Self-Supervised Learning for Interventional Image Analytics: Towards Robust Device Trackers
May 02, 2024Saahil Islam, Venkatesh N. Murthy, Dominik Neumann, Badhan Kumar Das, Puneet Sharma, Andreas Maier, Dorin Comaniciu, Florin C. Ghesu
An accurate detection and tracking of devices such as guiding catheters in live X-ray image acquisitions is an essential prerequisite for endovascular cardiac interventions. This information is leveraged for procedural guidance, e.g., directing stent placements. To ensure procedural safety and efficacy, there is a need for high robustness no failures during tracking. To achieve that, one needs to efficiently tackle challenges, such as: device obscuration by contrast agent or other external devices or wires, changes in field-of-view or acquisition angle, as well as the continuous movement due to cardiac and respiratory motion. To overcome the aforementioned challenges, we propose a novel approach to learn spatio-temporal features from a very large data cohort of over 16 million interventional X-ray frames using self-supervision for image sequence data. Our approach is based on a masked image modeling technique that leverages frame interpolation based reconstruction to learn fine inter-frame temporal correspondences. The features encoded in the resulting model are fine-tuned downstream. Our approach achieves state-of-the-art performance and in particular robustness compared to ultra optimized reference solutions (that use multi-stage feature fusion, multi-task and flow regularization). The experiments show that our method achieves 66.31% reduction in maximum tracking error against reference solutions (23.20% when flow regularization is used); achieving a success score of 97.95% at a 3x faster inference speed of 42 frames-per-second (on GPU). The results encourage the use of our approach in various other tasks within interventional image analytics that require effective understanding of spatio-temporal semantics.
General-Purpose vs. Domain-Adapted Large Language Models for Extraction of Data from Thoracic Radiology Reports
Dec 01, 2023Ali H. Dhanaliwala, Rikhiya Ghosh, Sanjeev Kumar Karn, Poikavila Ullaskrishnan, Oladimeji Farri, Dorin Comaniciu, Charles E. Kahn
Radiologists produce unstructured data that could be valuable for clinical care when consumed by information systems. However, variability in style limits usage. Study compares performance of system using domain-adapted language model (RadLing) and general-purpose large language model (GPT-4) in extracting common data elements (CDE) from thoracic radiology reports. Three radiologists annotated a retrospective dataset of 1300 thoracic reports (900 training, 400 test) and mapped to 21 pre-selected relevant CDEs. RadLing was used to generate embeddings for sentences and identify CDEs using cosine-similarity, which were mapped to values using light-weight mapper. GPT-4 system used OpenAI's general-purpose embeddings to identify relevant CDEs and used GPT-4 to map to values. The output CDE:value pairs were compared to the reference standard; an identical match was considered true positive. Precision (positive predictive value) was 96% (2700/2824) for RadLing and 99% (2034/2047) for GPT-4. Recall (sensitivity) was 94% (2700/2876) for RadLing and 70% (2034/2887) for GPT-4; the difference was statistically significant (P<.001). RadLing's domain-adapted embeddings were more sensitive in CDE identification (95% vs 71%) and its light-weight mapper had comparable precision in value assignment (95.4% vs 95.0%). RadLing system exhibited higher performance than GPT-4 system in extracting CDEs from radiology reports. RadLing system's domain-adapted embeddings outperform general-purpose embeddings from OpenAI in CDE identification and its light-weight value mapper achieves comparable precision to large GPT-4. RadLing system offers operational advantages including local deployment and reduced runtime costs. Domain-adapted RadLing system surpasses GPT-4 system in extracting common data elements from radiology reports, while providing benefits of local deployment and lower costs.
ConTrack: Contextual Transformer for Device Tracking in X-ray
Jul 14, 2023Marc Demoustier, Yue Zhang, Venkatesh Narasimha Murthy, Florin C. Ghesu, Dorin Comaniciu
Device tracking is an important prerequisite for guidance during endovascular procedures. Especially during cardiac interventions, detection and tracking of guiding the catheter tip in 2D fluoroscopic images is important for applications such as mapping vessels from angiography (high dose with contrast) to fluoroscopy (low dose without contrast). Tracking the catheter tip poses different challenges: the tip can be occluded by contrast during angiography or interventional devices; and it is always in continuous movement due to the cardiac and respiratory motions. To overcome these challenges, we propose ConTrack, a transformer-based network that uses both spatial and temporal contextual information for accurate device detection and tracking in both X-ray fluoroscopy and angiography. The spatial information comes from the template frames and the segmentation module: the template frames define the surroundings of the device, whereas the segmentation module detects the entire device to bring more context for the tip prediction. Using multiple templates makes the model more robust to the change in appearance of the device when it is occluded by the contrast agent. The flow information computed on the segmented catheter mask between the current and the previous frame helps in further refining the prediction by compensating for the respiratory and cardiac motions. The experiments show that our method achieves 45% or higher accuracy in detection and tracking when compared to state-of-the-art tracking models.
Generation of Radiology Findings in Chest X-Ray by Leveraging Collaborative Knowledge
Jun 18, 2023Manuela Daniela Danu, George Marica, Sanjeev Kumar Karn, Bogdan Georgescu, Awais Mansoor, Florin Ghesu, Lucian Mihai Itu, Constantin Suciu, Sasa Grbic, Oladimeji Farri, Dorin Comaniciu




Among all the sub-sections in a typical radiology report, the Clinical Indications, Findings, and Impression often reflect important details about the health status of a patient. The information included in Impression is also often covered in Findings. While Findings and Impression can be deduced by inspecting the image, Clinical Indications often require additional context. The cognitive task of interpreting medical images remains the most critical and often time-consuming step in the radiology workflow. Instead of generating an end-to-end radiology report, in this paper, we focus on generating the Findings from automated interpretation of medical images, specifically chest X-rays (CXRs). Thus, this work focuses on reducing the workload of radiologists who spend most of their time either writing or narrating the Findings. Unlike past research, which addresses radiology report generation as a single-step image captioning task, we have further taken into consideration the complexity of interpreting CXR images and propose a two-step approach: (a) detecting the regions with abnormalities in the image, and (b) generating relevant text for regions with abnormalities by employing a generative large language model (LLM). This two-step approach introduces a layer of interpretability and aligns the framework with the systematic reasoning that radiologists use when reviewing a CXR.
* Information Technology and Quantitative Management (ITQM 2023)
Self-supervised Learning from 100 Million Medical Images
Jan 04, 2022Florin C. Ghesu, Bogdan Georgescu, Awais Mansoor, Youngjin Yoo, Dominik Neumann, Pragneshkumar Patel, R. S. Vishwanath, James M. Balter, Yue Cao, Sasa Grbic, Dorin Comaniciu




Building accurate and robust artificial intelligence systems for medical image assessment requires not only the research and design of advanced deep learning models but also the creation of large and curated sets of annotated training examples. Constructing such datasets, however, is often very costly -- due to the complex nature of annotation tasks and the high level of expertise required for the interpretation of medical images (e.g., expert radiologists). To counter this limitation, we propose a method for self-supervised learning of rich image features based on contrastive learning and online feature clustering. For this purpose we leverage large training datasets of over 100,000,000 medical images of various modalities, including radiography, computed tomography (CT), magnetic resonance (MR) imaging and ultrasonography. We propose to use these features to guide model training in supervised and hybrid self-supervised/supervised regime on various downstream tasks. We highlight a number of advantages of this strategy on challenging image assessment problems in radiography, CT and MR: 1) Significant increase in accuracy compared to the state-of-the-art (e.g., AUC boost of 3-7% for detection of abnormalities from chest radiography scans and hemorrhage detection on brain CT); 2) Acceleration of model convergence during training by up to 85% compared to using no pretraining (e.g., 83% when training a model for detection of brain metastases in MR scans); 3) Increase in robustness to various image augmentations, such as intensity variations, rotations or scaling reflective of data variation seen in the field.
Robust Classification from Noisy Labels: Integrating Additional Knowledge for Chest Radiography Abnormality Assessment
Apr 21, 2021Sebastian Gündel, Arnaud A. A. Setio, Florin C. Ghesu, Sasa Grbic, Bogdan Georgescu, Andreas Maier, Dorin Comaniciu




Chest radiography is the most common radiographic examination performed in daily clinical practice for the detection of various heart and lung abnormalities. The large amount of data to be read and reported, with more than 100 studies per day for a single radiologist, poses a challenge in consistently maintaining high interpretation accuracy. The introduction of large-scale public datasets has led to a series of novel systems for automated abnormality classification. However, the labels of these datasets were obtained using natural language processed medical reports, yielding a large degree of label noise that can impact the performance. In this study, we propose novel training strategies that handle label noise from such suboptimal data. Prior label probabilities were measured on a subset of training data re-read by 4 board-certified radiologists and were used during training to increase the robustness of the training model to the label noise. Furthermore, we exploit the high comorbidity of abnormalities observed in chest radiography and incorporate this information to further reduce the impact of label noise. Additionally, anatomical knowledge is incorporated by training the system to predict lung and heart segmentation, as well as spatial knowledge labels. To deal with multiple datasets and images derived from various scanners that apply different post-processing techniques, we introduce a novel image normalization strategy. Experiments were performed on an extensive collection of 297,541 chest radiographs from 86,876 patients, leading to a state-of-the-art performance level for 17 abnormalities from 2 datasets. With an average AUC score of 0.880 across all abnormalities, our proposed training strategies can be used to significantly improve performance scores.
Automated detection and quantification of COVID-19 airspace disease on chest radiographs: A novel approach achieving radiologist-level performance using a CNN trained on digital reconstructed radiographs (DRRs) from CT-based ground-truth
Aug 13, 2020Eduardo Mortani Barbosa Jr., Warren B. Gefter, Rochelle Yang, Florin C. Ghesu, Siqi Liu, Boris Mailhe, Awais Mansoor, Sasa Grbic, Sebastian Piat, Guillaume Chabin, Vishwanath R S., Abishek Balachandran, Sebastian Vogt, Valentin Ziebandt, Steffen Kappler, Dorin Comaniciu




Purpose: To leverage volumetric quantification of airspace disease (AD) derived from a superior modality (CT) serving as ground truth, projected onto digitally reconstructed radiographs (DRRs) to: 1) train a convolutional neural network to quantify airspace disease on paired CXRs; and 2) compare the DRR-trained CNN to expert human readers in the CXR evaluation of patients with confirmed COVID-19. Materials and Methods: We retrospectively selected a cohort of 86 COVID-19 patients (with positive RT-PCR), from March-May 2020 at a tertiary hospital in the northeastern USA, who underwent chest CT and CXR within 48 hrs. The ground truth volumetric percentage of COVID-19 related AD (POv) was established by manual AD segmentation on CT. The resulting 3D masks were projected into 2D anterior-posterior digitally reconstructed radiographs (DRR) to compute area-based AD percentage (POa). A convolutional neural network (CNN) was trained with DRR images generated from a larger-scale CT dataset of COVID-19 and non-COVID-19 patients, automatically segmenting lungs, AD and quantifying POa on CXR. CNN POa results were compared to POa quantified on CXR by two expert readers and to the POv ground-truth, by computing correlations and mean absolute errors. Results: Bootstrap mean absolute error (MAE) and correlations between POa and POv were 11.98% [11.05%-12.47%] and 0.77 [0.70-0.82] for average of expert readers, and 9.56%-9.78% [8.83%-10.22%] and 0.78-0.81 [0.73-0.85] for the CNN, respectively. Conclusion: Our CNN trained with DRR using CT-derived airspace quantification achieved expert radiologist level of accuracy in the quantification of airspace disease on CXR, in patients with positive RT-PCR for COVID-19.
Extracting and Leveraging Nodule Features with Lung Inpainting for Local Feature Augmentation
Aug 05, 2020Sebastian Guendel, Arnaud Arindra Adiyoso Setio, Sasa Grbic, Andreas Maier, Dorin Comaniciu
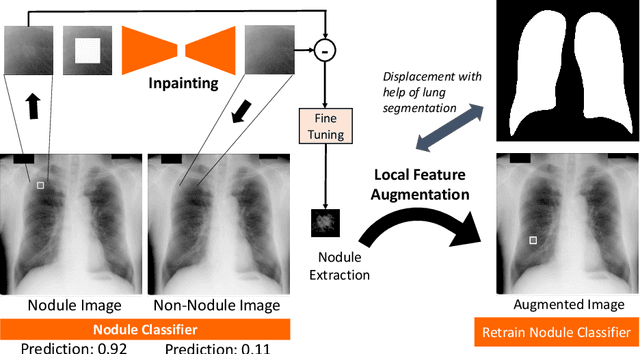
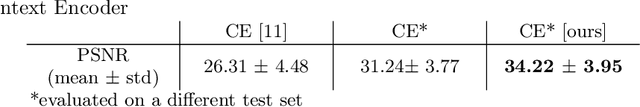
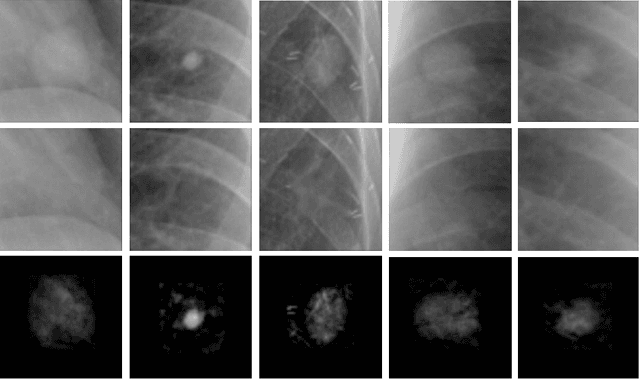
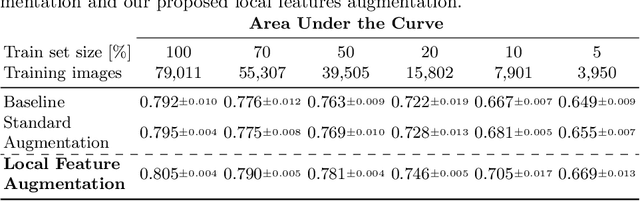
Chest X-ray (CXR) is the most common examination for fast detection of pulmonary abnormalities. Recently, automated algorithms have been developed to classify multiple diseases and abnormalities in CXR scans. However, because of the limited availability of scans containing nodules and the subtle properties of nodules in CXRs, state-of-the-art methods do not perform well on nodule classification. To create additional data for the training process, standard augmentation techniques are applied. However, the variance introduced by these methods are limited as the images are typically modified globally. In this paper, we propose a method for local feature augmentation by extracting local nodule features using a generative inpainting network. The network is applied to generate realistic, healthy tissue and structures in patches containing nodules. The nodules are entirely removed in the inpainted representation. The extraction of the nodule features is processed by subtraction of the inpainted patch from the nodule patch. With arbitrary displacement of the extracted nodules in the lung area across different CXR scans and further local modifications during training, we significantly increase the nodule classification performance and outperform state-of-the-art augmentation methods.
Quantifying and Leveraging Predictive Uncertainty for Medical Image Assessment
Jul 08, 2020Florin C. Ghesu, Bogdan Georgescu, Awais Mansoor, Youngjin Yoo, Eli Gibson, R. S. Vishwanath, Abishek Balachandran, James M. Balter, Yue Cao, Ramandeep Singh, Subba R. Digumarthy, Mannudeep K. Kalra, Sasa Grbic, Dorin Comaniciu
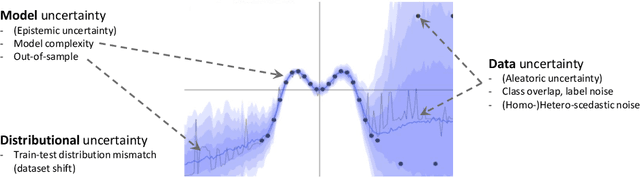

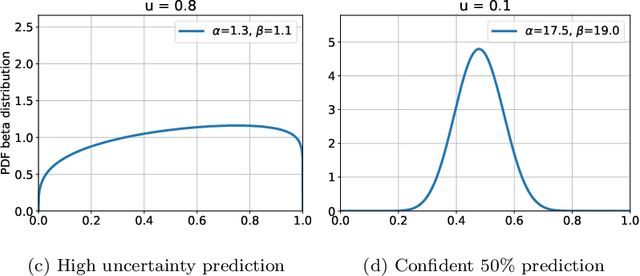
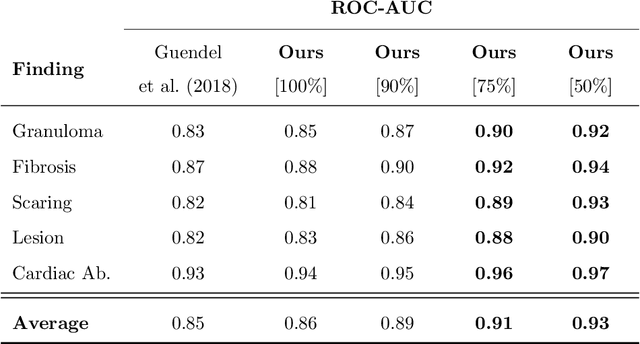
The interpretation of medical images is a challenging task, often complicated by the presence of artifacts, occlusions, limited contrast and more. Most notable is the case of chest radiography, where there is a high inter-rater variability in the detection and classification of abnormalities. This is largely due to inconclusive evidence in the data or subjective definitions of disease appearance. An additional example is the classification of anatomical views based on 2D Ultrasound images. Often, the anatomical context captured in a frame is not sufficient to recognize the underlying anatomy. Current machine learning solutions for these problems are typically limited to providing probabilistic predictions, relying on the capacity of underlying models to adapt to limited information and the high degree of label noise. In practice, however, this leads to overconfident systems with poor generalization on unseen data. To account for this, we propose a system that learns not only the probabilistic estimate for classification, but also an explicit uncertainty measure which captures the confidence of the system in the predicted output. We argue that this approach is essential to account for the inherent ambiguity characteristic of medical images from different radiologic exams including computed radiography, ultrasonography and magnetic resonance imaging. In our experiments we demonstrate that sample rejection based on the predicted uncertainty can significantly improve the ROC-AUC for various tasks, e.g., by 8% to 0.91 with an expected rejection rate of under 25% for the classification of different abnormalities in chest radiographs. In addition, we show that using uncertainty-driven bootstrapping to filter the training data, one can achieve a significant increase in robustness and accuracy.
Machine Learning Automatically Detects COVID-19 using Chest CTs in a Large Multicenter Cohort
Jun 11, 2020Bogdan Georgescu, Shikha Chaganti, Gorka Bastarrika Aleman, Eduardo Jose Mortani Barbosa Jr., Jordi Broncano Cabrero, Guillaume Chabin, Thomas Flohr, Philippe Grenier, Sasa Grbic, Nakul Gupta, François Mellot, Savvas Nicolaou, Thomas Re, Pina Sanelli, Alexander W. Sauter, Youngjin Yoo, Valentin Ziebandt, Dorin Comaniciu
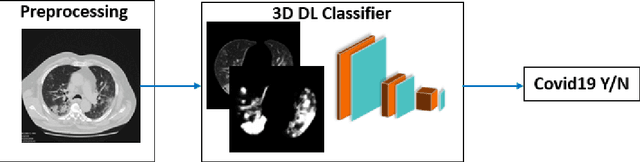
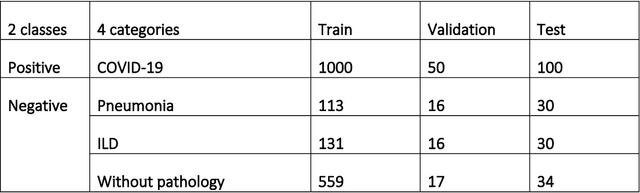
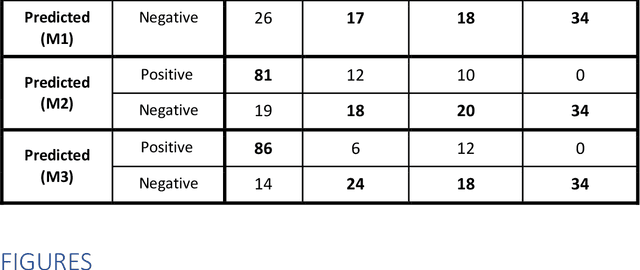
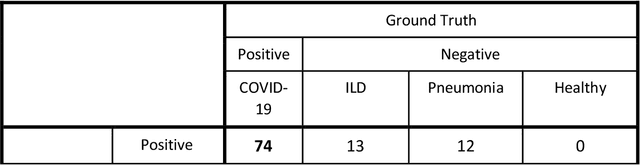

Purpose: To investigate if AI-based classifiers can distinguish COVID-19 from other pulmonary diseases and normal groups, using chest CT images. To study the interpretability of discriminative features for COVID19 detection. Materials and Methods: Our database consists of 2096 CT exams that include CTs from 1150 COVID-19 patients. Training was performed on 1000 COVID-19, 131 ILD, 113 other pneumonias, 559 normal CTs, and testing on 100 COVID-19, 30 ILD, 30 other pneumonias, and 34 normal CTs. A metric-based approach for classification of COVID-19 used interpretable features, relying on logistic regression and random forests. A deep learning-based classifier differentiated COVID-19 based on 3D features extracted directly from CT intensities and from the probability distribution of airspace opacities. Results: Most discriminative features of COVID-19 are percentage of airspace opacity, ground glass opacities, consolidations, and peripheral and basal opacities, which coincide with the typical characterization of COVID-19 in the literature. Unsupervised hierarchical clustering compares the distribution of these features across COVID-19 and control cohorts. The metrics-based classifier achieved AUC, sensitivity, and specificity of respectively 0.85, 0.81, and 0.77. The DL-based classifier achieved AUC, sensitivity, and specificity of respectively 0.90, 0.86, and 0.81. Most of ambiguity comes from non-COVID-19 pneumonia with manifestations that overlap with COVID-19, as well as COVID-19 cases in early stages. Conclusion: A new method discriminates COVID-19 from other types of pneumonia, ILD, and normal, using quantitative patterns from chest CT. Our models balance interpretability of results and classification performance, and therefore may be useful to expedite and improve diagnosis of COVID-19.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge