Iterative Subsampling in Solution Path Clustering of Noisy Big Data
Jul 16, 2015Yuliya Marchetti, Qing Zhou
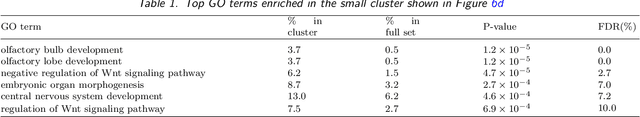
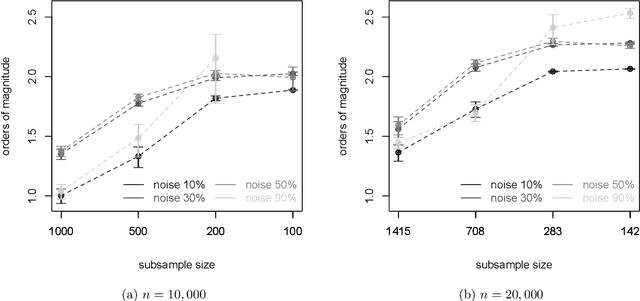

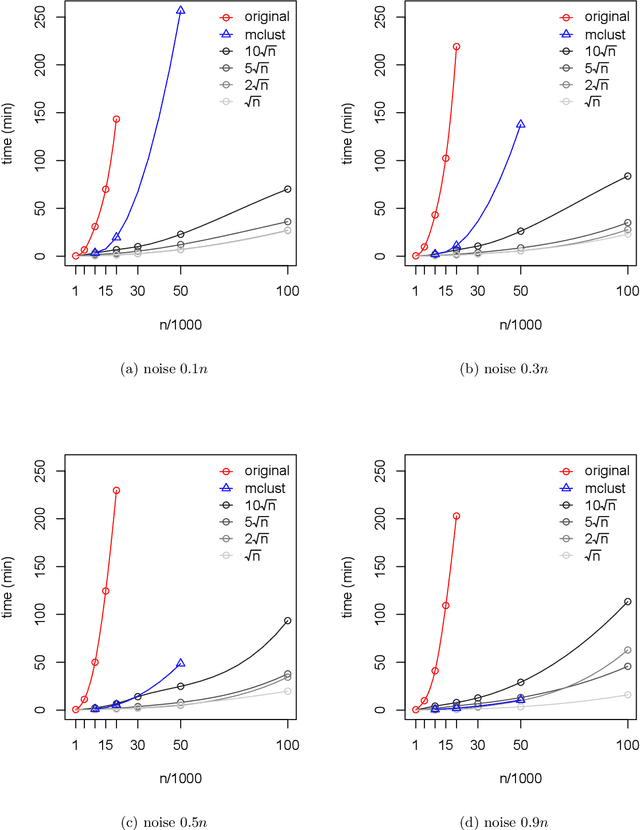
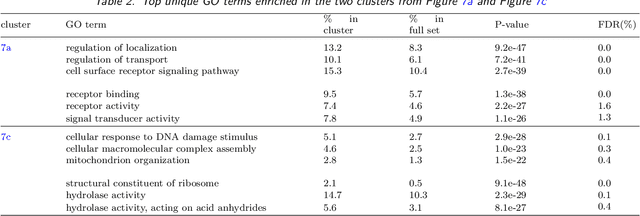
We develop an iterative subsampling approach to improve the computational efficiency of our previous work on solution path clustering (SPC). The SPC method achieves clustering by concave regularization on the pairwise distances between cluster centers. This clustering method has the important capability to recognize noise and to provide a short path of clustering solutions; however, it is not sufficiently fast for big datasets. Thus, we propose a method that iterates between clustering a small subsample of the full data and sequentially assigning the other data points to attain orders of magnitude of computational savings. The new method preserves the ability to isolate noise, includes a solution selection mechanism that ultimately provides one clustering solution with an estimated number of clusters, and is shown to be able to extract small tight clusters from noisy data. The method's relatively minor losses in accuracy are demonstrated through simulation studies, and its ability to handle large datasets is illustrated through applications to gene expression datasets. An R package, SPClustering, for the SPC method with iterative subsampling is available at http://www.stat.ucla.edu/~zhou/Software.html.
* 17 pages, 7 figures
Solution Path Clustering with Adaptive Concave Penalty
Apr 24, 2014Yuliya Marchetti, Qing Zhou
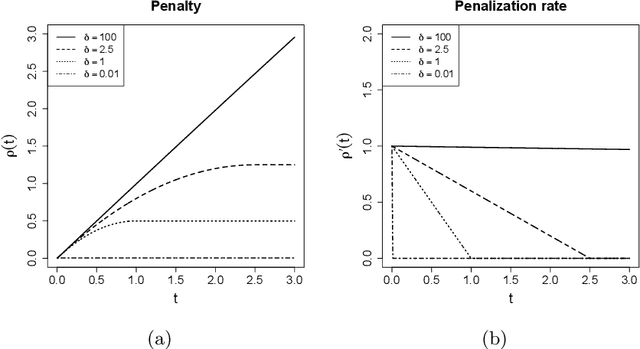
Fast accumulation of large amounts of complex data has created a need for more sophisticated statistical methodologies to discover interesting patterns and better extract information from these data. The large scale of the data often results in challenging high-dimensional estimation problems where only a minority of the data shows specific grouping patterns. To address these emerging challenges, we develop a new clustering methodology that introduces the idea of a regularization path into unsupervised learning. A regularization path for a clustering problem is created by varying the degree of sparsity constraint that is imposed on the differences between objects via the minimax concave penalty with adaptive tuning parameters. Instead of providing a single solution represented by a cluster assignment for each object, the method produces a short sequence of solutions that determines not only the cluster assignment but also a corresponding number of clusters for each solution. The optimization of the penalized loss function is carried out through an MM algorithm with block coordinate descent. The advantages of this clustering algorithm compared to other existing methods are as follows: it does not require the input of the number of clusters; it is capable of simultaneously separating irrelevant or noisy observations that show no grouping pattern, which can greatly improve data interpretation; it is a general methodology that can be applied to many clustering problems. We test this method on various simulated datasets and on gene expression data, where it shows better or competitive performance compared against several clustering methods.
* 36 pages
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge