Causal Imputation for Counterfactual SCMs: Bridging Graphs and Latent Factor Models
Feb 22, 2024Alvaro Ribot, Chandler Squires, Caroline Uhler
We consider the task of causal imputation, where we aim to predict the outcomes of some set of actions across a wide range of possible contexts. As a running example, we consider predicting how different drugs affect cells from different cell types. We study the index-only setting, where the actions and contexts are categorical variables with a finite number of possible values. Even in this simple setting, a practical challenge arises, since often only a small subset of possible action-context pairs have been studied. Thus, models must extrapolate to novel action-context pairs, which can be framed as a form of matrix completion with rows indexed by actions, columns indexed by contexts, and matrix entries corresponding to outcomes. We introduce a novel SCM-based model class, where the outcome is expressed as a counterfactual, actions are expressed as interventions on an instrumental variable, and contexts are defined based on the initial state of the system. We show that, under a linearity assumption, this setup induces a latent factor model over the matrix of outcomes, with an additional fixed effect term. To perform causal prediction based on this model class, we introduce simple extension to the Synthetic Interventions estimator (Agarwal et al., 2020). We evaluate several matrix completion approaches on the PRISM drug repurposing dataset, showing that our method outperforms all other considered matrix completion approaches.
Identifiability Guarantees for Causal Disentanglement from Soft Interventions
Jul 13, 2023Jiaqi Zhang, Chandler Squires, Kristjan Greenewald, Akash Srivastava, Karthikeyan Shanmugam, Caroline Uhler
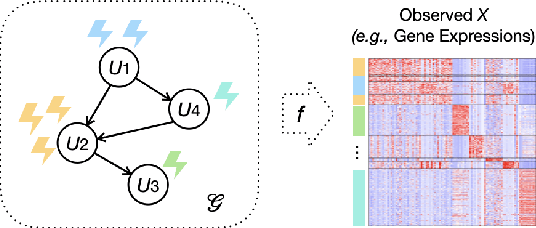
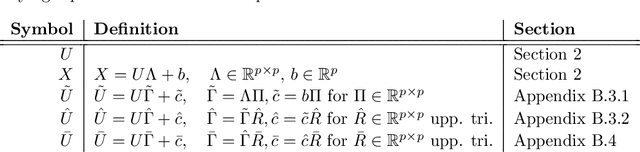
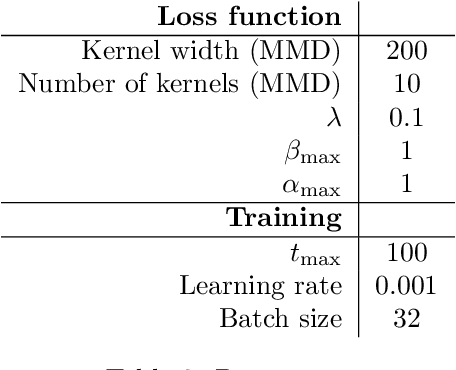
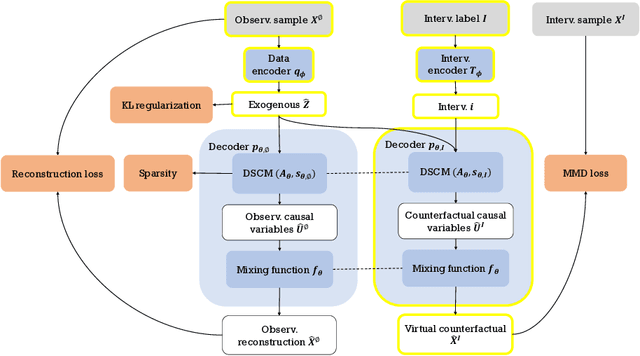
Causal disentanglement aims to uncover a representation of data using latent variables that are interrelated through a causal model. Such a representation is identifiable if the latent model that explains the data is unique. In this paper, we focus on the scenario where unpaired observational and interventional data are available, with each intervention changing the mechanism of a latent variable. When the causal variables are fully observed, statistically consistent algorithms have been developed to identify the causal model under faithfulness assumptions. We here show that identifiability can still be achieved with unobserved causal variables, given a generalized notion of faithfulness. Our results guarantee that we can recover the latent causal model up to an equivalence class and predict the effect of unseen combinations of interventions, in the limit of infinite data. We implement our causal disentanglement framework by developing an autoencoding variational Bayes algorithm and apply it to the problem of predicting combinatorial perturbation effects in genomics.
Unpaired Multi-Domain Causal Representation Learning
Feb 02, 2023Nils Sturma, Chandler Squires, Mathias Drton, Caroline Uhler
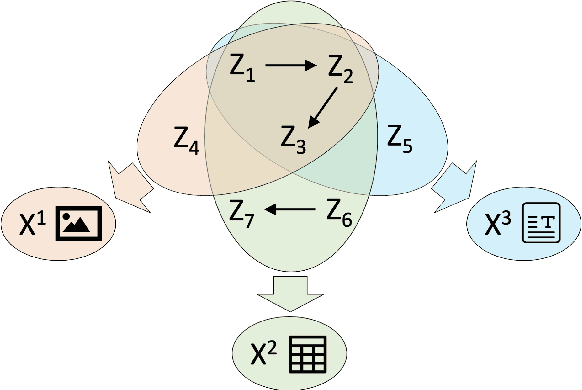
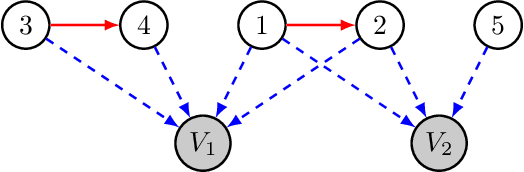
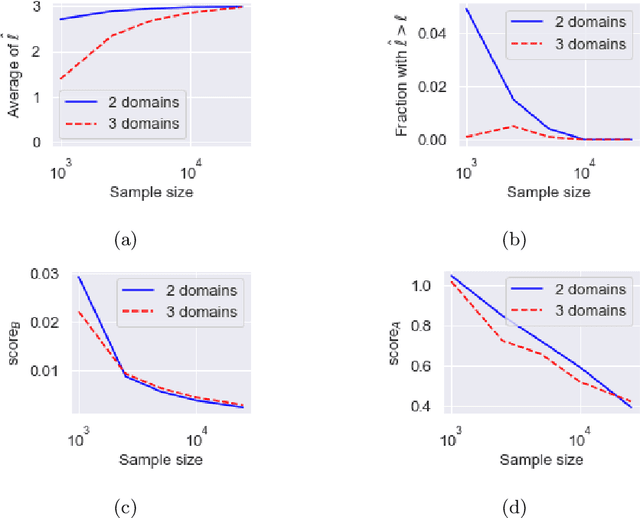
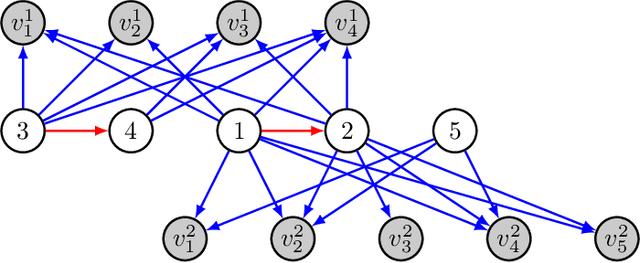
The goal of causal representation learning is to find a representation of data that consists of causally related latent variables. We consider a setup where one has access to data from multiple domains that potentially share a causal representation. Crucially, observations in different domains are assumed to be unpaired, that is, we only observe the marginal distribution in each domain but not their joint distribution. In this paper, we give sufficient conditions for identifiability of the joint distribution and the shared causal graph in a linear setup. Identifiability holds if we can uniquely recover the joint distribution and the shared causal representation from the marginal distributions in each domain. We transform our identifiability results into a practical method to recover the shared latent causal graph. Moreover, we study how multiple domains reduce errors in falsely detecting shared causal variables in the finite data setting.
Linear Causal Disentanglement via Interventions
Nov 29, 2022Anna Seigal, Chandler Squires, Caroline Uhler
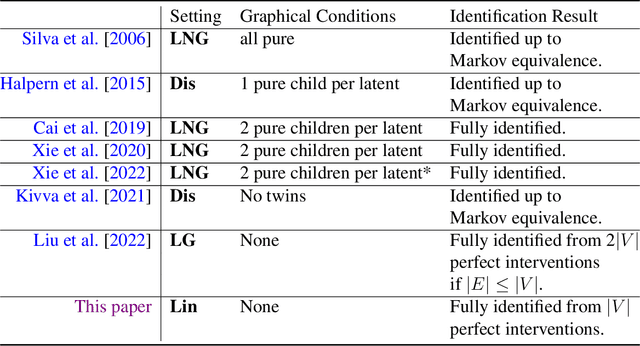
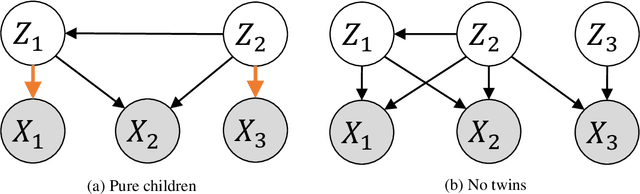
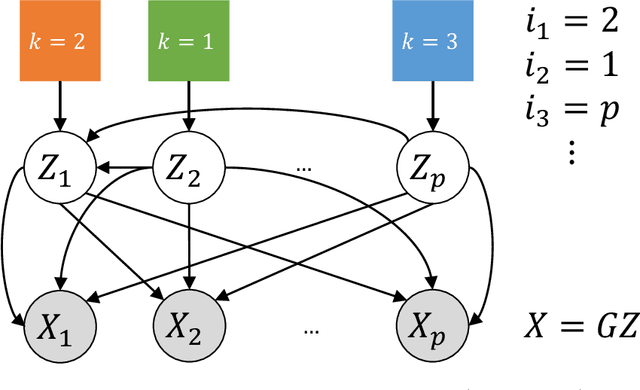
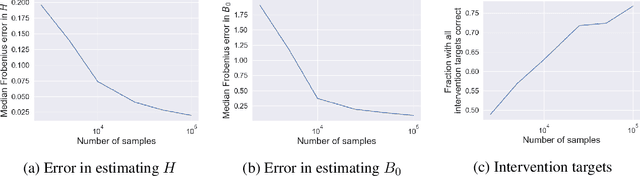
Causal disentanglement seeks a representation of data involving latent variables that relate to one another via a causal model. A representation is identifiable if both the latent model and the transformation from latent to observed variables are unique. In this paper, we study observed variables that are a linear transformation of a linear latent causal model. Data from interventions are necessary for identifiability: if one latent variable is missing an intervention, we show that there exist distinct models that cannot be distinguished. Conversely, we show that a single intervention on each latent variable is sufficient for identifiability. Our proof uses a generalization of the RQ decomposition of a matrix that replaces the usual orthogonal and upper triangular conditions with analogues depending on a partial order on the rows of the matrix, with partial order determined by a latent causal model. We corroborate our theoretical results with a method for causal disentanglement that accurately recovers a latent causal model.
Active Learning for Optimal Intervention Design in Causal Models
Sep 10, 2022Jiaqi Zhang, Louis Cammarata, Chandler Squires, Themistoklis P. Sapsis, Caroline Uhler
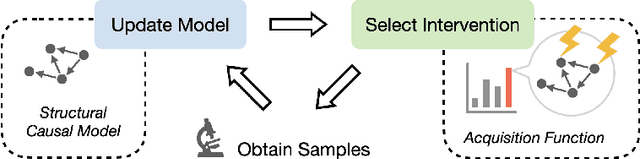
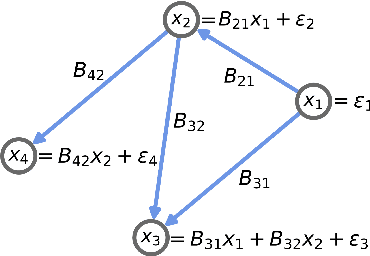
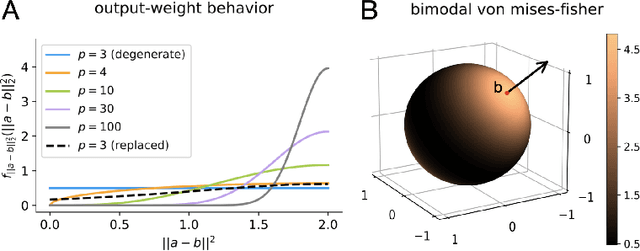
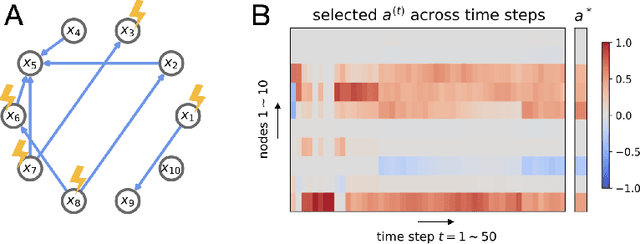
An important problem across disciplines is the discovery of interventions that produce a desired outcome. When the space of possible interventions is large, making an exhaustive search infeasible, experimental design strategies are needed. In this context, encoding the causal relationships between the variables, and thus the effect of interventions on the system, is critical in order to identify desirable interventions efficiently. We develop an iterative causal method to identify optimal interventions, as measured by the discrepancy between the post-interventional mean of the distribution and a desired target mean. We formulate an active learning strategy that uses the samples obtained so far from different interventions to update the belief about the underlying causal model, as well as to identify samples that are most informative about optimal interventions and thus should be acquired in the next batch. The approach employs a Bayesian update for the causal model and prioritizes interventions using a carefully designed, causally informed acquisition function. This acquisition function is evaluated in closed form, allowing for efficient optimization. The resulting algorithms are theoretically grounded with information-theoretic bounds and provable consistency results. We illustrate the method on both synthetic data and real-world biological data, namely gene expression data from Perturb-CITE-seq experiments, to identify optimal perturbations that induce a specific cell state transition; the proposed causal approach is observed to achieve better sample efficiency compared to several baselines. In both cases we observe that the causally informed acquisition function notably outperforms existing criteria allowing for optimal intervention design with significantly less experiments.
Causal Structure Learning: a Combinatorial Perspective
Jun 02, 2022Chandler Squires, Caroline Uhler
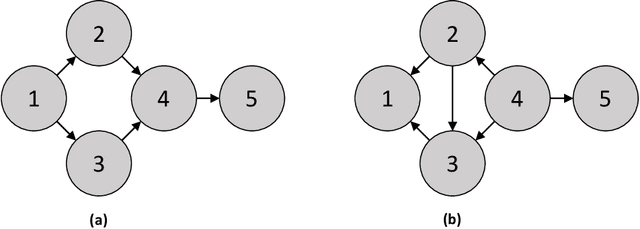

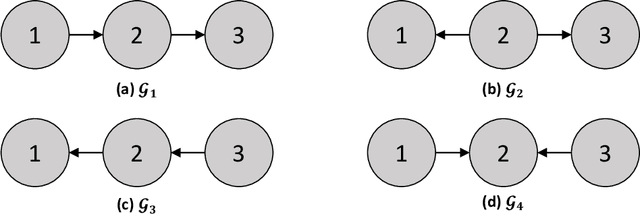
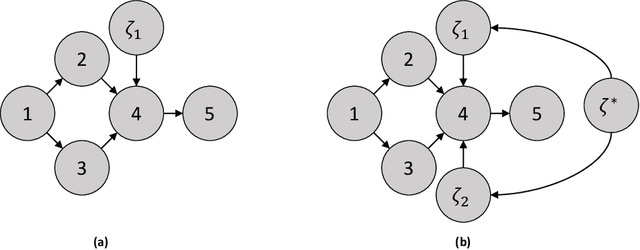
In this review, we discuss approaches for learning causal structure from data, also called causal discovery. In particular, we focus on approaches for learning directed acyclic graphs (DAGs) and various generalizations which allow for some variables to be unobserved in the available data. We devote special attention to two fundamental combinatorial aspects of causal structure learning. First, we discuss the structure of the search space over causal graphs. Second, we discuss the structure of equivalence classes over causal graphs, i.e., sets of graphs which represent what can be learned from observational data alone, and how these equivalence classes can be refined by adding interventional data.
Matching a Desired Causal State via Shift Interventions
Jul 05, 2021Jiaqi Zhang, Chandler Squires, Caroline Uhler
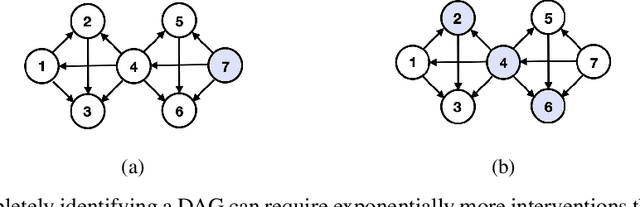
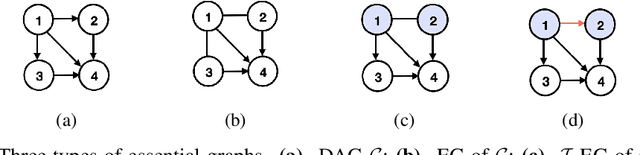
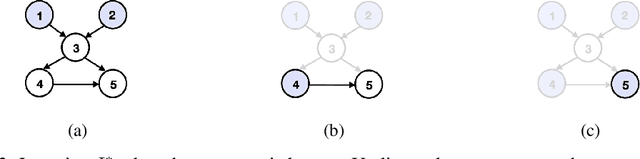
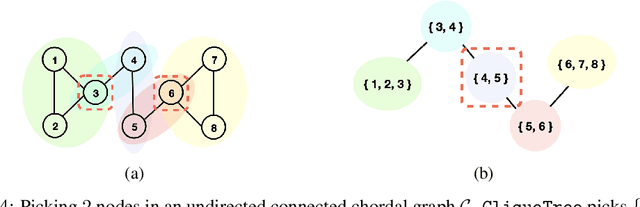
Transforming a causal system from a given initial state to a desired target state is an important task permeating multiple fields including control theory, biology, and materials science. In causal models, such transformations can be achieved by performing a set of interventions. In this paper, we consider the problem of identifying a shift intervention that matches the desired mean of a system through active learning. We define the Markov equivalence class that is identifiable from shift interventions and propose two active learning strategies that are guaranteed to exactly match a desired mean. We then derive a worst-case lower bound for the number of interventions required and show that these strategies are optimal for certain classes of graphs. In particular, we show that our strategies may require exponentially fewer interventions than the previously considered approaches, which optimize for structure learning in the underlying causal graph. In line with our theoretical results, we also demonstrate experimentally that our proposed active learning strategies require fewer interventions compared to several baselines.
Efficient Permutation Discovery in Causal DAGs
Nov 06, 2020Chandler Squires, Joshua Amaniampong, Caroline Uhler
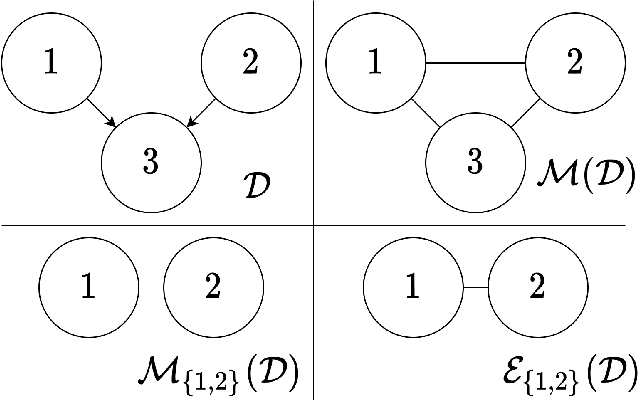
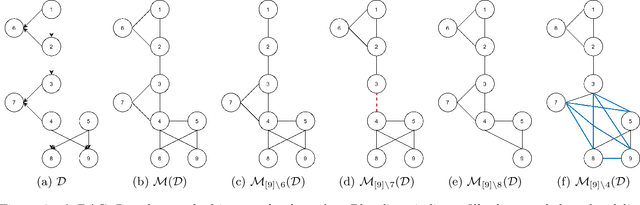
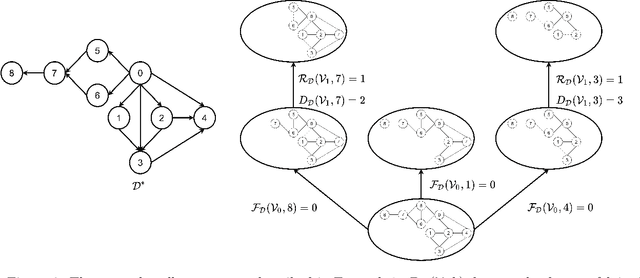
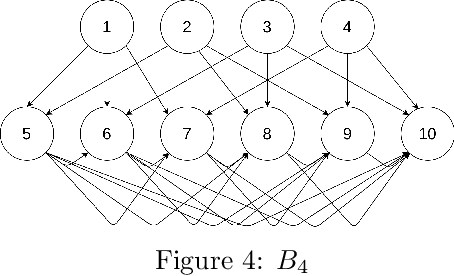
The problem of learning a directed acyclic graph (DAG) up to Markov equivalence is equivalent to the problem of finding a permutation of the variables that induces the sparsest graph. Without additional assumptions, this task is known to be NP-hard. Building on the minimum degree algorithm for sparse Cholesky decomposition, but utilizing DAG-specific problem structure, we introduce an efficient algorithm for finding such sparse permutations. We show that on jointly Gaussian distributions, our method with depth $w$ runs in $O(p^{w+3})$ time. We compare our method with $w = 1$ to algorithms for finding sparse elimination orderings of undirected graphs, and show that taking advantage of DAG-specific problem structure leads to a significant improvement in the discovered permutation. We also compare our algorithm to provably consistent causal structure learning algorithms, such as the PC algorithm, GES, and GSP, and show that our method achieves comparable performance with a shorter runtime. Thus, our method can be used on its own for causal structure discovery. Finally, we show that there exist dense graphs on which our method achieves almost perfect performance, so that unlike most existing causal structure learning algorithms, the situations in which our algorithm achieves both good performance and good runtime are not limited to sparse graphs.
Active Structure Learning of Causal DAGs via Directed Clique Tree
Nov 01, 2020Chandler Squires, Sara Magliacane, Kristjan Greenewald, Dmitriy Katz, Murat Kocaoglu, Karthikeyan Shanmugam
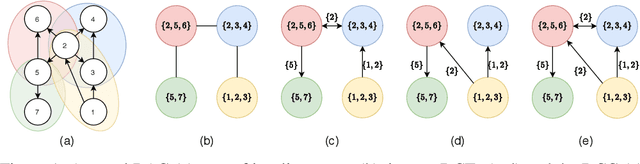
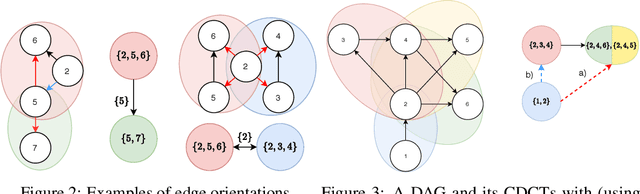
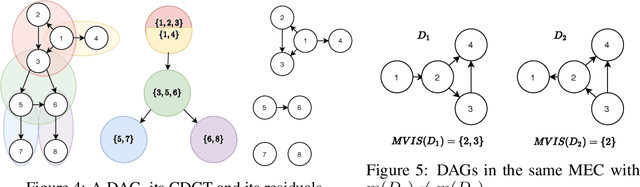
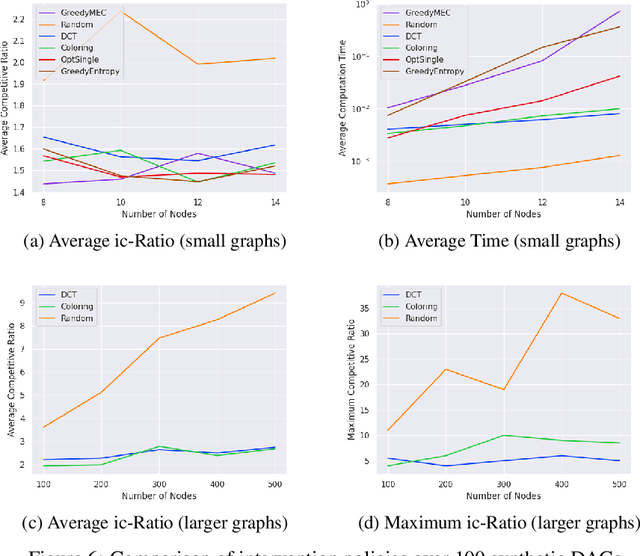
A growing body of work has begun to study intervention design for efficient structure learning of causal directed acyclic graphs (DAGs). A typical setting is a causally sufficient setting, i.e. a system with no latent confounders, selection bias, or feedback, when the essential graph of the observational equivalence class (EC) is given as an input and interventions are assumed to be noiseless. Most existing works focus on worst-case or average-case lower bounds for the number of interventions required to orient a DAG. These worst-case lower bounds only establish that the largest clique in the essential graph could make it difficult to learn the true DAG. In this work, we develop a universal lower bound for single-node interventions that establishes that the largest clique is always a fundamental impediment to structure learning. Specifically, we present a decomposition of a DAG into independently orientable components through directed clique trees and use it to prove that the number of single-node interventions necessary to orient any DAG in an EC is at least the sum of half the size of the largest cliques in each chain component of the essential graph. Moreover, we present a two-phase intervention design algorithm that, under certain conditions on the chordal skeleton, matches the optimal number of interventions up to a multiplicative logarithmic factor in the number of maximal cliques. We show via synthetic experiments that our algorithm can scale to much larger graphs than most of the related work and achieves better worst-case performance than other scalable approaches. A code base to recreate these results can be found at https://github.com/csquires/dct-policy
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge