Automatic hip osteoarthritis grading with uncertainty estimation from computed tomography using digitally-reconstructed radiographs
Dec 30, 2023Masachika Masuda, Mazen Soufi, Yoshito Otake, Keisuke Uemura, Sotaro Kono, Kazuma Takashima, Hidetoshi Hamada, Yi Gu, Masaki Takao, Seiji Okada, Nobuhiko Sugano, Yoshinobu Sato
Progression of hip osteoarthritis (hip OA) leads to pain and disability, likely leading to surgical treatment such as hip arthroplasty at the terminal stage. The severity of hip OA is often classified using the Crowe and Kellgren-Lawrence (KL) classifications. However, as the classification is subjective, we aimed to develop an automated approach to classify the disease severity based on the two grades using digitally-reconstructed radiographs (DRRs) from CT images. Automatic grading of the hip OA severity was performed using deep learning-based models. The models were trained to predict the disease grade using two grading schemes, i.e., predicting the Crowe and KL grades separately, and predicting a new ordinal label combining both grades and representing the disease progression of hip OA. The models were trained in classification and regression settings. In addition, the model uncertainty was estimated and validated as a predictor of classification accuracy. The models were trained and validated on a database of 197 hip OA patients, and externally validated on 52 patients. The model accuracy was evaluated using exact class accuracy (ECA), one-neighbor class accuracy (ONCA), and balanced accuracy.The deep learning models produced a comparable accuracy of approximately 0.65 (ECA) and 0.95 (ONCA) in the classification and regression settings. The model uncertainty was significantly larger in cases with large classification errors (P<6e-3). In this study, an automatic approach for grading hip OA severity from CT images was developed. The models have shown comparable performance with high ONCA, which facilitates automated grading in large-scale CT databases and indicates the potential for further disease progression analysis. Classification accuracy was correlated with the model uncertainty, which would allow for the prediction of classification errors.
Hybrid Representation-Enhanced Sampling for Bayesian Active Learning in Musculoskeletal Segmentation of Lower Extremities
Jul 26, 2023Ganping Li, Yoshito Otake, Mazen Soufi, Masashi Taniguchi, Masahide Yagi, Noriaki Ichihashi, Keisuke Uemura, Masaki Takao, Nobuhiko Sugano, Yoshinobu Sato




Purpose: Obtaining manual annotations to train deep learning (DL) models for auto-segmentation is often time-consuming. Uncertainty-based Bayesian active learning (BAL) is a widely-adopted method to reduce annotation efforts. Based on BAL, this study introduces a hybrid representation-enhanced sampling strategy that integrates density and diversity criteria to save manual annotation costs by efficiently selecting the most informative samples. Methods: The experiments are performed on two lower extremity (LE) datasets of MRI and CT images by a BAL framework based on Bayesian U-net. Our method selects uncertain samples with high density and diversity for manual revision, optimizing for maximal similarity to unlabeled instances and minimal similarity to existing training data. We assess the accuracy and efficiency using Dice and a proposed metric called reduced annotation cost (RAC), respectively. We further evaluate the impact of various acquisition rules on BAL performance and design an ablation study for effectiveness estimation. Results: The proposed method showed superiority or non-inferiority to other methods on both datasets across two acquisition rules, and quantitative results reveal the pros and cons of the acquisition rules. Our ablation study in volume-wise acquisition shows that the combination of density and diversity criteria outperforms solely using either of them in musculoskeletal segmentation. Conclusion: Our sampling method is proven efficient in reducing annotation costs in image segmentation tasks. The combination of the proposed method and our BAL framework provides a semi-automatic way for efficient annotation of medical image datasets.
Bone mineral density estimation from a plain X-ray image by learning decomposition into projections of bone-segmented computed tomography
Jul 21, 2023Yi Gu, Yoshito Otake, Keisuke Uemura, Mazen Soufi, Masaki Takao, Hugues Talbot, Seiji Okada, Nobuhiko Sugano, Yoshinobu Sato
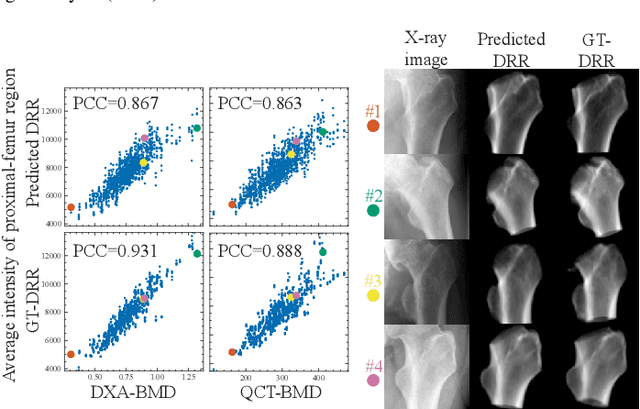
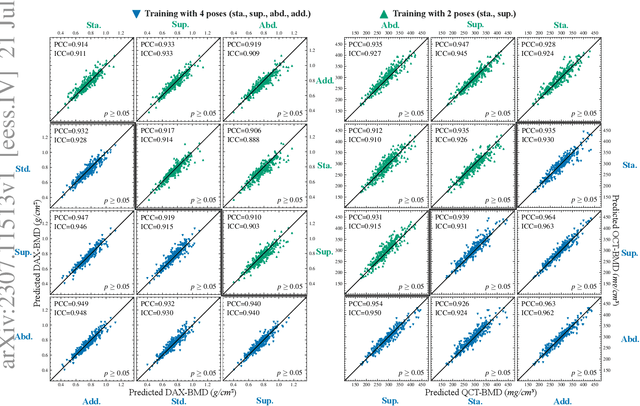
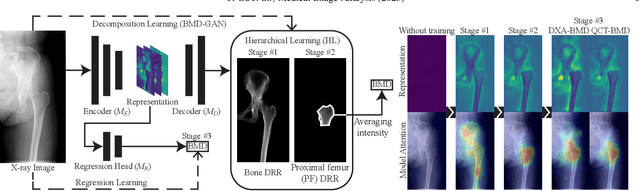
Osteoporosis is a prevalent bone disease that causes fractures in fragile bones, leading to a decline in daily living activities. Dual-energy X-ray absorptiometry (DXA) and quantitative computed tomography (QCT) are highly accurate for diagnosing osteoporosis; however, these modalities require special equipment and scan protocols. To frequently monitor bone health, low-cost, low-dose, and ubiquitously available diagnostic methods are highly anticipated. In this study, we aim to perform bone mineral density (BMD) estimation from a plain X-ray image for opportunistic screening, which is potentially useful for early diagnosis. Existing methods have used multi-stage approaches consisting of extraction of the region of interest and simple regression to estimate BMD, which require a large amount of training data. Therefore, we propose an efficient method that learns decomposition into projections of bone-segmented QCT for BMD estimation under limited datasets. The proposed method achieved high accuracy in BMD estimation, where Pearson correlation coefficients of 0.880 and 0.920 were observed for DXA-measured BMD and QCT-measured BMD estimation tasks, respectively, and the root mean square of the coefficient of variation values were 3.27 to 3.79% for four measurements with different poses. Furthermore, we conducted extensive validation experiments, including multi-pose, uncalibrated-CT, and compression experiments toward actual application in routine clinical practice.
MSKdeX: Musculoskeletal (MSK) decomposition from an X-ray image for fine-grained estimation of lean muscle mass and muscle volume
May 31, 2023Yi Gu, Yoshito Otake, Keisuke Uemura, Masaki Takao, Mazen Soufi, Yuta Hiasa, Hugues Talbot, Seiji Okata, Nobuhiko Sugano, Yoshinobu Sato
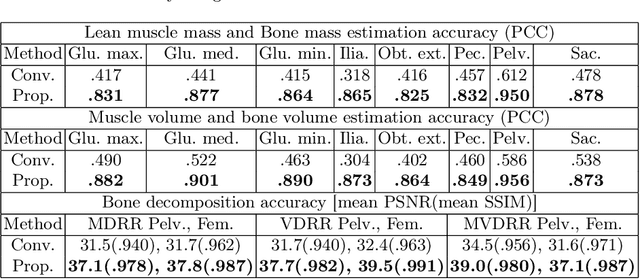
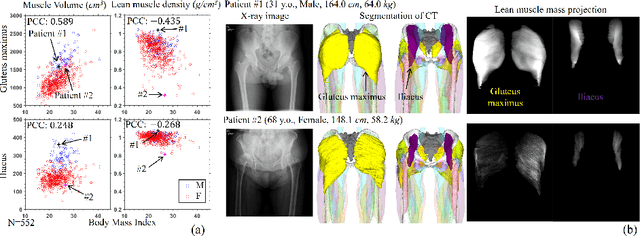

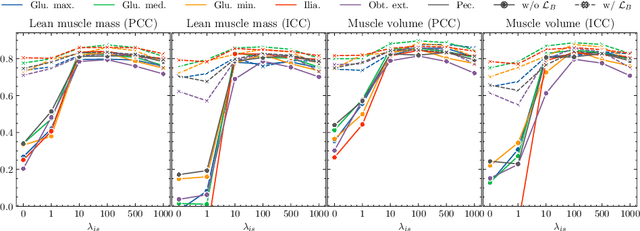
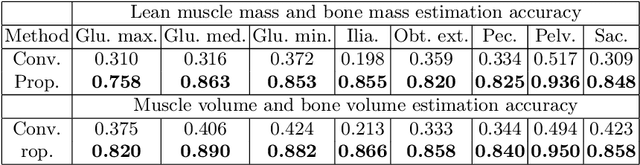
Musculoskeletal diseases such as sarcopenia and osteoporosis are major obstacles to health during aging. Although dual-energy X-ray absorptiometry (DXA) and computed tomography (CT) can be used to evaluate musculoskeletal conditions, frequent monitoring is difficult due to the cost and accessibility (as well as high radiation exposure in the case of CT). We propose a method (named MSKdeX) to estimate fine-grained muscle properties from a plain X-ray image, a low-cost, low-radiation, and highly accessible imaging modality, through musculoskeletal decomposition leveraging fine-grained segmentation in CT. We train a multi-channel quantitative image translation model to decompose an X-ray image into projections of CT of individual muscles to infer the lean muscle mass and muscle volume. We propose the object-wise intensity-sum loss, a simple yet surprisingly effective metric invariant to muscle deformation and projection direction, utilizing information in CT and X-ray images collected from the same patient. While our method is basically an unpaired image-to-image translation, we also exploit the nature of the bone's rigidity, which provides the paired data through 2D-3D rigid registration, adding strong pixel-wise supervision in unpaired training. Through the evaluation using a 539-patient dataset, we showed that the proposed method significantly outperformed conventional methods. The average Pearson correlation coefficient between the predicted and CT-derived ground truth metrics was increased from 0.460 to 0.863. We believe our method opened up a new musculoskeletal diagnosis method and has the potential to be extended to broader applications in multi-channel quantitative image translation tasks. Our source code will be released soon.
BMD-GAN: Bone mineral density estimation using x-ray image decomposition into projections of bone-segmented quantitative computed tomography using hierarchical learning
Jul 07, 2022Yi Gu, Yoshito Otake, Keisuke Uemura, Mazen Soufi, Masaki Takao, Nobuhiko Sugano, Yoshinobu Sato




We propose a method for estimating the bone mineral density (BMD) from a plain x-ray image. Dual-energy X-ray absorptiometry (DXA) and quantitative computed tomography (QCT) provide high accuracy in diagnosing osteoporosis; however, these modalities require special equipment and scan protocols. Measuring BMD from an x-ray image provides an opportunistic screening, which is potentially useful for early diagnosis. The previous methods that directly learn the relationship between x-ray images and BMD require a large training dataset to achieve high accuracy because of large intensity variations in the x-ray images. Therefore, we propose an approach using the QCT for training a generative adversarial network (GAN) and decomposing an x-ray image into a projection of bone-segmented QCT. The proposed hierarchical learning improved the robustness and accuracy of quantitatively decomposing a small-area target. The evaluation of 200 patients with osteoarthritis using the proposed method, which we named BMD-GAN, demonstrated a Pearson correlation coefficient of 0.888 between the predicted and ground truth DXA-measured BMD. Besides not requiring a large-scale training database, another advantage of our method is its extensibility to other anatomical areas, such as the vertebrae and rib bones.
Automated segmentation of an intensity calibration phantom in clinical CT images using a convolutional neural network
Dec 21, 2020Keisuke Uemura, Yoshito Otake, Masaki Takao, Mazen Soufi, Akihiro Kawasaki, Nobuhiko Sugano, Yoshinobu Sato




Purpose: To apply a convolutional neural network (CNN) to develop a system that segments intensity calibration phantom regions in computed tomography (CT) images, and to test the system in a large cohort to evaluate its robustness. Methods: A total of 1040 cases (520 cases each from two institutions), in which an intensity calibration phantom (B-MAS200, Kyoto Kagaku, Kyoto, Japan) was used, were included herein. A training dataset was created by manually segmenting the regions of the phantom for 40 cases (20 cases each). Segmentation accuracy of the CNN model was assessed with the Dice coefficient and the average symmetric surface distance (ASD) through the 4-fold cross validation. Further, absolute differences of radiodensity values (in Hounsfield units: HU) were compared between manually segmented regions and automatically segmented regions. The system was tested on the remaining 1000 cases. For each institution, linear regression was applied to calculate coefficients for the correlation between radiodensity and the densities of the phantom. Results: After training, the median Dice coefficient was 0.977, and the median ASD was 0.116 mm. When segmented regions were compared between manual segmentation and automated segmentation, the median absolute difference was 0.114 HU. For the test cases, the median correlation coefficient was 0.9998 for one institution and was 0.9999 for the other, with a minimum value of 0.9863. Conclusions: The CNN model successfully segmented the calibration phantom's regions in the CT images with excellent accuracy, and the automated method was found to be at least equivalent to the conventional manual method. Future study should integrate the system by automatically segmenting the region of interest in bones such that the bone mineral density can be fully automatically quantified from CT images.
Region-based Convolution Neural Network Approach for Accurate Segmentation of Pelvic Radiograph
Oct 29, 2019Ata Jodeiri, Reza A. Zoroofi, Yuta Hiasa, Masaki Takao, Nobuhiko Sugano, Yoshinobu Sato, Yoshito Otake




With the increasing usage of radiograph images as a most common medical imaging system for diagnosis, treatment planning, and clinical studies, it is increasingly becoming a vital factor to use machine learning-based systems to provide reliable information for surgical pre-planning. Segmentation of pelvic bone in radiograph images is a critical preprocessing step for some applications such as automatic pose estimation and disease detection. However, the encoder-decoder style network known as U-Net has demonstrated limited results due to the challenging complexity of the pelvic shapes, especially in severe patients. In this paper, we propose a novel multi-task segmentation method based on Mask R-CNN architecture. For training, the network weights were initialized by large non-medical dataset and fine-tuned with radiograph images. Furthermore, in the training process, augmented data was generated to improve network performance. Our experiments show that Mask R-CNN utilizing multi-task learning, transfer learning, and data augmentation techniques achieve 0.96 DICE coefficient, which significantly outperforms the U-Net. Notably, for a fair comparison, the same transfer learning and data augmentation techniques have been used for U-net training.
Estimation of Pelvic Sagittal Inclination from Anteroposterior Radiograph Using Convolutional Neural Networks: Proof-of-Concept Study
Oct 26, 2019Ata Jodeiri, Yoshito Otake, Reza A. Zoroofi, Yuta Hiasa, Masaki Takao, Keisuke Uemura, Nobuhiko Sugano, Yoshinobu Sato


Alignment of the bones in standing position provides useful information in surgical planning. In total hip arthroplasty (THA), pelvic sagittal inclination (PSI) angle in the standing position is an important factor in planning of cup alignment and has been estimated mainly from radiographs. Previous methods for PSI estimation used a patient-specific CT to create digitally reconstructed radiographs (DRRs) and compare them with the radiograph to estimate relative position between the pelvis and the x-ray detector. In this study, we developed a method that estimates PSI angle from a single anteroposterior radiograph using two convolutional neural networks (CNNs) without requiring the patient-specific CT, which reduces radiation exposure of the patient and opens up the possibility of application in a larger number of hospitals where CT is not acquired in a routine protocol.
Automated Muscle Segmentation from Clinical CT using Bayesian U-Net for Personalization of a Musculoskeletal Model
Jul 21, 2019Yuta Hiasa, Yoshito Otake, Masaki Takao, Takeshi Ogawa, Nobuhiko Sugano, Yoshinobu Sato




We propose a method for automatic segmentation of individual muscles from a clinical CT. The method uses Bayesian convolutional neural networks with the U-Net architecture, using Monte Carlo dropout that infers an uncertainty metric in addition to the segmentation label. We evaluated the performance of the proposed method using two data sets: 20 fully annotated CTs of the hip and thigh regions and 18 partially annotated CTs that are publicly available from The Cancer Imaging Archive (TCIA) database. The experiments showed a Dice coefficient (DC) of 0.891 +/- 0.016 (mean +/- std) and an average symmetric surface distance (ASD) of 0.994 +/- 0.230 mm over 19 muscles in the set of 20 CTs. These results were statistically significant improvements compared to the state-of-the-art hierarchical multi-atlas method which resulted in 0.845 +/- 0.031 DC and 1.556 +/- 0.444 mm ASD. We evaluated validity of the uncertainty metric in the multi-class organ segmentation problem and demonstrated a correlation between the pixels with high uncertainty and the segmentation failure. One application of the uncertainty metric in active-learning is demonstrated, and the proposed query pixel selection method considerably reduced the manual annotation cost for expanding the training data set. The proposed method allows an accurate patient-specific analysis of individual muscle shapes in a clinical routine. This would open up various applications including personalization of biomechanical simulation and quantitative evaluation of muscle atrophy.
Automated Segmentation of Hip and Thigh Muscles in Metal Artifact-Contaminated CT using Convolutional Neural Network-Enhanced Normalized Metal Artifact Reduction
Jun 27, 2019Mitsuki Sakamoto, Yuta Hiasa, Yoshito Otake, Masaki Takao, Yuki Suzuki, Nobuhiko Sugano, Yoshinobu Sato




In total hip arthroplasty, analysis of postoperative medical images is important to evaluate surgical outcome. Since Computed Tomography (CT) is most prevalent modality in orthopedic surgery, we aimed at the analysis of CT image. In this work, we focus on the metal artifact in postoperative CT caused by the metallic implant, which reduces the accuracy of segmentation especially in the vicinity of the implant. Our goal was to develop an automated segmentation method of the bones and muscles in the postoperative CT images. We propose a method that combines Normalized Metal Artifact Reduction (NMAR), which is one of the state-of-the-art metal artifact reduction methods, and a Convolutional Neural Network-based segmentation using two U-net architectures. The first U-net refines the result of NMAR and the muscle segmentation is performed by the second U-net. We conducted experiments using simulated images of 20 patients and real images of three patients to evaluate the segmentation accuracy of 19 muscles. In simulation study, the proposed method showed statistically significant improvement (p<0.05) in the average symmetric surface distance (ASD) metric for 14 muscles out of 19 muscles and the average ASD of all muscles from 1.17 +/- 0.543 mm (mean +/- std over all patients) to 1.10 +/- 0.509 mm over our previous method. The real image study using the manual trace of gluteus maximus and medius muscles showed ASD of 1.32 +/- 0.25 mm. Our future work includes training of a network in an end-to-end manner for both the metal artifact reduction and muscle segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge