Automatic classification of prostate MR series type using image content and metadata
Apr 16, 2024Deepa Krishnaswamy, Bálint Kovács, Stefan Denner, Steve Pieper, David Clunie, Christopher P. Bridge, Tina Kapur, Klaus H. Maier-Hein, Andrey Fedorov
With the wealth of medical image data, efficient curation is essential. Assigning the sequence type to magnetic resonance images is necessary for scientific studies and artificial intelligence-based analysis. However, incomplete or missing metadata prevents effective automation. We therefore propose a deep-learning method for classification of prostate cancer scanning sequences based on a combination of image data and DICOM metadata. We demonstrate superior results compared to metadata or image data alone, and make our code publicly available at https://github.com/deepakri201/DICOMScanClassification.
Enrichment of the NLST and NSCLC-Radiomics computed tomography collections with AI-derived annotations
May 31, 2023Deepa Krishnaswamy, Dennis Bontempi, Vamsi Thiriveedhi, Davide Punzo, David Clunie, Christopher P Bridge, Hugo JWL Aerts, Ron Kikinis, Andrey Fedorov
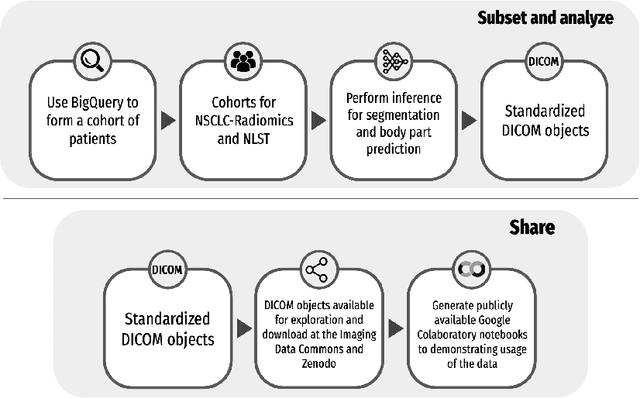
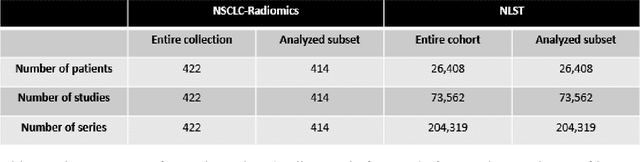
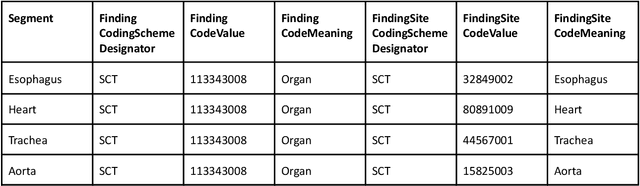

Public imaging datasets are critical for the development and evaluation of automated tools in cancer imaging. Unfortunately, many do not include annotations or image-derived features, complicating their downstream analysis. Artificial intelligence-based annotation tools have been shown to achieve acceptable performance and thus can be used to automatically annotate large datasets. As part of the effort to enrich public data available within NCI Imaging Data Commons (IDC), here we introduce AI-generated annotations for two collections of computed tomography images of the chest, NSCLC-Radiomics, and the National Lung Screening Trial. Using publicly available AI algorithms we derived volumetric annotations of thoracic organs at risk, their corresponding radiomics features, and slice-level annotations of anatomical landmarks and regions. The resulting annotations are publicly available within IDC, where the DICOM format is used to harmonize the data and achieve FAIR principles. The annotations are accompanied by cloud-enabled notebooks demonstrating their use. This study reinforces the need for large, publicly accessible curated datasets and demonstrates how AI can be used to aid in cancer imaging.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge