Goal-conditioned reinforcement learning for ultrasound navigation guidance
May 02, 2024Abdoul Aziz Amadou, Vivek Singh, Florin C. Ghesu, Young-Ho Kim, Laura Stanciulescu, Harshitha P. Sai, Puneet Sharma, Alistair Young, Ronak Rajani, Kawal Rhode
Transesophageal echocardiography (TEE) plays a pivotal role in cardiology for diagnostic and interventional procedures. However, using it effectively requires extensive training due to the intricate nature of image acquisition and interpretation. To enhance the efficiency of novice sonographers and reduce variability in scan acquisitions, we propose a novel ultrasound (US) navigation assistance method based on contrastive learning as goal-conditioned reinforcement learning (GCRL). We augment the previous framework using a novel contrastive patient batching method (CPB) and a data-augmented contrastive loss, both of which we demonstrate are essential to ensure generalization to anatomical variations across patients. The proposed framework enables navigation to both standard diagnostic as well as intricate interventional views with a single model. Our method was developed with a large dataset of 789 patients and obtained an average error of 6.56 mm in position and 9.36 degrees in angle on a testing dataset of 140 patients, which is competitive or superior to models trained on individual views. Furthermore, we quantitatively validate our method's ability to navigate to interventional views such as the Left Atrial Appendage (LAA) view used in LAA closure. Our approach holds promise in providing valuable guidance during transesophageal ultrasound examinations, contributing to the advancement of skill acquisition for cardiac ultrasound practitioners.
Self-Supervised Learning for Interventional Image Analytics: Towards Robust Device Trackers
May 02, 2024Saahil Islam, Venkatesh N. Murthy, Dominik Neumann, Badhan Kumar Das, Puneet Sharma, Andreas Maier, Dorin Comaniciu, Florin C. Ghesu
An accurate detection and tracking of devices such as guiding catheters in live X-ray image acquisitions is an essential prerequisite for endovascular cardiac interventions. This information is leveraged for procedural guidance, e.g., directing stent placements. To ensure procedural safety and efficacy, there is a need for high robustness no failures during tracking. To achieve that, one needs to efficiently tackle challenges, such as: device obscuration by contrast agent or other external devices or wires, changes in field-of-view or acquisition angle, as well as the continuous movement due to cardiac and respiratory motion. To overcome the aforementioned challenges, we propose a novel approach to learn spatio-temporal features from a very large data cohort of over 16 million interventional X-ray frames using self-supervision for image sequence data. Our approach is based on a masked image modeling technique that leverages frame interpolation based reconstruction to learn fine inter-frame temporal correspondences. The features encoded in the resulting model are fine-tuned downstream. Our approach achieves state-of-the-art performance and in particular robustness compared to ultra optimized reference solutions (that use multi-stage feature fusion, multi-task and flow regularization). The experiments show that our method achieves 66.31% reduction in maximum tracking error against reference solutions (23.20% when flow regularization is used); achieving a success score of 97.95% at a 3x faster inference speed of 42 frames-per-second (on GPU). The results encourage the use of our approach in various other tasks within interventional image analytics that require effective understanding of spatio-temporal semantics.
Towards Establishing Dense Correspondence on Multiview Coronary Angiography: From Point-to-Point to Curve-to-Curve Query Matching
Dec 18, 2023Yifan Wu, Rohit Jena, Mehmet Gulsun, Vivek Singh, Puneet Sharma, James C. Gee
Coronary angiography is the gold standard imaging technique for studying and diagnosing coronary artery disease. However, the resulting 2D X-ray projections lose 3D information and exhibit visual ambiguities. In this work, we aim to establish dense correspondence in multi-view angiography, serving as a fundamental basis for various clinical applications and downstream tasks. To overcome the challenge of unavailable annotated data, we designed a data simulation pipeline using 3D Coronary Computed Tomography Angiography (CCTA). We formulated the problem of dense correspondence estimation as a query matching task over all points of interest in the given views. We established point-to-point query matching and advanced it to curve-to-curve correspondence, significantly reducing errors by minimizing ambiguity and improving topological awareness. The method was evaluated on a set of 1260 image pairs from different views across 8 clinically relevant angulation groups, demonstrating compelling results and indicating the feasibility of establishing dense correspondence in multi-view angiography.
AI-based, automated chamber volumetry from gated, non-contrast CT
Oct 25, 2023Athira J Jacob, Ola Abdelkarim, Salma Zook, Kristian Hay Kragholm, Prantik Gupta, Myra Cocker, Juan Ramirez Giraldo, Jim O Doherty, Max Schoebinger, Chris Schwemmer, Mehmet A Gulsun, Saikiran Rapaka, Puneet Sharma, Su-Min Chang
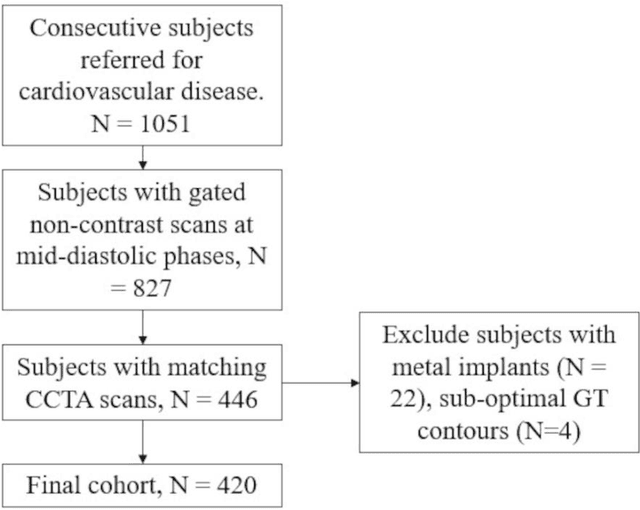
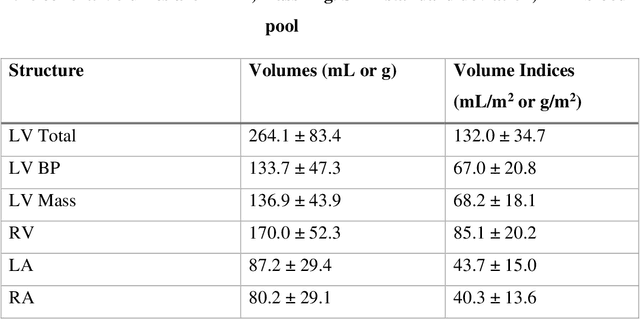
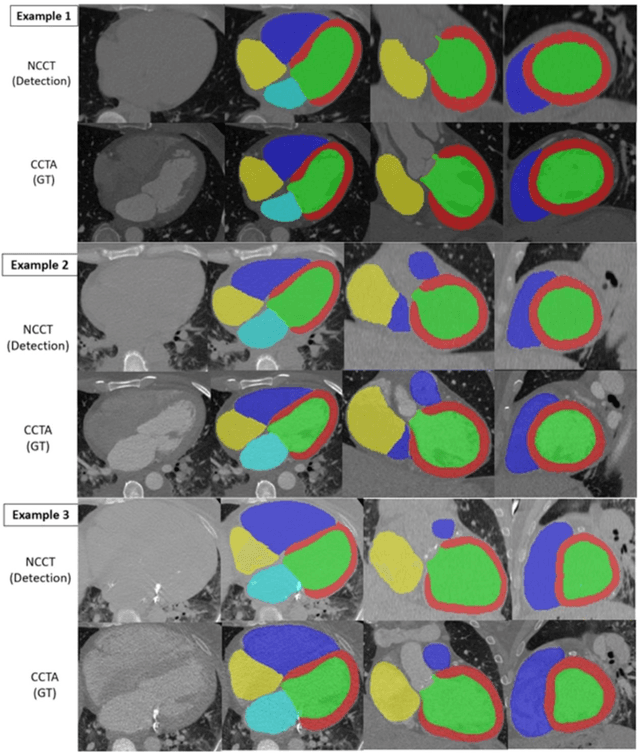
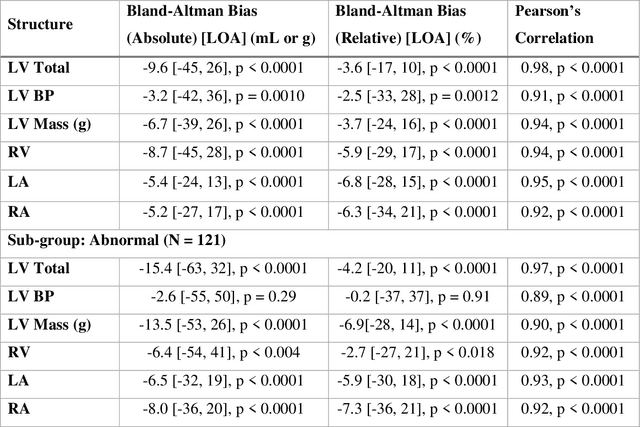
Background: Accurate chamber volumetry from gated, non-contrast cardiac CT (NCCT) scans can be useful for potential screening of heart failure. Objectives: To validate a new, fully automated, AI-based method for cardiac volume and myocardial mass quantification from NCCT scans compared to contrasted CT Angiography (CCTA). Methods: Of a retrospectively collected cohort of 1051 consecutive patients, 420 patients had both NCCT and CCTA scans at mid-diastolic phase, excluding patients with cardiac devices. Ground truth values were obtained from the CCTA scans. Results: The NCCT volume computation shows good agreement with ground truth values. Volume differences [95% CI ] and correlation coefficients were: -9.6 [-45; 26] mL, r = 0.98 for LV Total, -5.4 [-24; 13] mL, r = 0.95 for LA, -8.7 [-45; 28] mL, r = 0.94 for RV, -5.2 [-27; 17] mL, r = 0.92 for RA, -3.2 [-42; 36] mL, r = 0.91 for LV blood pool, and -6.7 [-39; 26] g, r = 0.94 for LV wall mass, respectively. Mean relative volume errors of less than 7% were obtained for all chambers. Conclusions: Fully automated assessment of chamber volumes from NCCT scans is feasible and correlates well with volumes obtained from contrast study.
Deep Conditional Shape Models for 3D cardiac image segmentation
Oct 16, 2023Athira J Jacob, Puneet Sharma, Daniel Ruckert

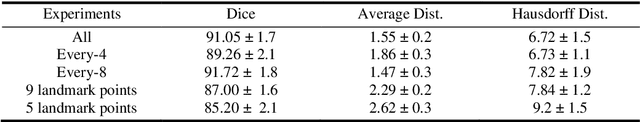


Delineation of anatomical structures is often the first step of many medical image analysis workflows. While convolutional neural networks achieve high performance, these do not incorporate anatomical shape information. We introduce a novel segmentation algorithm that uses Deep Conditional Shape models (DCSMs) as a core component. Using deep implicit shape representations, the algorithm learns a modality-agnostic shape model that can generate the signed distance functions for any anatomy of interest. To fit the generated shape to the image, the shape model is conditioned on anatomic landmarks that can be automatically detected or provided by the user. Finally, we add a modality-dependent, lightweight refinement network to capture any fine details not represented by the implicit function. The proposed DCSM framework is evaluated on the problem of cardiac left ventricle (LV) segmentation from multiple 3D modalities (contrast-enhanced CT, non-contrasted CT, 3D echocardiography-3DE). We demonstrate that the automatic DCSM outperforms the baseline for non-contrasted CT without the local refinement, and with the refinement for contrasted CT and 3DE, especially with significant improvement in the Hausdorff distance. The semi-automatic DCSM with user-input landmarks, while only trained on contrasted CT, achieves greater than 92% Dice for all modalities. Both automatic DCSM with refinement and semi-automatic DCSM achieve equivalent or better performance compared to inter-user variability for these modalities.
Anatomically aware dual-hop learning for pulmonary embolism detection in CT pulmonary angiograms
Mar 30, 2023Florin Condrea, Saikiran Rapaka, Lucian Itu, Puneet Sharma, Jonathan Sperl, A Mohamed Ali, Marius Leordeanu
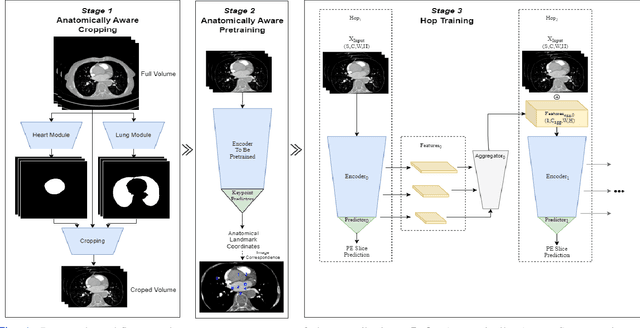
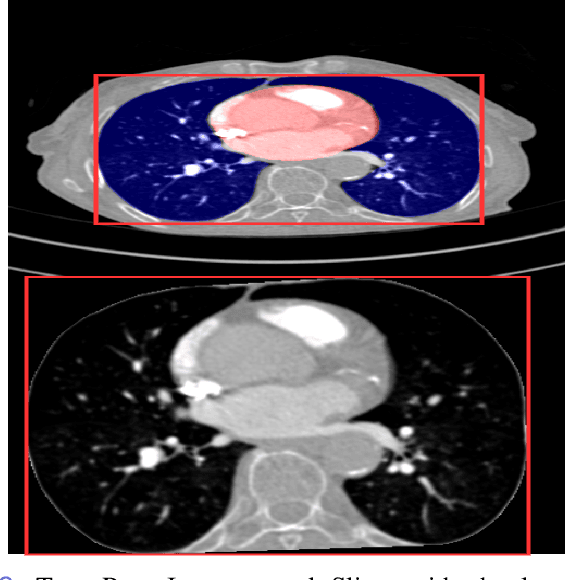
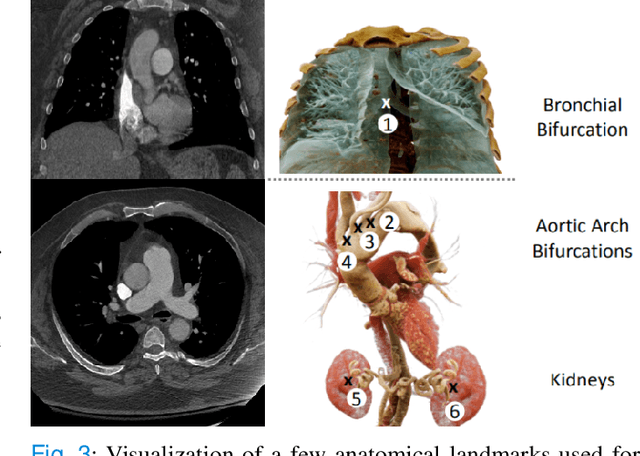
Pulmonary Embolisms (PE) represent a leading cause of cardiovascular death. While medical imaging, through computed tomographic pulmonary angiography (CTPA), represents the gold standard for PE diagnosis, it is still susceptible to misdiagnosis or significant diagnosis delays, which may be fatal for critical cases. Despite the recently demonstrated power of deep learning to bring a significant boost in performance in a wide range of medical imaging tasks, there are still very few published researches on automatic pulmonary embolism detection. Herein we introduce a deep learning based approach, which efficiently combines computer vision and deep neural networks for pulmonary embolism detection in CTPA. Our method features novel improvements along three orthogonal axes: 1) automatic detection of anatomical structures; 2) anatomical aware pretraining, and 3) a dual-hop deep neural net for PE detection. We obtain state-of-the-art results on the publicly available multicenter large-scale RSNA dataset.
Automated Cardiac Resting Phase Detection Targeted on the Right Coronary Artery
Sep 06, 2021Seung Su Yoon, Elisabeth Preuhs, Michaela Schmidt, Christoph Forman, Teodora Chitiboi, Puneet Sharma, Juliano Lara Fernandes, Christoph Tillmanns, Jens Wetzl, Andreas Maier
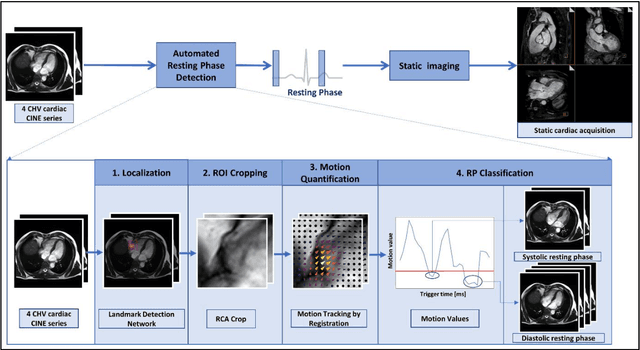

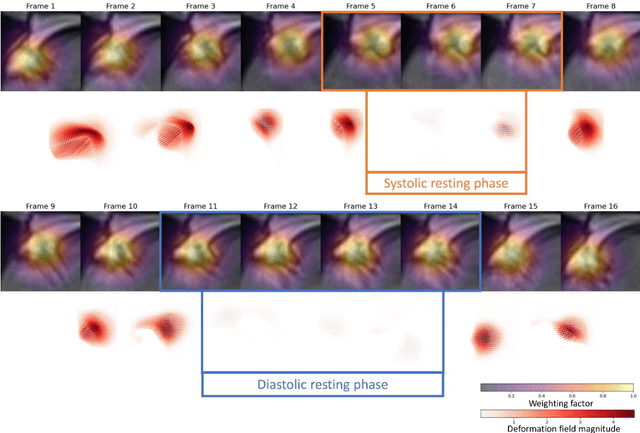
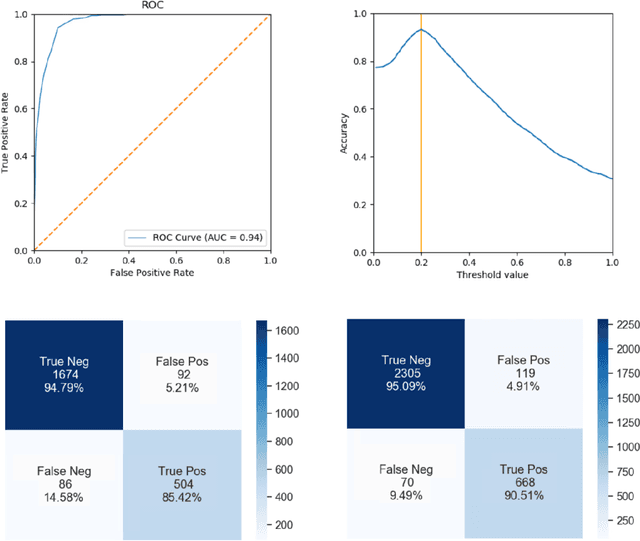
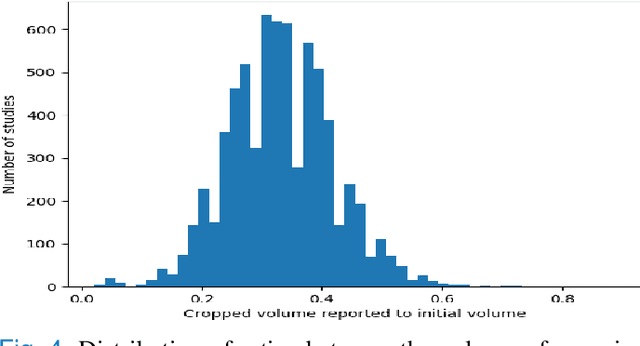
Purpose: Static cardiac imaging such as late gadolinium enhancement, mapping, or 3-D coronary angiography require prior information, e.g., the phase during a cardiac cycle with least motion, called resting phase (RP). The purpose of this work is to propose a fully automated framework that allows the detection of the right coronary artery (RCA) RP within CINE series. Methods: The proposed prototype system consists of three main steps. First, the localization of the regions of interest (ROI) is performed. Second, as CINE series are time-resolved, the cropped ROI series over all time points are taken for tracking motions quantitatively. Third, the output motion values are used to classify RPs. In this work, we focused on the detection of the area with the outer edge of the cross-section of the RCA as our target. The proposed framework was evaluated on 102 clinically acquired dataset at 1.5T and 3T. The automatically classified RPs were compared with the ground truth RPs annotated manually by a medical expert for testing the robustness and feasibility of the framework. Results: The predicted RCA RPs showed high agreement with the experts annotated RPs with 92.7% accuracy, 90.5% sensitivity and 95.0% specificity for the unseen study dataset. The mean absolute difference of the start and end RP was 13.6 ${\pm}$ 18.6 ms for the validation study dataset (n=102). Conclusion: In this work, automated RP detection has been introduced by the proposed framework and demonstrated feasibility, robustness, and applicability for diverse static imaging acquisitions.
Spatial Sharing of GPU for Autotuning DNN models
Aug 08, 2020Aditya Dhakal, Junguk Cho, Sameer G. Kulkarni, K. K. Ramakrishnan, Puneet Sharma
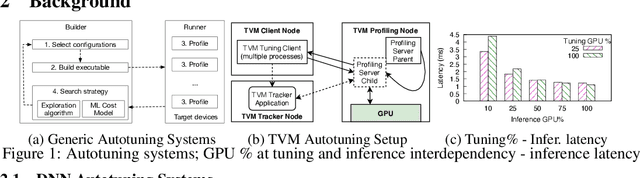
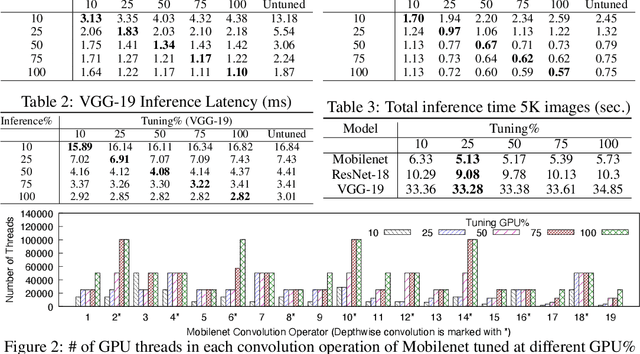
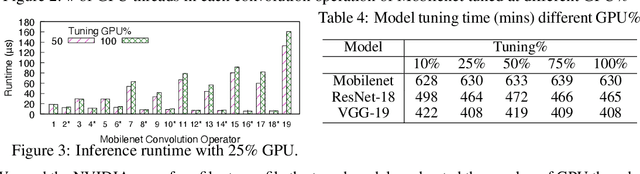
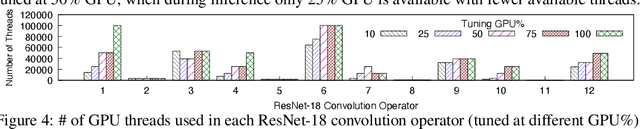
GPUs are used for training, inference, and tuning the machine learning models. However, Deep Neural Network (DNN) vary widely in their ability to exploit the full power of high-performance GPUs. Spatial sharing of GPU enables multiplexing several DNNs on the GPU and can improve GPU utilization, thus improving throughput and lowering latency. DNN models given just the right amount of GPU resources can still provide low inference latency, just as much as dedicating all of the GPU for their inference task. An approach to improve DNN inference is tuning of the DNN model. Autotuning frameworks find the optimal low-level implementation for a certain target device based on the trained machine learning model, thus reducing the DNN's inference latency and increasing inference throughput. We observe an interdependency between the tuned model and its inference latency. A DNN model tuned with specific GPU resources provides the best inference latency when inferred with close to the same amount of GPU resources. While a model tuned with the maximum amount of the GPU's resources has poorer inference latency once the GPU resources are limited for inference. On the other hand, a model tuned with an appropriate amount of GPU resources still achieves good inference latency across a wide range of GPU resource availability. We explore the causes that impact the tuning of a model at different amounts of GPU resources. We present many techniques to maximize resource utilization and improve tuning performance. We enable controlled spatial sharing of GPU to multiplex several tuning applications on the GPU. We scale the tuning server instances and shard the tuning model across multiple client instances for concurrent tuning of different operators of a model, achieving better GPU multiplexing. With our improvements, we decrease DNN autotuning time by up to 75 percent and increase throughput by a factor of 5.
Assertion Detection in Multi-Label Clinical Text using Scope Localization
May 19, 2020Rajeev Bhatt Ambati, Ahmed Ada Hanifi, Ramya Vunikili, Puneet Sharma, Oladimeji Farri
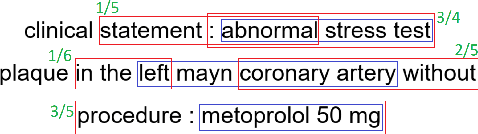




Multi-label sentences (text) in the clinical domain result from the rich description of scenarios during patient care. The state-of-theart methods for assertion detection mostly address this task in the setting of a single assertion label per sentence (text). In addition, few rules based and deep learning methods perform negation/assertion scope detection on single-label text. It is a significant challenge extending these methods to address multi-label sentences without diminishing performance. Therefore, we developed a convolutional neural network (CNN) architecture to localize multiple labels and their scopes in a single stage end-to-end fashion, and demonstrate that our model performs atleast 12% better than the state-of-the-art on multi-label clinical text.
Deep Neural Networks for ECG-free Cardiac Phase and End-Diastolic Frame Detection on Coronary Angiographies
Nov 07, 2018Costin Ciusdel, Alexandru Turcea, Andrei Puiu, Lucian Itu, Lucian Calmac, Emma Weiss, Cornelia Margineanu, Elisabeta Badila, Martin Berger, Thomas Redel, Tiziano Passerini, Mehmet Gulsun, Puneet Sharma

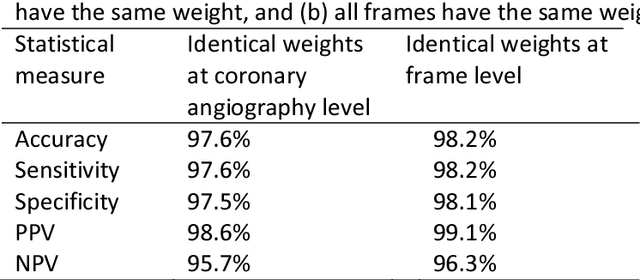
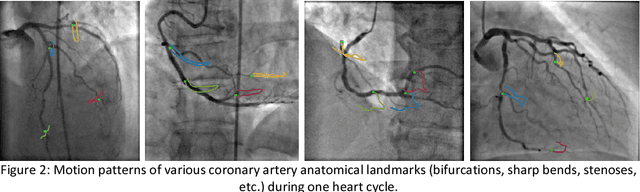
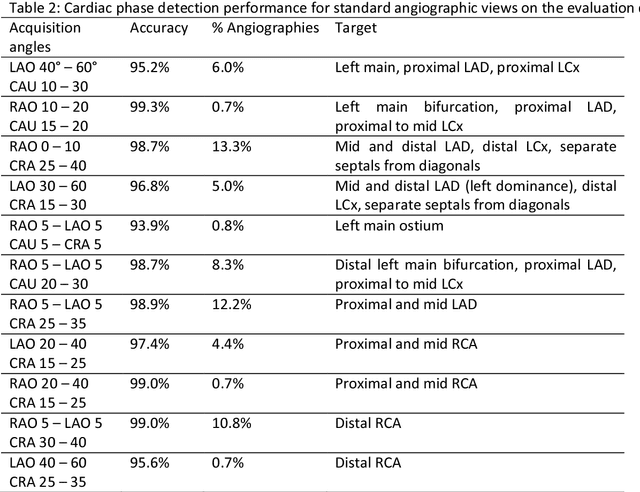
Invasive coronary angiography (ICA) is the gold standard in Coronary Artery Disease (CAD) imaging. Detection of the end-diastolic frame (EDF) and, in general, cardiac phase detection on each temporal frame of a coronary angiography acquisition is of significant importance for the anatomical and non-invasive functional assessment of CAD. This task is generally performed via manual frame selection or semi-automated selection based on simultaneously acquired ECG signals - thus introducing the requirement of simultaneous ECG recordings. We evaluate the performance of a purely image based workflow based on deep neural networks for fully automated cardiac phase and EDF detection on coronary angiographies. A first deep neural network (DNN), trained to detect coronary arteries, is employed to preselect a subset of frames in which coronary arteries are well visible. A second DNN predicts cardiac phase labels for each frame. Only in the training and evaluation phases for the second DNN, ECG signals are used to provide ground truth labels for each angiographic frame. The networks were trained on 17800 coronary angiographies from 3900 patients and evaluated on 27900 coronary angiographies from 6250 patients. No exclusion criteria related to patient state, previous interventions, or pathology were formulated. Cardiac phase detection had an accuracy of 97.6%, a sensitivity of 97.6% and a specificity of 97.5% on the evaluation set. EDF prediction had a precision of 97.4% and a recall of 96.9%. Several sub-group analyses were performed, indicating that the cardiac phase detection performance is largely independent from acquisition angles and the heart rate of the patient. The average execution time of cardiac phase detection for one angiographic series was on average less than five seconds on a standard workstation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge