A Selective Review on Statistical Methods for Massive Data Computation: Distributed Computing, Subsampling, and Minibatch Techniques
Mar 17, 2024Xuetong Li, Yuan Gao, Hong Chang, Danyang Huang, Yingying Ma, Rui Pan, Haobo Qi, Feifei Wang, Shuyuan Wu, Ke Xu, Jing Zhou, Xuening Zhu, Yingqiu Zhu, Hansheng Wang
This paper presents a selective review of statistical computation methods for massive data analysis. A huge amount of statistical methods for massive data computation have been rapidly developed in the past decades. In this work, we focus on three categories of statistical computation methods: (1) distributed computing, (2) subsampling methods, and (3) minibatch gradient techniques. The first class of literature is about distributed computing and focuses on the situation, where the dataset size is too huge to be comfortably handled by one single computer. In this case, a distributed computation system with multiple computers has to be utilized. The second class of literature is about subsampling methods and concerns about the situation, where the sample size of dataset is small enough to be placed on one single computer but too large to be easily processed by its memory as a whole. The last class of literature studies those minibatch gradient related optimization techniques, which have been extensively used for optimizing various deep learning models.
Improved Naive Bayes with Mislabeled Data
Apr 13, 2023Qianhan Zeng, Yingqiu Zhu, Xuening Zhu, Feifei Wang, Weichen Zhao, Shuning Sun, Meng Su, Hansheng Wang
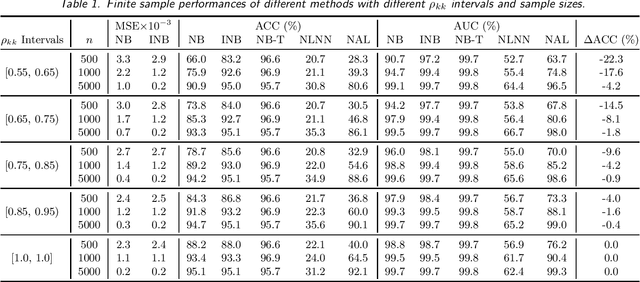



Labeling mistakes are frequently encountered in real-world applications. If not treated well, the labeling mistakes can deteriorate the classification performances of a model seriously. To address this issue, we propose an improved Naive Bayes method for text classification. It is analytically simple and free of subjective judgements on the correct and incorrect labels. By specifying the generating mechanism of incorrect labels, we optimize the corresponding log-likelihood function iteratively by using an EM algorithm. Our simulation and experiment results show that the improved Naive Bayes method greatly improves the performances of the Naive Bayes method with mislabeled data.
CoGANPPIS: Coevolution-enhanced Global Attention Neural Network for Protein-Protein Interaction Site Prediction
Apr 03, 2023Jiaxing Guo, Xuening Zhu, Zixin Hu, Xiaoxi Hu

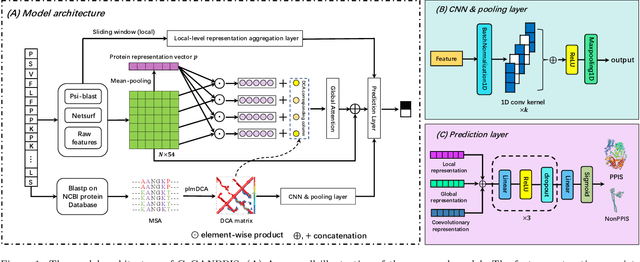

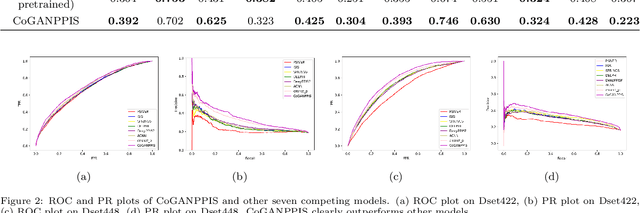
Protein-protein interactions are essential in biochemical processes. Accurate prediction of the protein-protein interaction sites (PPIs) deepens our understanding of biological mechanism and is crucial for new drug design. However, conventional experimental methods for PPIs prediction are costly and time-consuming so that many computational approaches, especially ML-based methods, have been developed recently. Although these approaches have achieved gratifying results, there are still two limitations: (1) Most models have excavated some useful input features, but failed to take coevolutionary features into account, which could provide clues for inter-residue relationships; (2) The attention-based models only allocate attention weights for neighboring residues, instead of doing it globally, neglecting that some residues being far away from the target residues might also matter. We propose a coevolution-enhanced global attention neural network, a sequence-based deep learning model for PPIs prediction, called CoGANPPIS. It utilizes three layers in parallel for feature extraction: (1) Local-level representation aggregation layer, which aggregates the neighboring residues' features; (2) Global-level representation learning layer, which employs a novel coevolution-enhanced global attention mechanism to allocate attention weights to all the residues on the same protein sequences; (3) Coevolutionary information learning layer, which applies CNN & pooling to coevolutionary information to obtain the coevolutionary profile representation. Then, the three outputs are concatenated and passed into several fully connected layers for the final prediction. Application on two benchmark datasets demonstrated a state-of-the-art performance of our model. The source code is publicly available at https://github.com/Slam1423/CoGANPPIS_source_code.
Least Squares Approximation for a Distributed System
Aug 14, 2019Xuening Zhu, Feng Li, Hansheng Wang
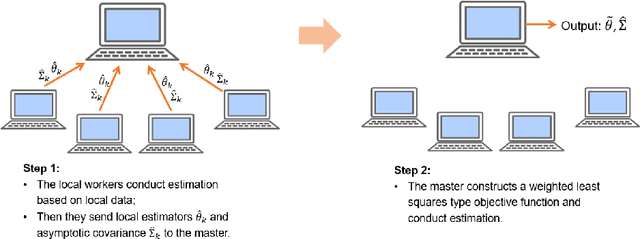
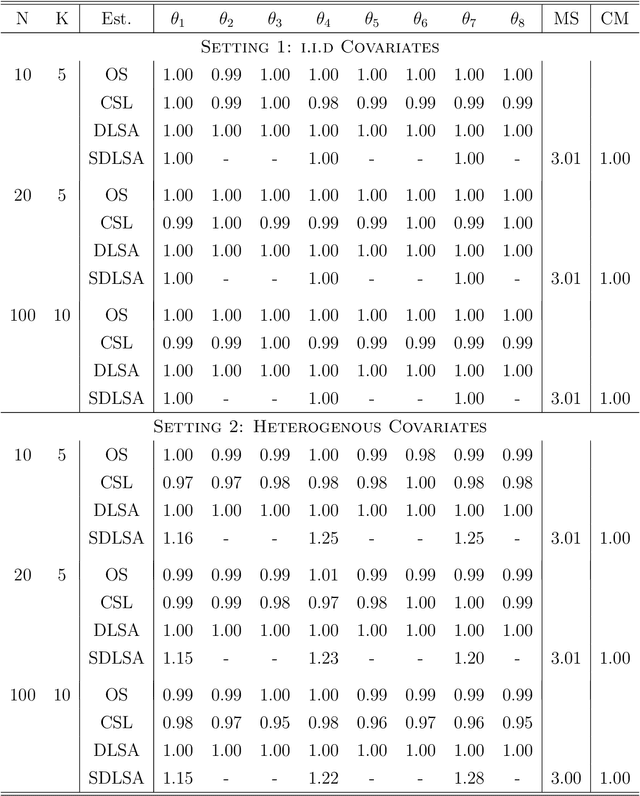
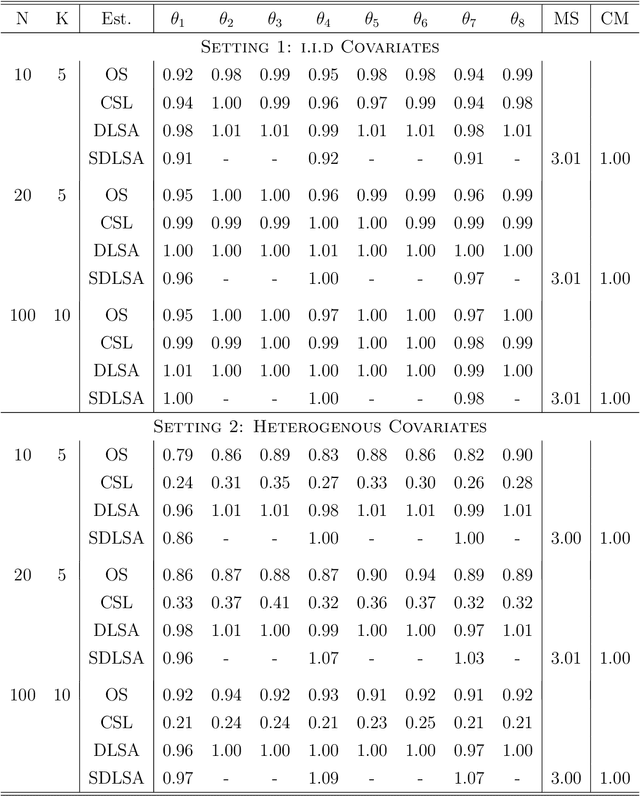
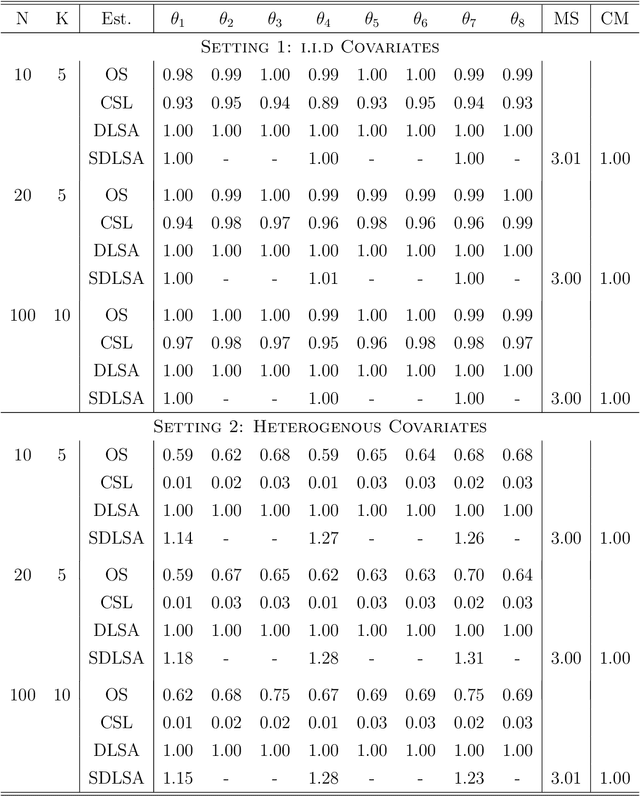
In this work we develop a distributed least squares approximation (DLSA) method, which is able to solve a large family of regression problems (e.g., linear regression, logistic regression, Cox's model) on a distributed system. By approximating the local objective function using a local quadratic form, we are able to obtain a combined estimator by taking a weighted average of local estimators. The resulting estimator is proved to be statistically as efficient as the global estimator. In the meanwhile it requires only one round of communication. We further conduct the shrinkage estimation based on the DLSA estimation by using an adaptive Lasso approach. The solution can be easily obtained by using the LARS algorithm on the master node. It is theoretically shown that the resulting estimator enjoys the oracle property and is selection consistent by using a newly designed distributed Bayesian Information Criterion (DBIC). The finite sample performance as well as the computational efficiency are further illustrated by extensive numerical study and an airline dataset. The airline dataset is 52GB in memory size. The entire methodology has been implemented by Python for a de-facto standard Spark system. By using the proposed DLSA algorithm on the Spark system, it takes 26 minutes to obtain a logistic regression estimator whereas a full likelihood algorithm takes 15 hours to reaches an inferior result.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge