Neural Ordinary Differential Equation based Sequential Image Registration for Dynamic Characterization
Apr 02, 2024Yifan Wu, Mengjin Dong, Rohit Jena, Chen Qin, James C. Gee
Deformable image registration (DIR) is crucial in medical image analysis, enabling the exploration of biological dynamics such as organ motions and longitudinal changes in imaging. Leveraging Neural Ordinary Differential Equations (ODE) for registration, this extension work discusses how this framework can aid in the characterization of sequential biological processes. Utilizing the Neural ODE's ability to model state derivatives with neural networks, our Neural Ordinary Differential Equation Optimization-based (NODEO) framework considers voxels as particles within a dynamic system, defining deformation fields through the integration of neural differential equations. This method learns dynamics directly from data, bypassing the need for physical priors, making it exceptionally suitable for medical scenarios where such priors are unavailable or inapplicable. Consequently, the framework can discern underlying dynamics and use sequence data to regularize the transformation trajectory. We evaluated our framework on two clinical datasets: one for cardiac motion tracking and another for longitudinal brain MRI analysis. Demonstrating its efficacy in both 2D and 3D imaging scenarios, our framework offers flexibility and model agnosticism, capable of managing image sequences and facilitating label propagation throughout these sequences. This study provides a comprehensive understanding of how the Neural ODE-based framework uniquely benefits the image registration challenge.
FireANTs: Adaptive Riemannian Optimization for Multi-Scale Diffeomorphic Registration
Apr 01, 2024Rohit Jena, Pratik Chaudhari, James C. Gee
Diffeomorphic Image Registration is a critical part of the analysis in various imaging modalities and downstream tasks like image translation, segmentation, and atlas building. Registration algorithms based on optimization have stood the test of time in terms of accuracy, reliability, and robustness across a wide spectrum of modalities and acquisition settings. However, these algorithms converge slowly, are prohibitively expensive to run, and their usage requires a steep learning curve, limiting their scalability to larger clinical and scientific studies. In this paper, we develop multi-scale Adaptive Riemannian Optimization algorithms for diffeomorphic image registration. We demonstrate compelling improvements on image registration across a spectrum of modalities and anatomies by measuring structural and landmark overlap of the registered image volumes. Our proposed framework leads to a consistent improvement in performance, and from 300x up to 2000x speedup over existing algorithms. Our modular library design makes it easy to use and allows customization via user-defined cost functions.
A Concept-based Interpretable Model for the Diagnosis of Choroid Neoplasias using Multimodal Data
Mar 08, 2024Yifan Wu, Yang Liu, Yue Yang, Michael S. Yao, Wenli Yang, Xuehui Shi, Lihong Yang, Dongjun Li, Yueming Liu, James C. Gee, Xuan Yang, Wenbin Wei, Shi Gu
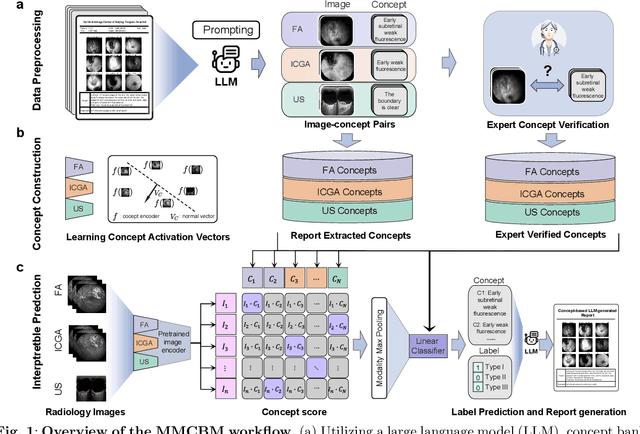
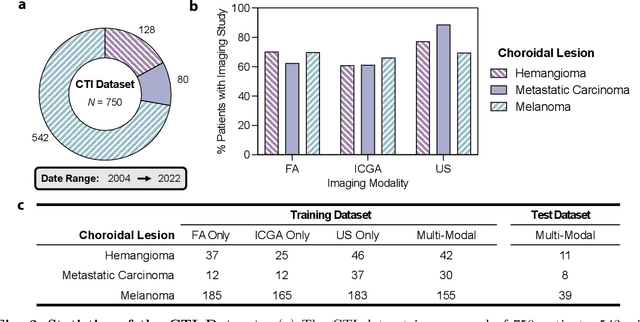
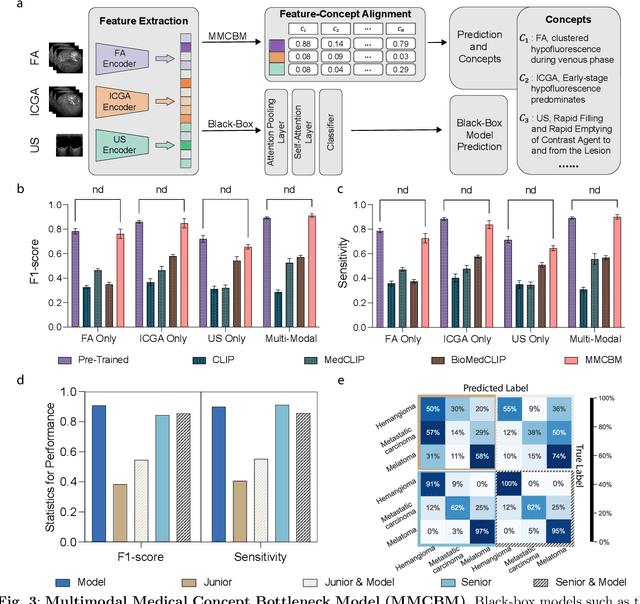
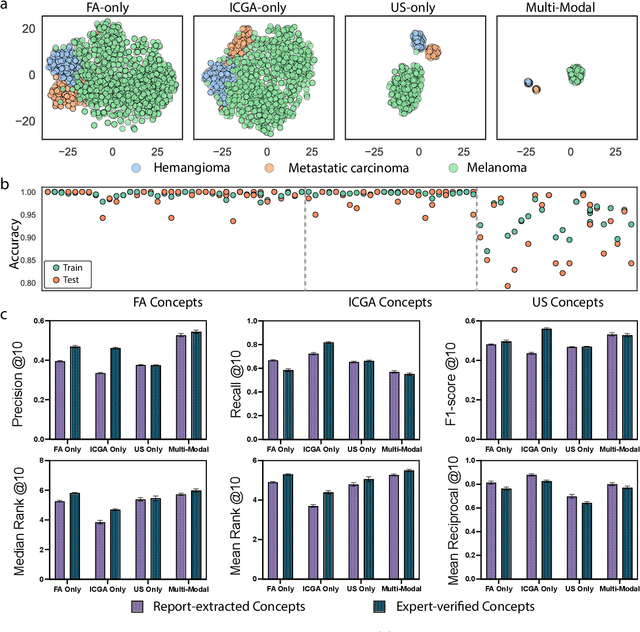
Diagnosing rare diseases presents a common challenge in clinical practice, necessitating the expertise of specialists for accurate identification. The advent of machine learning offers a promising solution, while the development of such technologies is hindered by the scarcity of data on rare conditions and the demand for models that are both interpretable and trustworthy in a clinical context. Interpretable AI, with its capacity for human-readable outputs, can facilitate validation by clinicians and contribute to medical education. In the current work, we focus on choroid neoplasias, the most prevalent form of eye cancer in adults, albeit rare with 5.1 per million. We built the so-far largest dataset consisting of 750 patients, incorporating three distinct imaging modalities collected from 2004 to 2022. Our work introduces a concept-based interpretable model that distinguishes between three types of choroidal tumors, integrating insights from domain experts via radiological reports. Remarkably, this model not only achieves an F1 score of 0.91, rivaling that of black-box models, but also boosts the diagnostic accuracy of junior doctors by 42%. This study highlights the significant potential of interpretable machine learning in improving the diagnosis of rare diseases, laying a groundwork for future breakthroughs in medical AI that could tackle a wider array of complex health scenarios.
Generative Adversarial Bayesian Optimization for Surrogate Objectives
Feb 09, 2024Michael S. Yao, Yimeng Zeng, Hamsa Bastani, Jacob Gardner, James C. Gee, Osbert Bastani
Offline model-based policy optimization seeks to optimize a learned surrogate objective function without querying the true oracle objective during optimization. However, inaccurate surrogate model predictions are frequently encountered along the optimization trajectory. To address this limitation, we propose generative adversarial Bayesian optimization (GABO) using adaptive source critic regularization, a task-agnostic framework for Bayesian optimization that employs a Lipschitz-bounded source critic model to constrain the optimization trajectory to regions where the surrogate function is reliable. We show that under certain assumptions for the continuous input space prior, our algorithm dynamically adjusts the strength of the source critic regularization. GABO outperforms existing baselines on a number of different offline optimization tasks across a variety of scientific domains. Our code is available at https://github.com/michael-s-yao/gabo
Towards Establishing Dense Correspondence on Multiview Coronary Angiography: From Point-to-Point to Curve-to-Curve Query Matching
Dec 18, 2023Yifan Wu, Rohit Jena, Mehmet Gulsun, Vivek Singh, Puneet Sharma, James C. Gee
Coronary angiography is the gold standard imaging technique for studying and diagnosing coronary artery disease. However, the resulting 2D X-ray projections lose 3D information and exhibit visual ambiguities. In this work, we aim to establish dense correspondence in multi-view angiography, serving as a fundamental basis for various clinical applications and downstream tasks. To overcome the challenge of unavailable annotated data, we designed a data simulation pipeline using 3D Coronary Computed Tomography Angiography (CCTA). We formulated the problem of dense correspondence estimation as a query matching task over all points of interest in the given views. We established point-to-point query matching and advanced it to curve-to-curve correspondence, significantly reducing errors by minimizing ambiguity and improving topological awareness. The method was evaluated on a set of 1260 image pairs from different views across 8 clinically relevant angulation groups, demonstrating compelling results and indicating the feasibility of establishing dense correspondence in multi-view angiography.
The Role of Chain-of-Thought in Complex Vision-Language Reasoning Task
Nov 15, 2023Yifan Wu, Pengchuan Zhang, Wenhan Xiong, Barlas Oguz, James C. Gee, Yixin Nie
The study explores the effectiveness of the Chain-of-Thought approach, known for its proficiency in language tasks by breaking them down into sub-tasks and intermediate steps, in improving vision-language tasks that demand sophisticated perception and reasoning. We present the "Description then Decision" strategy, which is inspired by how humans process signals. This strategy significantly improves probing task performance by 50%, establishing the groundwork for future research on reasoning paradigms in complex vision-language tasks.
Deformable Image Registration using Neural ODEs
Aug 27, 2021Yifan Wu, Tom Z. Jiahao, Jiancong Wang, Paul A. Yushkevich, James C. Gee, M. Ani Hsieh




Deformable image registration, aiming to find spatial correspondence between a given image pair, is one of the most critical problems in the domain of medical image analysis. In this paper, we present a generic, fast, and accurate diffeomorphic image registration framework that leverages neural ordinary differential equations (NODEs). We model each voxel as a moving particle and consider the set of all voxels in a 3D image as a high-dimensional dynamical system whose trajectory determines the targeted deformation field. Compared with traditional optimization-based methods, our framework reduces the running time from tens of minutes to tens of seconds. Compared with recent data-driven deep learning methods, our framework is more accessible since it does not require large amounts of training data. Our experiments show that the registration results of our method outperform state-of-the-arts under various metrics, indicating that our modeling approach is well fitted for the task of deformable image registration.
Nested Scale Editing for Conditional Image Synthesis
Jun 03, 2020Lingzhi Zhang, Jiancong Wang, Yinshuang Xu, Jie Min, Tarmily Wen, James C. Gee, Jianbo Shi

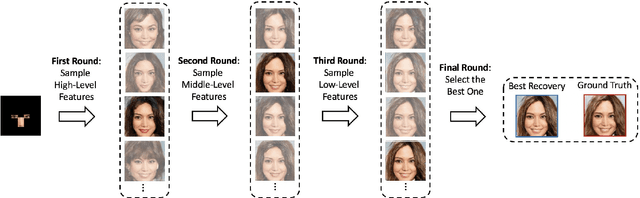
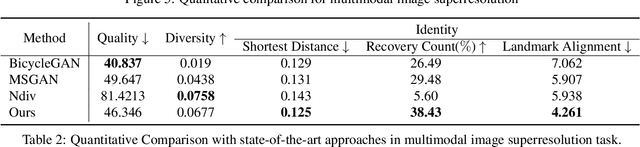
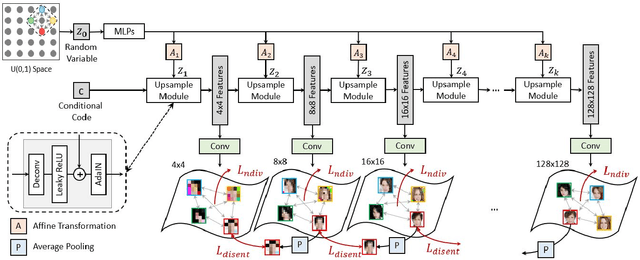
We propose an image synthesis approach that provides stratified navigation in the latent code space. With a tiny amount of partial or very low-resolution image, our approach can consistently out-perform state-of-the-art counterparts in terms of generating the closest sampled image to the ground truth. We achieve this through scale-independent editing while expanding scale-specific diversity. Scale-independence is achieved with a nested scale disentanglement loss. Scale-specific diversity is created by incorporating a progressive diversification constraint. We introduce semantic persistency across the scales by sharing common latent codes. Together they provide better control of the image synthesis process. We evaluate the effectiveness of our proposed approach through various tasks, including image outpainting, image superresolution, and cross-domain image translation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge