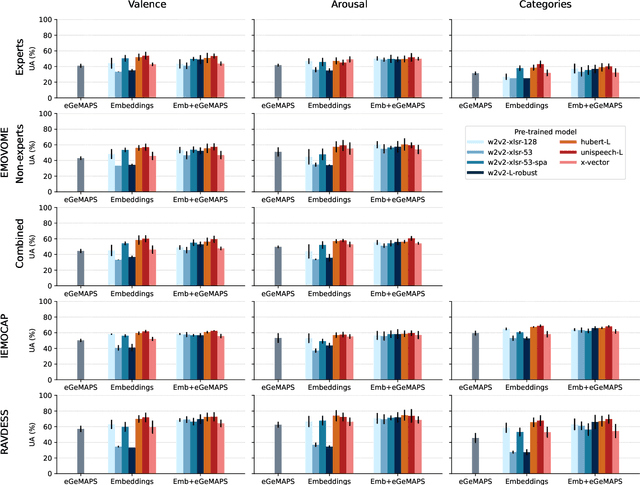
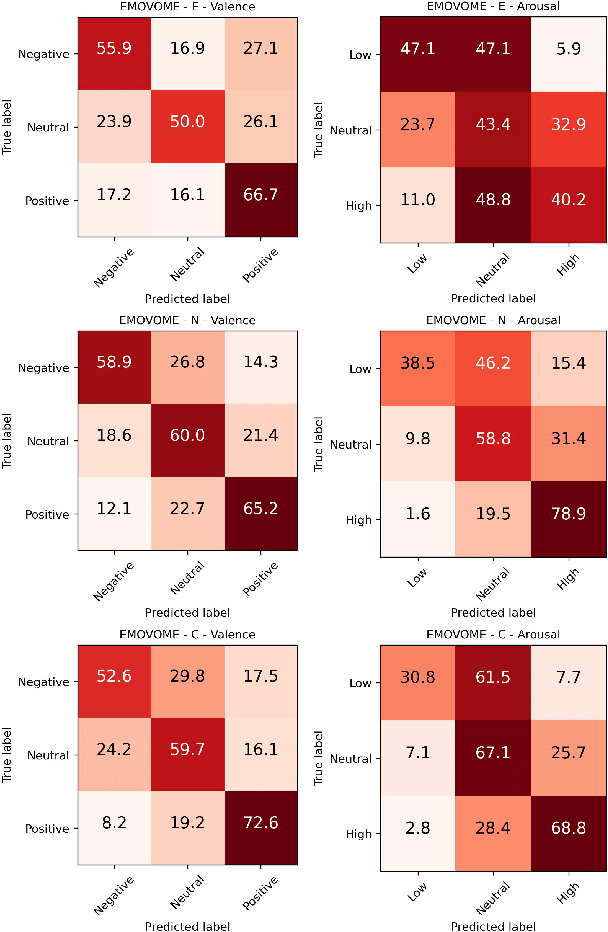

Emotion datasets used for Speech Emotion Recognition (SER) often contain acted or elicited speech, limiting their applicability in real-world scenarios. In this work, we used the Emotional Voice Messages (EMOVOME) database, including spontaneous voice messages from conversations of 100 Spanish speakers on a messaging app, labeled in continuous and discrete emotions by expert and non-expert annotators. We created speaker-independent SER models using the eGeMAPS features, transformer-based models and their combination. We compared the results with reference databases and analyzed the influence of annotators and gender fairness. The pre-trained Unispeech-L model and its combination with eGeMAPS achieved the highest results, with 61.64% and 55.57% Unweighted Accuracy (UA) for 3-class valence and arousal prediction respectively, a 10% improvement over baseline models. For the emotion categories, 42.58% UA was obtained. EMOVOME performed lower than the acted RAVDESS database. The elicited IEMOCAP database also outperformed EMOVOME in the prediction of emotion categories, while similar results were obtained in valence and arousal. Additionally, EMOVOME outcomes varied with annotator labels, showing superior results and better fairness when combining expert and non-expert annotations. This study significantly contributes to the evaluation of SER models in real-life situations, advancing in the development of applications for analyzing spontaneous voice messages.
Emotional Voice Messages (EMOVOME) database: emotion recognition in spontaneous voice messages
Feb 27, 2024Lucía Gómez Zaragozá, Rocío del Amor, Elena Parra Vargas, Valery Naranjo, Mariano Alcañiz Raya, Javier Marín-Morales
Emotional Voice Messages (EMOVOME) is a spontaneous speech dataset containing 999 audio messages from real conversations on a messaging app from 100 Spanish speakers, gender balanced. Voice messages were produced in-the-wild conditions before participants were recruited, avoiding any conscious bias due to laboratory environment. Audios were labeled in valence and arousal dimensions by three non-experts and two experts, which were then combined to obtain a final label per dimension. The experts also provided an extra label corresponding to seven emotion categories. To set a baseline for future investigations using EMOVOME, we implemented emotion recognition models using both speech and audio transcriptions. For speech, we used the standard eGeMAPS feature set and support vector machines, obtaining 49.27% and 44.71% unweighted accuracy for valence and arousal respectively. For text, we fine-tuned a multilingual BERT model and achieved 61.15% and 47.43% unweighted accuracy for valence and arousal respectively. This database will significantly contribute to research on emotion recognition in the wild, while also providing a unique natural and freely accessible resource for Spanish.
Siamese Content-based Search Engine for a More Transparent Skin and Breast Cancer Diagnosis through Histological Imaging
Jan 16, 2024Zahra Tabatabaei, Adrián Colomer, JAvier Oliver Moll, Valery Naranjo
Computer Aid Diagnosis (CAD) has developed digital pathology with Deep Learning (DL)-based tools to assist pathologists in decision-making. Content-Based Histopathological Image Retrieval (CBHIR) is a novel tool to seek highly correlated patches in terms of similarity in histopathological features. In this work, we proposed two CBHIR approaches on breast (Breast-twins) and skin cancer (Skin-twins) data sets for robust and accurate patch-level retrieval, integrating a custom-built Siamese network as a feature extractor. The proposed Siamese network is able to generalize for unseen images by focusing on the similar histopathological features of the input pairs. The proposed CBHIR approaches are evaluated on the Breast (public) and Skin (private) data sets with top K accuracy. Finding the optimum amount of K is challenging, but also, as much as K increases, the dissimilarity between the query and the returned images increases which might mislead the pathologists. To the best of the author's belief, this paper is tackling this issue for the first time on histopathological images by evaluating the top first retrieved images. The Breast-twins model achieves 70% of the F1score at the top first, which exceeds the other state-of-the-art methods at a higher amount of K such as 5 and 400. Skin-twins overpasses the recently proposed Convolutional Auto Encoder (CAE) by 67%, increasing the precision. Besides, the Skin-twins model tackles the challenges of Spitzoid Tumors of Uncertain Malignant Potential (STUMP) to assist pathologists with retrieving top K images and their corresponding labels. So, this approach can offer a more explainable CAD tool to pathologists in terms of transparency, trustworthiness, or reliability among other characteristics.
Attention to detail: inter-resolution knowledge distillation
Jan 11, 2024Rocío del Amor, Julio Silva-Rodríguez, Adrián Colomer, Valery Naranjo
The development of computer vision solutions for gigapixel images in digital pathology is hampered by significant computational limitations due to the large size of whole slide images. In particular, digitizing biopsies at high resolutions is a time-consuming process, which is necessary due to the worsening results from the decrease in image detail. To alleviate this issue, recent literature has proposed using knowledge distillation to enhance the model performance at reduced image resolutions. In particular, soft labels and features extracted at the highest magnification level are distilled into a model that takes lower-magnification images as input. However, this approach fails to transfer knowledge about the most discriminative image regions in the classification process, which may be lost when the resolution is decreased. In this work, we propose to distill this information by incorporating attention maps during training. In particular, our formulation leverages saliency maps of the target class via grad-CAMs, which guides the lower-resolution Student model to match the Teacher distribution by minimizing the l2 distance between them. Comprehensive experiments on prostate histology image grading demonstrate that the proposed approach substantially improves the model performance across different image resolutions compared to previous literature.
HistoColAi: An Open-Source Web Platform for Collaborative Digital Histology Image Annotation with AI-Driven Predictive Integration
Jul 11, 2023Cristian Camilo Pulgarín-Ospina, Rocío del Amor, Adrián Colomera, Julio Silva-Rodríguez, Valery Naranjo
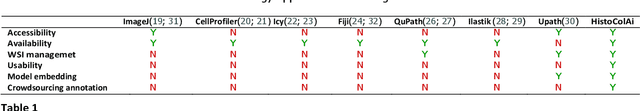
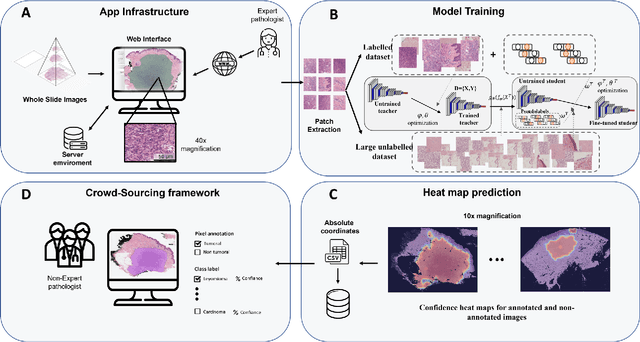
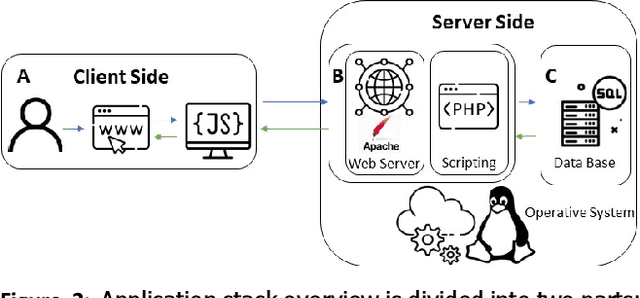
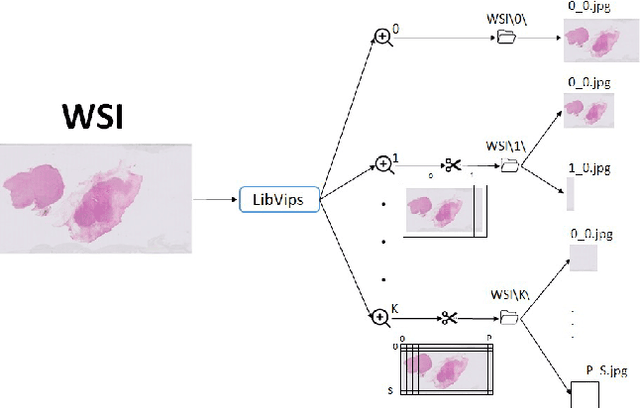
Digital pathology has become a standard in the pathology workflow due to its many benefits. These include the level of detail of the whole slide images generated and the potential immediate sharing of cases between hospitals. Recent advances in deep learning-based methods for image analysis make them of potential aid in digital pathology. However, a major limitation in developing computer-aided diagnostic systems for pathology is the lack of an intuitive and open web application for data annotation. This paper proposes a web service that efficiently provides a tool to visualize and annotate digitized histological images. In addition, to show and validate the tool, in this paper we include a use case centered on the diagnosis of spindle cell skin neoplasm for multiple annotators. A usability study of the tool is also presented, showing the feasibility of the developed tool.
Towards More Transparent and Accurate Cancer Diagnosis with an Unsupervised CAE Approach
May 19, 2023Zahra Tabatabaei, Adrian Colomer, Javier Oliver Moll, Valery Naranjo

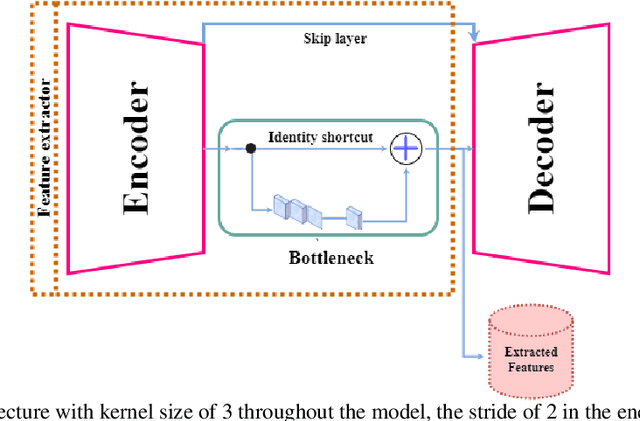
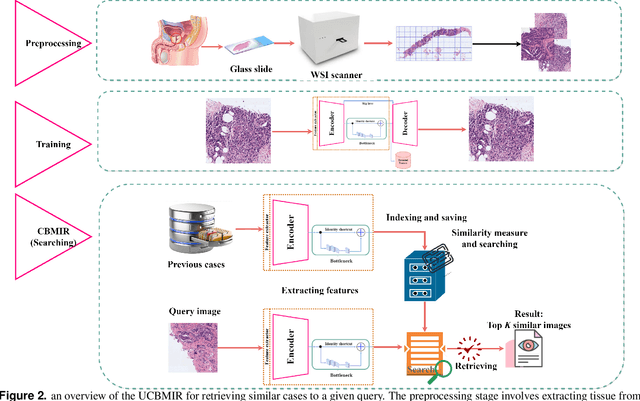
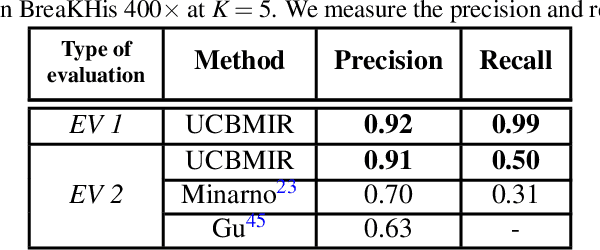
Digital pathology has revolutionized cancer diagnosis by leveraging Content-Based Medical Image Retrieval (CBMIR) for analyzing histopathological Whole Slide Images (WSIs). CBMIR enables searching for similar content, enhancing diagnostic reliability and accuracy. In 2020, breast and prostate cancer constituted 11.7% and 14.1% of cases, respectively, as reported by the Global Cancer Observatory (GCO). The proposed Unsupervised CBMIR (UCBMIR) replicates the traditional cancer diagnosis workflow, offering a dependable method to support pathologists in WSI-based diagnostic conclusions. This approach alleviates pathologists' workload, potentially enhancing diagnostic efficiency. To address the challenge of the lack of labeled histopathological images in CBMIR, a customized unsupervised Convolutional Auto Encoder (CAE) was developed, extracting 200 features per image for the search engine component. UCBMIR was evaluated using widely-used numerical techniques in CBMIR, alongside visual evaluation and comparison with a classifier. The validation involved three distinct datasets, with an external evaluation demonstrating its effectiveness. UCBMIR outperformed previous studies, achieving a top 5 recall of 99% and 80% on BreaKHis and SICAPv2, respectively, using the first evaluation technique. Precision rates of 91% and 70% were achieved for BreaKHis and SICAPv2, respectively, using the second evaluation technique. Furthermore, UCBMIR demonstrated the capability to identify various patterns in patches, achieving an 81% accuracy in the top 5 when tested on an external image from Arvaniti.
WWFedCBMIR: World-Wide Federated Content-Based Medical Image Retrieval
May 05, 2023Zahra Tabatabaei, Yuandou Wang, Adrián Colomer, Javier Oliver Moll, Zhiming Zhao, Valery Naranjo




The paper proposes a Federated Content-Based Medical Image Retrieval (FedCBMIR) platform that utilizes Federated Learning (FL) to address the challenges of acquiring a diverse medical data set for training CBMIR models. CBMIR assists pathologists in diagnosing breast cancer more rapidly by identifying similar medical images and relevant patches in prior cases compared to traditional cancer detection methods. However, CBMIR in histopathology necessitates a pool of Whole Slide Images (WSIs) to train to extract an optimal embedding vector that leverages search engine performance, which may not be available in all centers. The strict regulations surrounding data sharing in medical data sets also hinder research and model development, making it difficult to collect a rich data set. The proposed FedCBMIR distributes the model to collaborative centers for training without sharing the data set, resulting in shorter training times than local training. FedCBMIR was evaluated in two experiments with three scenarios on BreaKHis and Camelyon17 (CAM17). The study shows that the FedCBMIR method increases the F1-Score (F1S) of each client to 98%, 96%, 94%, and 97% in the BreaKHis experiment with a generalized model of four magnifications and does so in 6.30 hours less time than total local training. FedCBMIR also achieves 98% accuracy with CAM17 in 2.49 hours less training time than local training, demonstrating that our FedCBMIR is both fast and accurate for both pathologists and engineers. In addition, our FedCBMIR provides similar images with higher magnification for non-developed countries where participate in the worldwide FedCBMIR with developed countries to facilitate mitosis measuring in breast cancer diagnosis. We evaluate this scenario by scattering BreaKHis into four centers with different magnifications.
Self-supervised learning of a tailored Convolutional Auto Encoder for histopathological prostate grading
Mar 21, 2023Zahra Tabatabaei, Adrian colomer, Kjersti Engan, Javier Oliver, Valery Naranjo
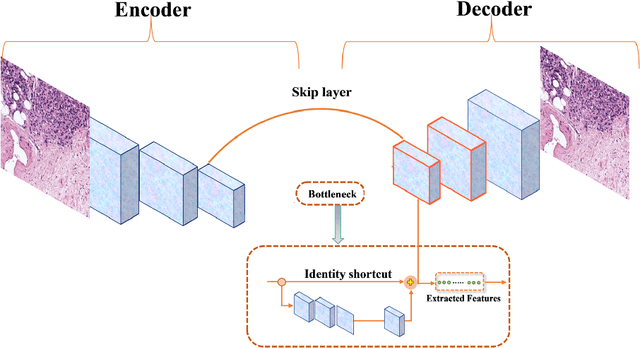

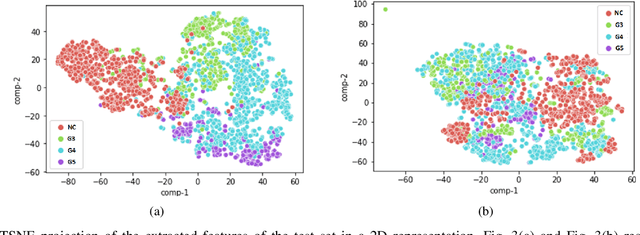
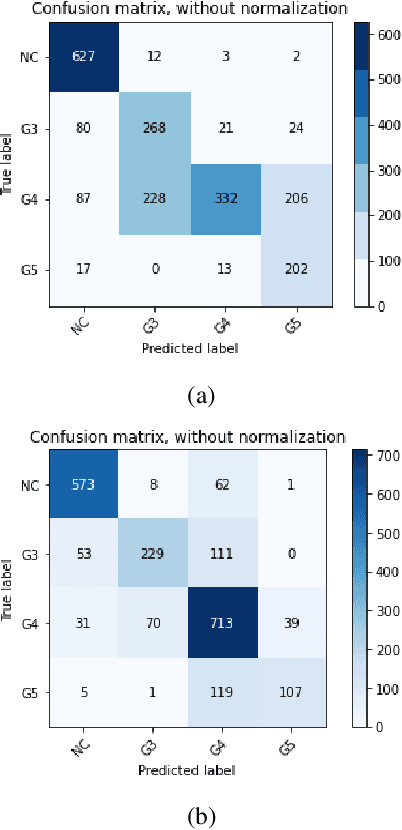
According to GLOBOCAN 2020, prostate cancer is the second most common cancer in men worldwide and the fourth most prevalent cancer overall. For pathologists, grading prostate cancer is challenging, especially when discriminating between Grade 3 (G3) and Grade 4 (G4). This paper proposes a Self-Supervised Learning (SSL) framework to classify prostate histopathological images when labeled images are scarce. In particular, a tailored Convolutional Auto Encoder (CAE) is trained to reconstruct 128x128x3 patches of prostate cancer Whole Slide Images (WSIs) as a pretext task. The downstream task of the proposed SSL paradigm is the automatic grading of histopathological patches of prostate cancer. The presented framework reports promising results on the validation set, obtaining an overall accuracy of 83% and on the test set, achieving an overall accuracy value of 76% with F1-score of 77% in G4.
Challenging mitosis detection algorithms: Global labels allow centroid localization
Nov 30, 2022Claudio Fernandez-Martín, Umay Kiraz, Julio Silva-Rodríguez, Sandra Morales, Emiel Janssen, Valery Naranjo
Mitotic activity is a crucial proliferation biomarker for the diagnosis and prognosis of different types of cancers. Nevertheless, mitosis counting is a cumbersome process for pathologists, prone to low reproducibility, due to the large size of augmented biopsy slides, the low density of mitotic cells, and pattern heterogeneity. To improve reproducibility, deep learning methods have been proposed in the last years using convolutional neural networks. However, these methods have been hindered by the process of data labelling, which usually solely consist of the mitosis centroids. Therefore, current literature proposes complex algorithms with multiple stages to refine the labels at pixel level, and to reduce the number of false positives. In this work, we propose to avoid complex scenarios, and we perform the localization task in a weakly supervised manner, using only image-level labels on patches. The results obtained on the publicly available TUPAC16 dataset are competitive with state-of-the-art methods, using only one training phase. Our method achieves an F1-score of 0.729 and challenges the efficiency of previous methods, which required multiple stages and strong mitosis location information.
* Presented at IDEAL 2022
DCASE 2022: Comparative Analysis Of CNNs For Acoustic Scene Classification Under Low-Complexity Considerations
Jun 16, 2022Josep Zaragoza-Paredes, Javier Naranjo-Alcazar, Valery Naranjo, Pedro Zuccarello
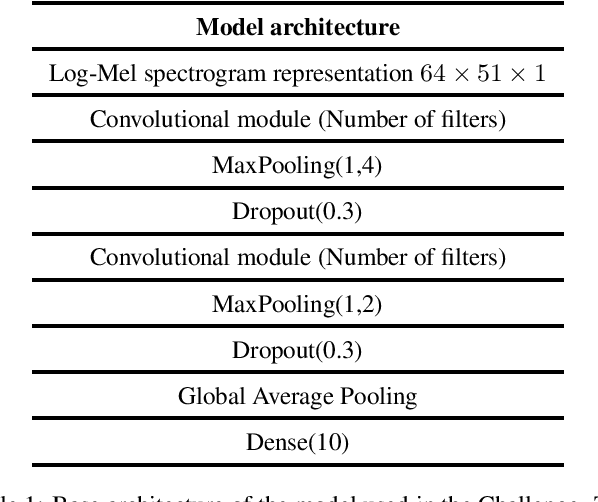

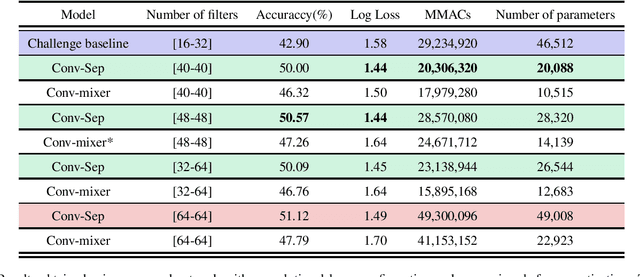
Acoustic scene classification is an automatic listening problem that aims to assign an audio recording to a pre-defined scene based on its audio data. Over the years (and in past editions of the DCASE) this problem has often been solved with techniques known as ensembles (use of several machine learning models to combine their predictions in the inference phase). While these solutions can show performance in terms of accuracy, they can be very expensive in terms of computational capacity, making it impossible to deploy them in IoT devices. Due to the drift in this field of study, this task has two limitations in terms of model complexity. It should be noted that there is also the added complexity of mismatching devices (the audios provided are recorded by different sources of information). This technical report makes a comparative study of two different network architectures: conventional CNN and Conv-mixer. Although both networks exceed the baseline required by the competition, the conventional CNN shows a higher performance, exceeding the baseline by 8 percentage points. Solutions based on Conv-mixer architectures show worse performance although they are much lighter solutions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge