DeepVar: An End-to-End Deep Learning Approach for Genomic Variant Recognition in Biomedical Literature
Jun 05, 2020Chaoran Cheng, Fei Tan, Zhi Wei
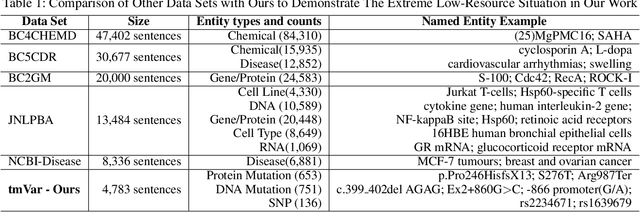
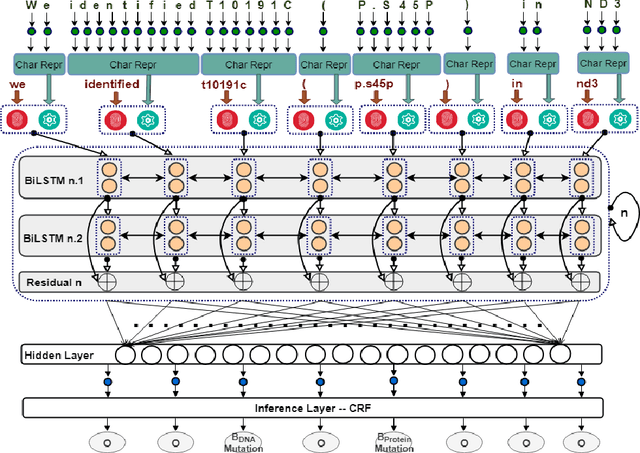

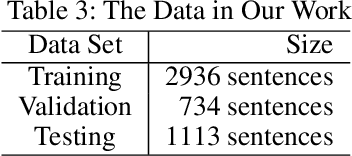
We consider the problem of Named Entity Recognition (NER) on biomedical scientific literature, and more specifically the genomic variants recognition in this work. Significant success has been achieved for NER on canonical tasks in recent years where large data sets are generally available. However, it remains a challenging problem on many domain-specific areas, especially the domains where only small gold annotations can be obtained. In addition, genomic variant entities exhibit diverse linguistic heterogeneity, differing much from those that have been characterized in existing canonical NER tasks. The state-of-the-art machine learning approaches in such tasks heavily rely on arduous feature engineering to characterize those unique patterns. In this work, we present the first successful end-to-end deep learning approach to bridge the gap between generic NER algorithms and low-resource applications through genomic variants recognition. Our proposed model can result in promising performance without any hand-crafted features or post-processing rules. Our extensive experiments and results may shed light on other similar low-resource NER applications.
A Blended Deep Learning Approach for Predicting User Intended Actions
Oct 11, 2018Fei Tan, Zhi Wei, Jun He, Xiang Wu, Bo Peng, Haoran Liu, Zhenyu Yan
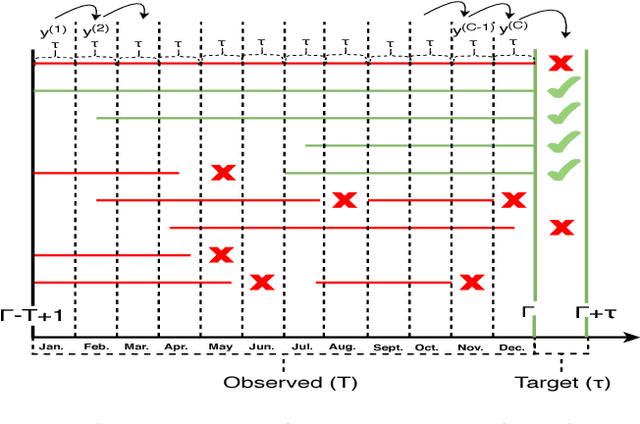
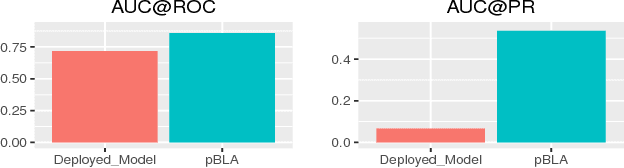
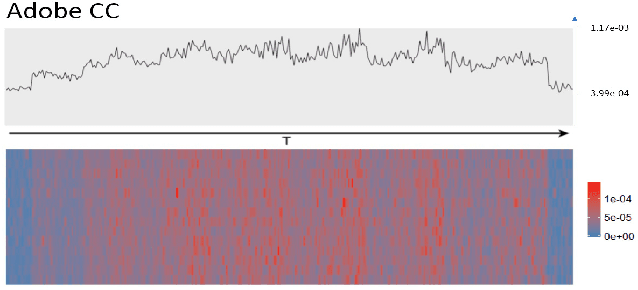
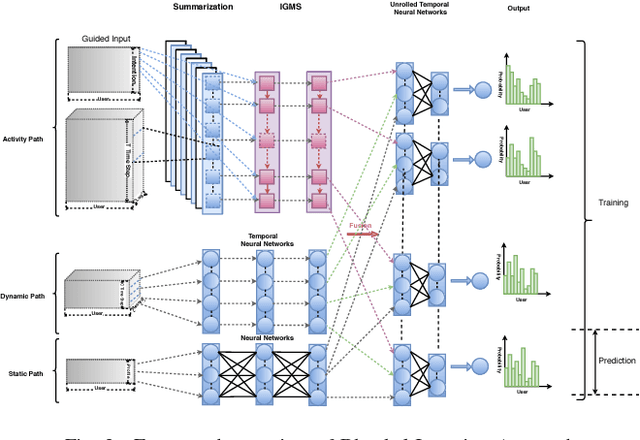
User intended actions are widely seen in many areas. Forecasting these actions and taking proactive measures to optimize business outcome is a crucial step towards sustaining the steady business growth. In this work, we focus on pre- dicting attrition, which is one of typical user intended actions. Conventional attrition predictive modeling strategies suffer a few inherent drawbacks. To overcome these limitations, we propose a novel end-to-end learning scheme to keep track of the evolution of attrition patterns for the predictive modeling. It integrates user activity logs, dynamic and static user profiles based on multi-path learning. It exploits historical user records by establishing a decaying multi-snapshot technique. And finally it employs the precedent user intentions via guiding them to the subsequent learning procedure. As a result, it addresses all disadvantages of conventional methods. We evaluate our methodology on two public data repositories and one private user usage dataset provided by Adobe Creative Cloud. The extensive experiments demonstrate that it can offer the appealing performance in comparison with several existing approaches as rated by different popular metrics. Furthermore, we introduce an advanced interpretation and visualization strategy to effectively characterize the periodicity of user activity logs. It can help to pinpoint important factors that are critical to user attrition and retention and thus suggests actionable improvement targets for business practice. Our work will provide useful insights into the prediction and elucidation of other user intended actions as well.
A Noise-Filtering Approach for Cancer Drug Sensitivity Prediction
Dec 05, 2016Turki Turki, Zhi Wei
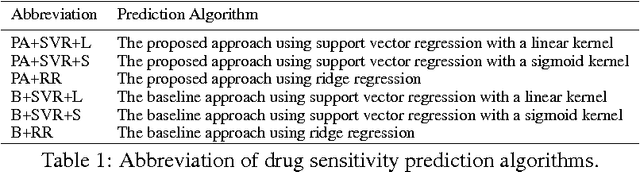
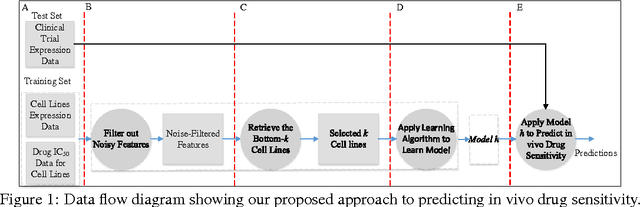

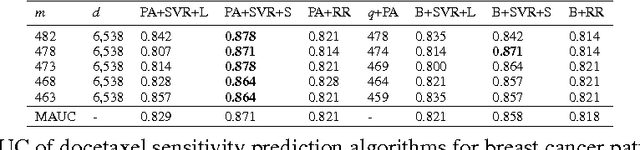
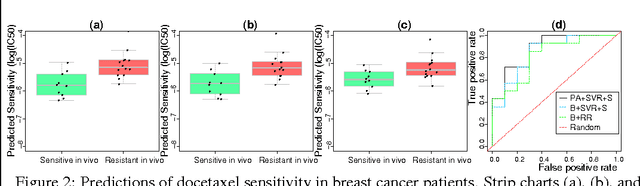
Accurately predicting drug responses to cancer is an important problem hindering oncologists' efforts to find the most effective drugs to treat cancer, which is a core goal in precision medicine. The scientific community has focused on improving this prediction based on genomic, epigenomic, and proteomic datasets measured in human cancer cell lines. Real-world cancer cell lines contain noise, which degrades the performance of machine learning algorithms. This problem is rarely addressed in the existing approaches. In this paper, we present a noise-filtering approach that integrates techniques from numerical linear algebra and information retrieval targeted at filtering out noisy cancer cell lines. By filtering out noisy cancer cell lines, we can train machine learning algorithms on better quality cancer cell lines. We evaluate the performance of our approach and compare it with an existing approach using the Area Under the ROC Curve (AUC) on clinical trial data. The experimental results show that our proposed approach is stable and also yields the highest AUC at a statistically significant level.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge