Generating Medical Prescriptions with Conditional Transformer
Nov 18, 2023Samuel Belkadi, Nicolo Micheletti, Lifeng Han, Warren Del-Pinto, Goran Nenadic
Access to real-world medication prescriptions is essential for medical research and healthcare quality improvement. However, access to real medication prescriptions is often limited due to the sensitive nature of the information expressed. Additionally, manually labelling these instructions for training and fine-tuning Natural Language Processing (NLP) models can be tedious and expensive. We introduce a novel task-specific model architecture, Label-To-Text-Transformer (\textbf{LT3}), tailored to generate synthetic medication prescriptions based on provided labels, such as a vocabulary list of medications and their attributes. LT3 is trained on a set of around 2K lines of medication prescriptions extracted from the MIMIC-III database, allowing the model to produce valuable synthetic medication prescriptions. We evaluate LT3's performance by contrasting it with a state-of-the-art Pre-trained Language Model (PLM), T5, analysing the quality and diversity of generated texts. We deploy the generated synthetic data to train the SpacyNER model for the Named Entity Recognition (NER) task over the n2c2-2018 dataset. The experiments show that the model trained on synthetic data can achieve a 96-98\% F1 score at Label Recognition on Drug, Frequency, Route, Strength, and Form. LT3 codes and data will be shared at \url{https://github.com/HECTA-UoM/Label-To-Text-Transformer}
Exploring the Consistency, Quality and Challenges in Manual and Automated Coding of Free-text Diagnoses from Hospital Outpatient Letters
Nov 17, 2023Warren Del-Pinto, George Demetriou, Meghna Jani, Rikesh Patel, Leanne Gray, Alex Bulcock, Niels Peek, Andrew S. Kanter, William G Dixon, Goran Nenadic
Coding of unstructured clinical free-text to produce interoperable structured data is essential to improve direct care, support clinical communication and to enable clinical research.However, manual clinical coding is difficult and time consuming, which motivates the development and use of natural language processing for automated coding. This work evaluates the quality and consistency of both manual and automated clinical coding of diagnoses from hospital outpatient letters. Using 100 randomly selected letters, two human clinicians performed coding of diagnosis lists to SNOMED CT. Automated coding was also performed using IMO's Concept Tagger. A gold standard was constructed by a panel of clinicians from a subset of the annotated diagnoses. This was used to evaluate the quality and consistency of both manual and automated coding via (1) a distance-based metric, treating SNOMED CT as a graph, and (2) a qualitative metric agreed upon by the panel of clinicians. Correlation between the two metrics was also evaluated. Comparing human and computer-generated codes to the gold standard, the results indicate that humans slightly out-performed automated coding, while both performed notably better when there was only a single diagnosis contained in the free-text description. Automated coding was considered acceptable by the panel of clinicians in approximately 90% of cases.
Signature-Based Abduction for Expressive Description Logics -- Technical Report
Jul 08, 2020Patrick Koopmann, Warren Del-Pinto, Sophie Tourret, Renate A. Schmidt
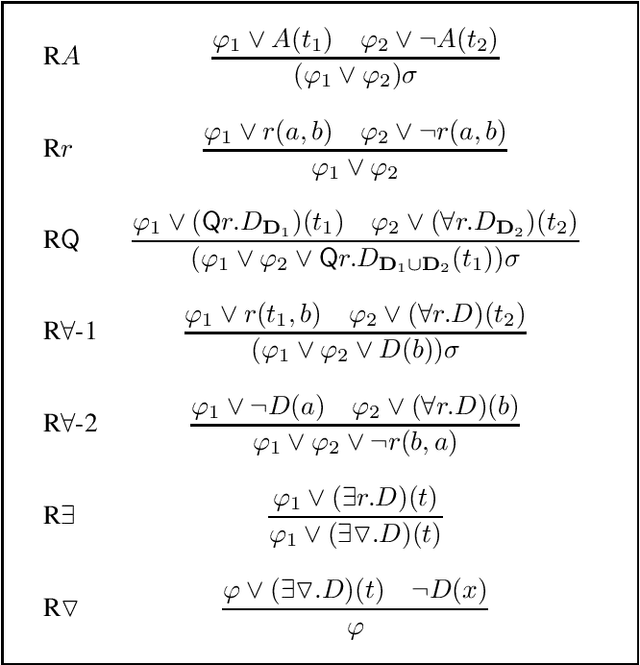

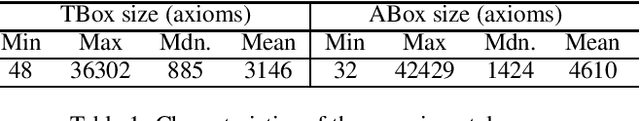

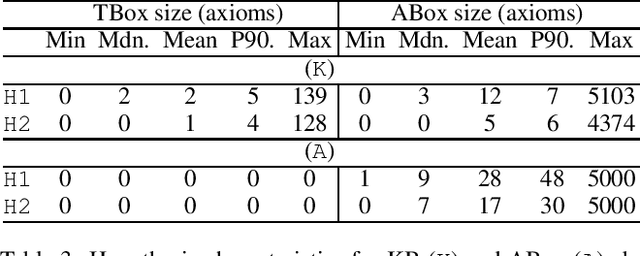
Signature-based abduction aims at building hypotheses over a specified set of names, the signature, that explain an observation relative to some background knowledge. This type of abduction is useful for tasks such as diagnosis, where the vocabulary used for observed symptoms differs from the vocabulary expected to explain those symptoms. We present the first complete method solving signature-based abduction for observations expressed in the expressive description logic ALC, which can include TBox and ABox axioms, thereby solving the knowledge base abduction problem. The method is guaranteed to compute a finite and complete set of hypotheses, and is evaluated on a set of realistic knowledge bases.
ABox Abduction via Forgetting in ALC (Long Version)
Nov 13, 2018Warren Del-Pinto, Renate A. Schmidt
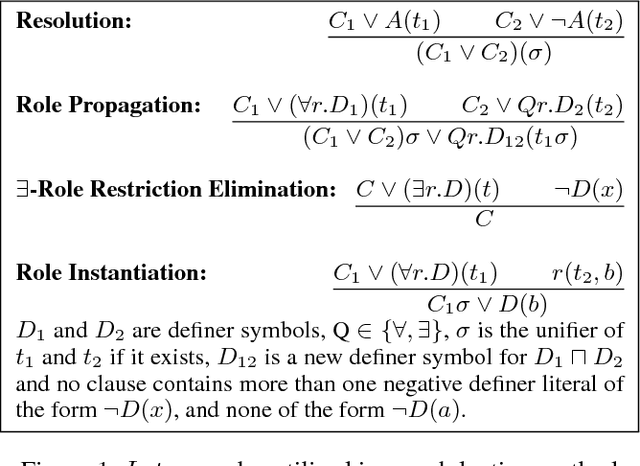

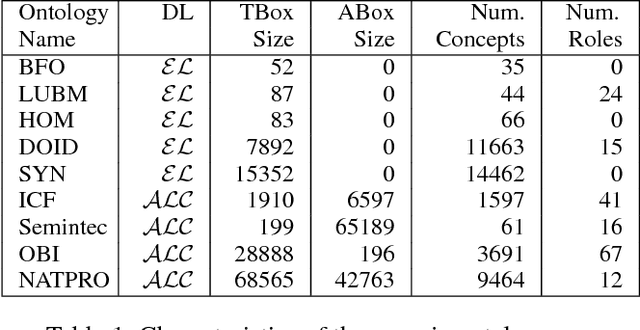
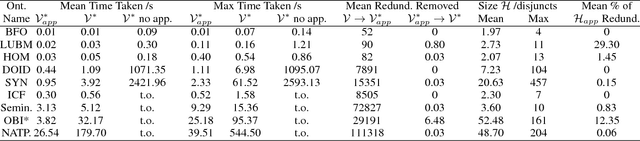
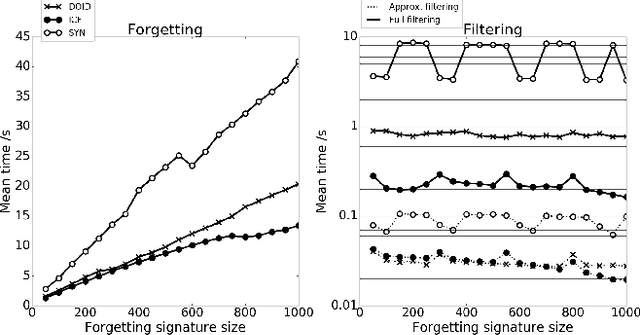
Abductive reasoning generates explanatory hypotheses for new observations using prior knowledge. This paper investigates the use of forgetting, also known as uniform interpolation, to perform ABox abduction in description logic (ALC) ontologies. Non-abducibles are specified by a forgetting signature which can contain concept, but not role, symbols. The resulting hypotheses are semantically minimal and each consist of a set of disjuncts. These disjuncts are each independent explanations, and are not redundant with respect to the background ontology or the other disjuncts, representing a form of hypothesis space. The observations and hypotheses handled by the method can contain both atomic or complex ALC concepts, excluding role assertions, and are not restricted to Horn clauses. Two approaches to redundancy elimination are explored for practical use: full and approximate. Using a prototype implementation, experiments were performed over a corpus of real world ontologies to investigate the practicality of both approaches across several settings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge