Learning truly monotone operators with applications to nonlinear inverse problems
Mar 30, 2024Younes Belkouchi, Jean-Christophe Pesquet, Audrey Repetti, Hugues Talbot
This article introduces a novel approach to learning monotone neural networks through a newly defined penalization loss. The proposed method is particularly effective in solving classes of variational problems, specifically monotone inclusion problems, commonly encountered in image processing tasks. The Forward-Backward-Forward (FBF) algorithm is employed to address these problems, offering a solution even when the Lipschitz constant of the neural network is unknown. Notably, the FBF algorithm provides convergence guarantees under the condition that the learned operator is monotone. Building on plug-and-play methodologies, our objective is to apply these newly learned operators to solving non-linear inverse problems. To achieve this, we initially formulate the problem as a variational inclusion problem. Subsequently, we train a monotone neural network to approximate an operator that may not inherently be monotone. Leveraging the FBF algorithm, we then show simulation examples where the non-linear inverse problem is successfully solved.
A foundation for exact binarized morphological neural networks
Jan 08, 2024Theodore Aouad, Hugues Talbot
Training and running deep neural networks (NNs) often demands a lot of computation and energy-intensive specialized hardware (e.g. GPU, TPU...). One way to reduce the computation and power cost is to use binary weight NNs, but these are hard to train because the sign function has a non-smooth gradient. We present a model based on Mathematical Morphology (MM), which can binarize ConvNets without losing performance under certain conditions, but these conditions may not be easy to satisfy in real-world scenarios. To solve this, we propose two new approximation methods and develop a robust theoretical framework for ConvNets binarization using MM. We propose as well regularization losses to improve the optimization. We empirically show that our model can learn a complex morphological network, and explore its performance on a classification task.
On the detection of Out-Of-Distribution samples in Multiple Instance Learning
Sep 11, 2023Loïc Le Bescond, Maria Vakalopoulou, Stergios Christodoulidis, Fabrice André, Hugues Talbot

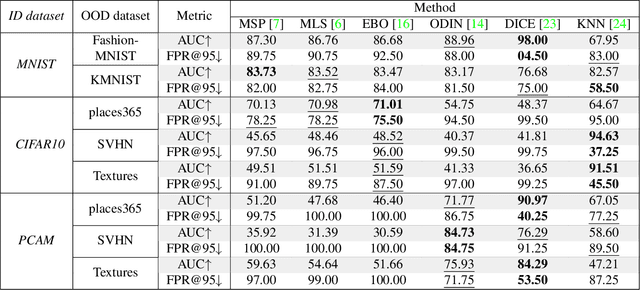
The deployment of machine learning solutions in real-world scenarios often involves addressing the challenge of out-of-distribution (OOD) detection. While significant efforts have been devoted to OOD detection in classical supervised settings, the context of weakly supervised learning, particularly the Multiple Instance Learning (MIL) framework, remains under-explored. In this study, we tackle this challenge by adapting post-hoc OOD detection methods to the MIL setting while introducing a novel benchmark specifically designed to assess OOD detection performance in weakly supervised scenarios. Extensive experiments based on diverse public datasets do not reveal a single method with a clear advantage over the others. Although DICE emerges as the best-performing method overall, it exhibits significant shortcomings on some datasets, emphasizing the complexity of this under-explored and challenging topic. Our findings shed light on the complex nature of OOD detection under the MIL framework, emphasizing the importance of developing novel, robust, and reliable methods that can generalize effectively in a weakly supervised context. The code for the paper is available here: https://github.com/loic-lb/OOD_MIL.
Bone mineral density estimation from a plain X-ray image by learning decomposition into projections of bone-segmented computed tomography
Jul 21, 2023Yi Gu, Yoshito Otake, Keisuke Uemura, Mazen Soufi, Masaki Takao, Hugues Talbot, Seiji Okada, Nobuhiko Sugano, Yoshinobu Sato
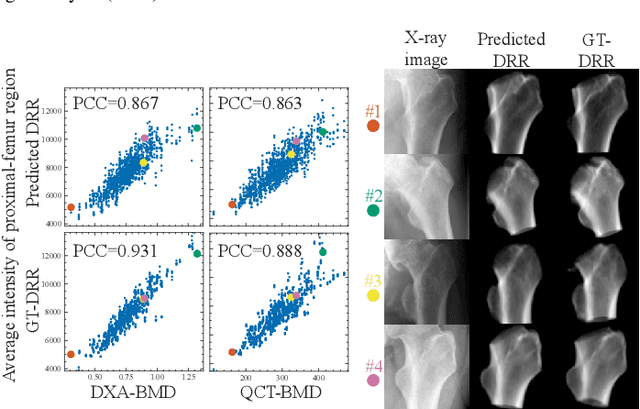
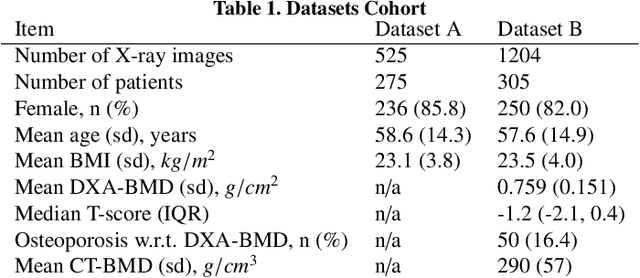
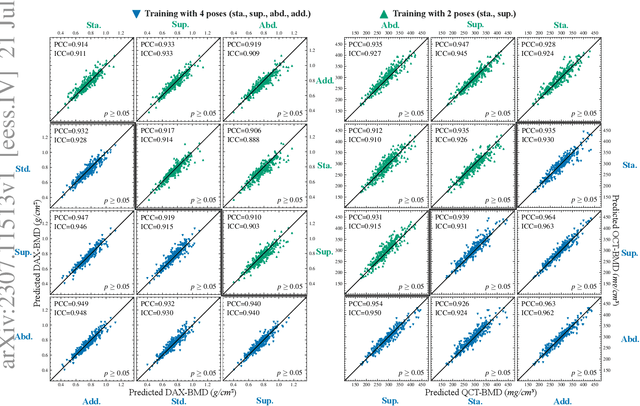
Osteoporosis is a prevalent bone disease that causes fractures in fragile bones, leading to a decline in daily living activities. Dual-energy X-ray absorptiometry (DXA) and quantitative computed tomography (QCT) are highly accurate for diagnosing osteoporosis; however, these modalities require special equipment and scan protocols. To frequently monitor bone health, low-cost, low-dose, and ubiquitously available diagnostic methods are highly anticipated. In this study, we aim to perform bone mineral density (BMD) estimation from a plain X-ray image for opportunistic screening, which is potentially useful for early diagnosis. Existing methods have used multi-stage approaches consisting of extraction of the region of interest and simple regression to estimate BMD, which require a large amount of training data. Therefore, we propose an efficient method that learns decomposition into projections of bone-segmented QCT for BMD estimation under limited datasets. The proposed method achieved high accuracy in BMD estimation, where Pearson correlation coefficients of 0.880 and 0.920 were observed for DXA-measured BMD and QCT-measured BMD estimation tasks, respectively, and the root mean square of the coefficient of variation values were 3.27 to 3.79% for four measurements with different poses. Furthermore, we conducted extensive validation experiments, including multi-pose, uncalibrated-CT, and compression experiments toward actual application in routine clinical practice.
MSKdeX: Musculoskeletal (MSK) decomposition from an X-ray image for fine-grained estimation of lean muscle mass and muscle volume
May 31, 2023Yi Gu, Yoshito Otake, Keisuke Uemura, Masaki Takao, Mazen Soufi, Yuta Hiasa, Hugues Talbot, Seiji Okata, Nobuhiko Sugano, Yoshinobu Sato
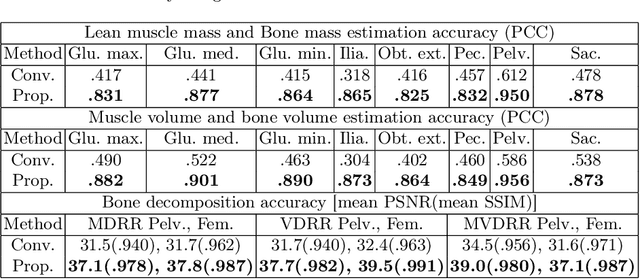
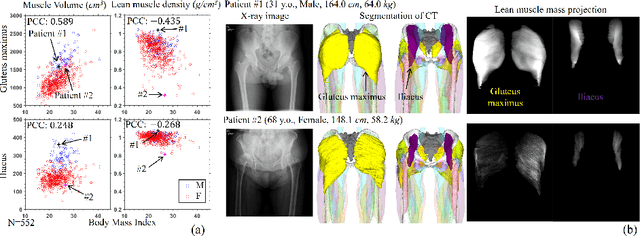

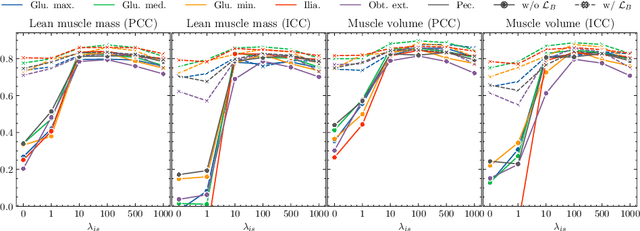
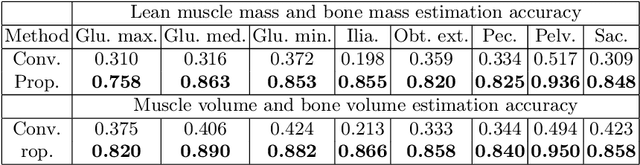
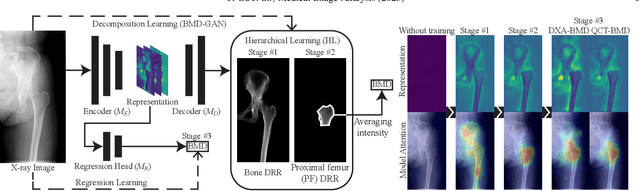
Musculoskeletal diseases such as sarcopenia and osteoporosis are major obstacles to health during aging. Although dual-energy X-ray absorptiometry (DXA) and computed tomography (CT) can be used to evaluate musculoskeletal conditions, frequent monitoring is difficult due to the cost and accessibility (as well as high radiation exposure in the case of CT). We propose a method (named MSKdeX) to estimate fine-grained muscle properties from a plain X-ray image, a low-cost, low-radiation, and highly accessible imaging modality, through musculoskeletal decomposition leveraging fine-grained segmentation in CT. We train a multi-channel quantitative image translation model to decompose an X-ray image into projections of CT of individual muscles to infer the lean muscle mass and muscle volume. We propose the object-wise intensity-sum loss, a simple yet surprisingly effective metric invariant to muscle deformation and projection direction, utilizing information in CT and X-ray images collected from the same patient. While our method is basically an unpaired image-to-image translation, we also exploit the nature of the bone's rigidity, which provides the paired data through 2D-3D rigid registration, adding strong pixel-wise supervision in unpaired training. Through the evaluation using a 539-patient dataset, we showed that the proposed method significantly outperformed conventional methods. The average Pearson correlation coefficient between the predicted and CT-derived ground truth metrics was increased from 0.460 to 0.863. We believe our method opened up a new musculoskeletal diagnosis method and has the potential to be extended to broader applications in multi-channel quantitative image translation tasks. Our source code will be released soon.
Binary Multi Channel Morphological Neural Network
Apr 19, 2022Theodore Aouad, Hugues Talbot




Neural networks and particularly Deep learning have been comparatively little studied from the theoretical point of view. Conversely, Mathematical Morphology is a discipline with solid theoretical foundations. We combine these domains to propose a new type of neural architecture that is theoretically more explainable. We introduce a Binary Morphological Neural Network (BiMoNN) built upon the convolutional neural network. We design it for learning morphological networks with binary inputs and outputs. We demonstrate an equivalence between BiMoNNs and morphological operators that we can use to binarize entire networks. These can learn classical morphological operators and show promising results on a medical imaging application.
Binary Morphological Neural Network
Mar 23, 2022Theodore Aouad, Hugues Talbot




In the last ten years, Convolutional Neural Networks (CNNs) have formed the basis of deep-learning architectures for most computer vision tasks. However, they are not necessarily optimal. For example, mathematical morphology is known to be better suited to deal with binary images. In this work, we create a morphological neural network that handles binary inputs and outputs. We propose their construction inspired by CNNs to formulate layers adapted to such images by replacing convolutions with erosions and dilations. We give explainable theoretical results on whether or not the resulting learned networks are indeed morphological operators. We present promising experimental results designed to learn basic binary operators, and we have made our code publicly available online.
Geo-Spatiotemporal Features and Shape-Based Prior Knowledge for Fine-grained Imbalanced Data Classification
Mar 21, 2021Charles A. Kantor, Marta Skreta, Brice Rauby, Léonard Boussioux, Emmanuel Jehanno, Alexandra Luccioni, David Rolnick, Hugues Talbot
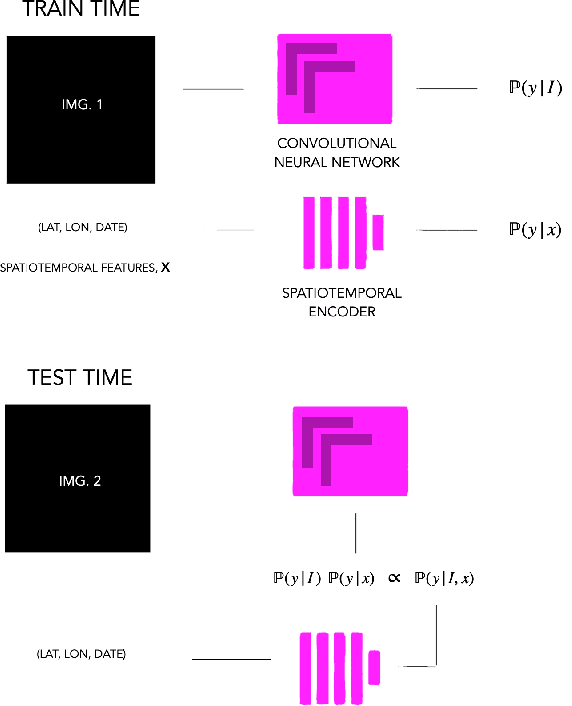
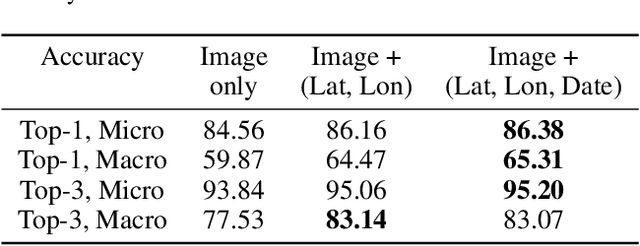

Fine-grained classification aims at distinguishing between items with similar global perception and patterns, but that differ by minute details. Our primary challenges come from both small inter-class variations and large intra-class variations. In this article, we propose to combine several innovations to improve fine-grained classification within the use-case of wildlife, which is of practical interest for experts. We utilize geo-spatiotemporal data to enrich the picture information and further improve the performance. We also investigate state-of-the-art methods for handling the imbalanced data issue.
* Copyright by the authors. All rights reserved to authors only. Correspondence to: ckantor (at) stanford [dot] edu
Atlas-based automated detection of swim bladder in Medaka embryo
Feb 16, 2019Diane Genest, Marc Léonard, Jean Cousty, Noémie De Crozé, Hugues Talbot




Fish embryo models are increasingly being used both for the assessment of chemicals efficacy and potential toxicity. This article proposes a methodology to automatically detect the swim bladder on 2D images of Medaka fish embryos seen either in dorsal view or in lateral view. After embryo segmentation and for each studied orientation, the method builds an atlas of a healthy embryo. This atlas is then used to define the region of interest and to guide the swim bladder segmentation with a discrete globally optimal active contour. Descriptors are subsequently designed from this segmentation. An automated random forest clas-sifier is built from these descriptors in order to classify embryos with and without a swim bladder. The proposed method is assessed on a dataset of 261 images, containing 202 embryos with a swim bladder (where 196 are in dorsal view and 6 are in lateral view) and 59 without (where 43 are in dorsal view and 16 are in lateral view). We obtain an average precision rate of 95% in the total dataset following 5-fold cross-validation.
Spatially variant PSF modeling in confocal macroscopy
Jul 14, 2017Anna Jezierska, Hugues Talbot, Jean-Christophe Pesquet, Gilbert Engler




Point spread function (PSF) plays an essential role in image reconstruction. In the context of confocal microscopy, optical performance degrades towards the edge of the field of view as astigmatism, coma and vignetting. Thus, one should expect the related artifacts to be even stronger in macroscopy, where the field of view is much larger. The field aberrations in macroscopy fluorescence imaging system was observed to be symmetrical and to increase with the distance from the center of the field of view. In this paper we propose an experiment and an optimization method for assessing the center of the field of view. The obtained results constitute a step towards reducing the number of parameters in macroscopy PSF model.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge