Cohort-Individual Cooperative Learning for Multimodal Cancer Survival Analysis
Apr 03, 2024Huajun Zhou, Fengtao Zhou, Hao Chen
Recently, we have witnessed impressive achievements in cancer survival analysis by integrating multimodal data, e.g., pathology images and genomic profiles. However, the heterogeneity and high dimensionality of these modalities pose significant challenges for extracting discriminative representations while maintaining good generalization. In this paper, we propose a Cohort-individual Cooperative Learning (CCL) framework to advance cancer survival analysis by collaborating knowledge decomposition and cohort guidance. Specifically, first, we propose a Multimodal Knowledge Decomposition (MKD) module to explicitly decompose multimodal knowledge into four distinct components: redundancy, synergy and uniqueness of the two modalities. Such a comprehensive decomposition can enlighten the models to perceive easily overlooked yet important information, facilitating an effective multimodal fusion. Second, we propose a Cohort Guidance Modeling (CGM) to mitigate the risk of overfitting task-irrelevant information. It can promote a more comprehensive and robust understanding of the underlying multimodal data, while avoiding the pitfalls of overfitting and enhancing the generalization ability of the model. By cooperating the knowledge decomposition and cohort guidance methods, we develop a robust multimodal survival analysis model with enhanced discrimination and generalization abilities. Extensive experimental results on five cancer datasets demonstrate the effectiveness of our model in integrating multimodal data for survival analysis.
iMD4GC: Incomplete Multimodal Data Integration to Advance Precise Treatment Response Prediction and Survival Analysis for Gastric Cancer
Apr 01, 2024Fengtao Zhou, Yingxue Xu, Yanfen Cui, Shenyan Zhang, Yun Zhu, Weiyang He, Jiguang Wang, Xin Wang, Ronald Chan, Louis Ho Shing Lau, Chu Han, Dafu Zhang, Zhenhui Li, Hao Chen
Gastric cancer (GC) is a prevalent malignancy worldwide, ranking as the fifth most common cancer with over 1 million new cases and 700 thousand deaths in 2020. Locally advanced gastric cancer (LAGC) accounts for approximately two-thirds of GC diagnoses, and neoadjuvant chemotherapy (NACT) has emerged as the standard treatment for LAGC. However, the effectiveness of NACT varies significantly among patients, with a considerable subset displaying treatment resistance. Ineffective NACT not only leads to adverse effects but also misses the optimal therapeutic window, resulting in lower survival rate. However, existing multimodal learning methods assume the availability of all modalities for each patient, which does not align with the reality of clinical practice. The limited availability of modalities for each patient would cause information loss, adversely affecting predictive accuracy. In this study, we propose an incomplete multimodal data integration framework for GC (iMD4GC) to address the challenges posed by incomplete multimodal data, enabling precise response prediction and survival analysis. Specifically, iMD4GC incorporates unimodal attention layers for each modality to capture intra-modal information. Subsequently, the cross-modal interaction layers explore potential inter-modal interactions and capture complementary information across modalities, thereby enabling information compensation for missing modalities. To evaluate iMD4GC, we collected three multimodal datasets for GC study: GastricRes (698 cases) for response prediction, GastricSur (801 cases) for survival analysis, and TCGA-STAD (400 cases) for survival analysis. The scale of our datasets is significantly larger than previous studies. The iMD4GC achieved impressive performance with an 80.2% AUC on GastricRes, 71.4% C-index on GastricSur, and 66.1% C-index on TCGA-STAD, significantly surpassing other compared methods.
Feature Re-Embedding: Towards Foundation Model-Level Performance in Computational Pathology
Feb 27, 2024Wenhao Tang, Fengtao Zhou, Sheng Huang, Xiang Zhu, Yi Zhang, Bo Liu
Multiple instance learning (MIL) is the most widely used framework in computational pathology, encompassing sub-typing, diagnosis, prognosis, and more. However, the existing MIL paradigm typically requires an offline instance feature extractor, such as a pre-trained ResNet or a foundation model. This approach lacks the capability for feature fine-tuning within the specific downstream tasks, limiting its adaptability and performance. To address this issue, we propose a Re-embedded Regional Transformer (R$^2$T) for re-embedding the instance features online, which captures fine-grained local features and establishes connections across different regions. Unlike existing works that focus on pre-training powerful feature extractor or designing sophisticated instance aggregator, R$^2$T is tailored to re-embed instance features online. It serves as a portable module that can seamlessly integrate into mainstream MIL models. Extensive experimental results on common computational pathology tasks validate that: 1) feature re-embedding improves the performance of MIL models based on ResNet-50 features to the level of foundation model features, and further enhances the performance of foundation model features; 2) the R$^2$T can introduce more significant performance improvements to various MIL models; 3) R$^2$T-MIL, as an R$^2$T-enhanced AB-MIL, outperforms other latest methods by a large margin. The code is available at:~\href{https://github.com/DearCaat/RRT-MIL}{https://github.com/DearCaat/RRT-MIL}.
Cross-Modal Translation and Alignment for Survival Analysis
Sep 22, 2023Fengtao Zhou, Hao Chen




With the rapid advances in high-throughput sequencing technologies, the focus of survival analysis has shifted from examining clinical indicators to incorporating genomic profiles with pathological images. However, existing methods either directly adopt a straightforward fusion of pathological features and genomic profiles for survival prediction, or take genomic profiles as guidance to integrate the features of pathological images. The former would overlook intrinsic cross-modal correlations. The latter would discard pathological information irrelevant to gene expression. To address these issues, we present a Cross-Modal Translation and Alignment (CMTA) framework to explore the intrinsic cross-modal correlations and transfer potential complementary information. Specifically, we construct two parallel encoder-decoder structures for multi-modal data to integrate intra-modal information and generate cross-modal representation. Taking the generated cross-modal representation to enhance and recalibrate intra-modal representation can significantly improve its discrimination for comprehensive survival analysis. To explore the intrinsic crossmodal correlations, we further design a cross-modal attention module as the information bridge between different modalities to perform cross-modal interactions and transfer complementary information. Our extensive experiments on five public TCGA datasets demonstrate that our proposed framework outperforms the state-of-the-art methods.
Multiple Instance Learning Framework with Masked Hard Instance Mining for Whole Slide Image Classification
Jul 31, 2023Wenhao Tang, Sheng Huang, Xiaoxian Zhang, Fengtao Zhou, Yi Zhang, Bo Liu




The whole slide image (WSI) classification is often formulated as a multiple instance learning (MIL) problem. Since the positive tissue is only a small fraction of the gigapixel WSI, existing MIL methods intuitively focus on identifying salient instances via attention mechanisms. However, this leads to a bias towards easy-to-classify instances while neglecting hard-to-classify instances. Some literature has revealed that hard examples are beneficial for modeling a discriminative boundary accurately. By applying such an idea at the instance level, we elaborate a novel MIL framework with masked hard instance mining (MHIM-MIL), which uses a Siamese structure (Teacher-Student) with a consistency constraint to explore the potential hard instances. With several instance masking strategies based on attention scores, MHIM-MIL employs a momentum teacher to implicitly mine hard instances for training the student model, which can be any attention-based MIL model. This counter-intuitive strategy essentially enables the student to learn a better discriminating boundary. Moreover, the student is used to update the teacher with an exponential moving average (EMA), which in turn identifies new hard instances for subsequent training iterations and stabilizes the optimization. Experimental results on the CAMELYON-16 and TCGA Lung Cancer datasets demonstrate that MHIM-MIL outperforms other latest methods in terms of performance and training cost. The code is available at: https://github.com/DearCaat/MHIM-MIL.
Boosting Multi-Label Image Classification with Complementary Parallel Self-Distillation
May 23, 2022Jiazhi Xu, Sheng Huang, Fengtao Zhou, Luwen Huangfu, Daniel Zeng, Bo Liu




Multi-Label Image Classification (MLIC) approaches usually exploit label correlations to achieve good performance. However, emphasizing correlation like co-occurrence may overlook discriminative features of the target itself and lead to model overfitting, thus undermining the performance. In this study, we propose a generic framework named Parallel Self-Distillation (PSD) for boosting MLIC models. PSD decomposes the original MLIC task into several simpler MLIC sub-tasks via two elaborated complementary task decomposition strategies named Co-occurrence Graph Partition (CGP) and Dis-occurrence Graph Partition (DGP). Then, the MLIC models of fewer categories are trained with these sub-tasks in parallel for respectively learning the joint patterns and the category-specific patterns of labels. Finally, knowledge distillation is leveraged to learn a compact global ensemble of full categories with these learned patterns for reconciling the label correlation exploitation and model overfitting. Extensive results on MS-COCO and NUS-WIDE datasets demonstrate that our framework can be easily plugged into many MLIC approaches and improve performances of recent state-of-the-art approaches. The explainable visual study also further validates that our method is able to learn both the category-specific and co-occurring features. The source code is released at https://github.com/Robbie-Xu/CPSD.
Deep Semantic Dictionary Learning for Multi-label Image Classification
Dec 23, 2020Fengtao Zhou, Sheng Huang, Yun Xing

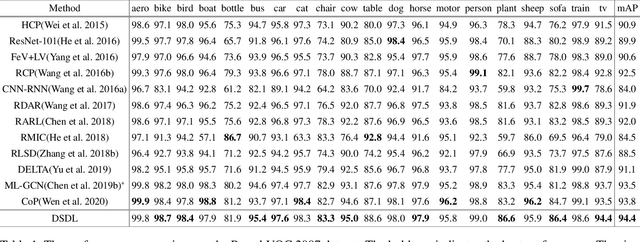

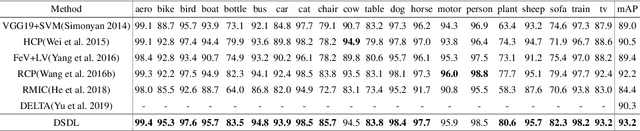
Compared with single-label image classification, multi-label image classification is more practical and challenging. Some recent studies attempted to leverage the semantic information of categories for improving multi-label image classification performance. However, these semantic-based methods only take semantic information as type of complements for visual representation without further exploitation. In this paper, we present a innovative path towards the solution of the multi-label image classification which considers it as a dictionary learning task. A novel end-to-end model named Deep Semantic Dictionary Learning (DSDL) is designed. In DSDL, an auto-encoder is applied to generate the semantic dictionary from class-level semantics and then such dictionary is utilized for representing the visual features extracted by Convolutional Neural Network (CNN) with label embeddings. The DSDL provides a simple but elegant way to exploit and reconcile the label, semantic and visual spaces simultaneously via conducting the dictionary learning among them. Moreover, inspired by iterative optimization of traditional dictionary learning, we further devise a novel training strategy named Alternately Parameters Update Strategy (APUS) for optimizing DSDL, which alteratively optimizes the representation coefficients and the semantic dictionary in forward and backward propagation. Extensive experimental results on three popular benchmarks demonstrate that our method achieves promising performances in comparison with the state-of-the-arts. Our codes and models are available at https://github.com/ZFT-CQU/DSDL.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge