Lamarckian Inheritance Improves Robot Evolution in Dynamic Environments
Mar 28, 2024Jie Luo, Karine Miras, Carlo Longhi, Oliver Weissl, Agoston E. Eiben

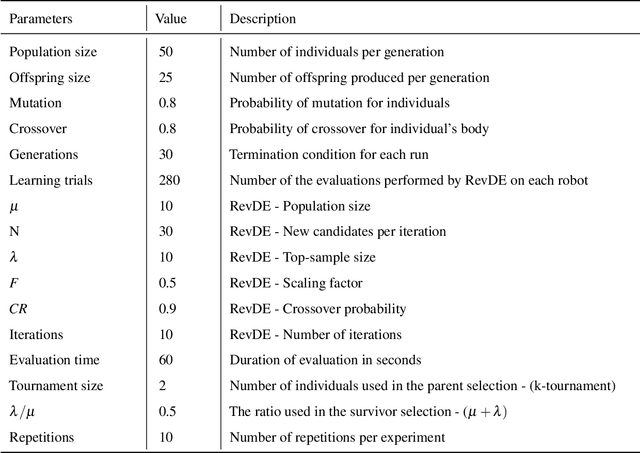
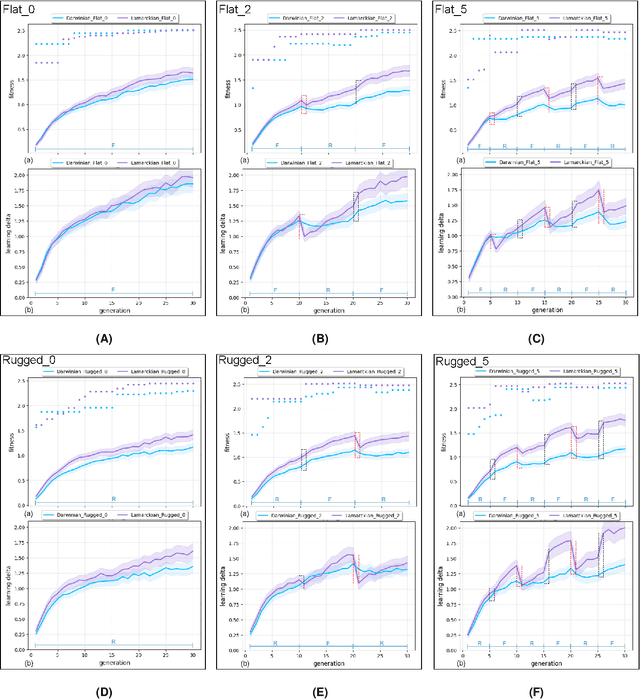
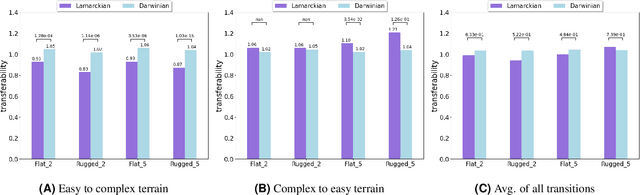
This study explores the integration of Lamarckian system into evolutionary robotics (ER), comparing it with the traditional Darwinian model across various environments. By adopting Lamarckian principles, where robots inherit learned traits, alongside Darwinian learning without inheritance, we investigate adaptation in dynamic settings. Our research, conducted in six distinct environmental setups, demonstrates that Lamarckian systems outperform Darwinian ones in adaptability and efficiency, particularly in challenging conditions. Our analysis highlights the critical role of the interplay between controller \& morphological evolution and environment adaptation, with parent-offspring similarities and newborn \&survivors before and after learning providing insights into the effectiveness of trait inheritance. Our findings suggest Lamarckian principles could significantly advance autonomous system design, highlighting the potential for more adaptable and robust robotic solutions in complex, real-world applications. These theoretical insights were validated using real physical robots, bridging the gap between simulation and practical application.
A Fourier Transform Framework for Domain Adaptation
Mar 21, 2024Le Luo, Bingrong Xu, Qingyong Zhang, Cheng Lian, Jie Luo
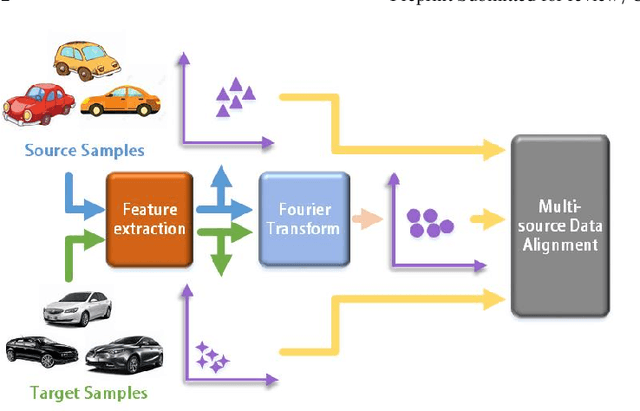
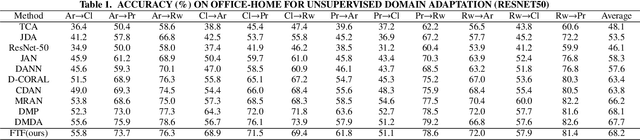
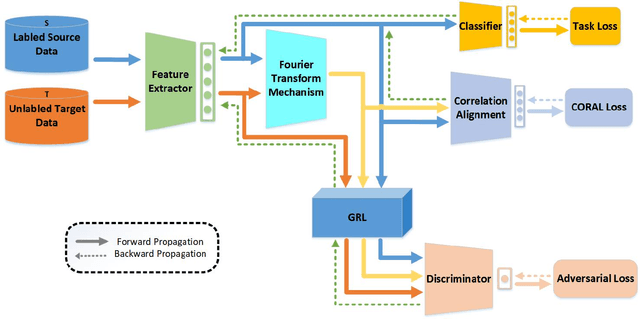
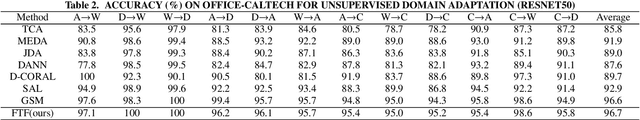
By using unsupervised domain adaptation (UDA), knowledge can be transferred from a label-rich source domain to a target domain that contains relevant information but lacks labels. Many existing UDA algorithms suffer from directly using raw images as input, resulting in models that overly focus on redundant information and exhibit poor generalization capability. To address this issue, we attempt to improve the performance of unsupervised domain adaptation by employing the Fourier method (FTF).Specifically, FTF is inspired by the amplitude of Fourier spectra, which primarily preserves low-level statistical information. In FTF, we effectively incorporate low-level information from the target domain into the source domain by fusing the amplitudes of both domains in the Fourier domain. Additionally, we observe that extracting features from batches of images can eliminate redundant information while retaining class-specific features relevant to the task. Building upon this observation, we apply the Fourier Transform at the data stream level for the first time. To further align multiple sources of data, we introduce the concept of correlation alignment. To evaluate the effectiveness of our FTF method, we conducted evaluations on four benchmark datasets for domain adaptation, including Office-31, Office-Home, ImageCLEF-DA, and Office-Caltech. Our results demonstrate superior performance.
TinyLLaVA: A Framework of Small-scale Large Multimodal Models
Feb 22, 2024Baichuan Zhou, Ying Hu, Xi Weng, Junlong Jia, Jie Luo, Xien Liu, Ji Wu, Lei Huang
We present the TinyLLaVA framework that provides a unified perspective in designing and analyzing the small-scale Large Multimodal Models (LMMs). We empirically study the effects of different vision encoders, connection modules, language models, training data and training recipes. Our extensive experiments showed that better quality of data combined with better training recipes, smaller LMMs can consistently achieve on-par performances compared to bigger LMMs. Under our framework, we train a family of small-scale LMMs. Our best model, TinyLLaVA-3.1B, achieves better overall performance against existing 7B models such as LLaVA-1.5 and Qwen-VL. We hope our findings can serve as baselines for future research in terms of data scaling, training setups and model selections. Our model weights and codes will be made public.
Accurate LoRA-Finetuning Quantization of LLMs via Information Retention
Feb 08, 2024Haotong Qin, Xudong Ma, Xingyu Zheng, Xiaoyang Li, Yang Zhang, Shouda Liu, Jie Luo, Xianglong Liu, Michele Magno
The LoRA-finetuning quantization of LLMs has been extensively studied to obtain accurate yet compact LLMs for deployment on resource-constrained hardware. However, existing methods cause the quantized LLM to severely degrade and even fail to benefit from the finetuning of LoRA. This paper proposes a novel IR-QLoRA for pushing quantized LLMs with LoRA to be highly accurate through information retention. The proposed IR-QLoRA mainly relies on two technologies derived from the perspective of unified information: (1) statistics-based Information Calibration Quantization allows the quantized parameters of LLM to retain original information accurately; (2) finetuning-based Information Elastic Connection makes LoRA utilizes elastic representation transformation with diverse information. Comprehensive experiments show that IR-QLoRA can significantly improve accuracy across LLaMA and LLaMA2 families under 2-4 bit-widths, e.g., 4- bit LLaMA-7B achieves 1.4% improvement on MMLU compared with the state-of-the-art methods. The significant performance gain requires only a tiny 0.31% additional time consumption, revealing the satisfactory efficiency of our IRQLoRA. We highlight that IR-QLoRA enjoys excellent versatility, compatible with various frameworks (e.g., NormalFloat and Integer quantization) and brings general accuracy gains. The code is available at https://github.com/htqin/ir-qlora.
Unsupervised Generation of Pseudo Normal PET from MRI with Diffusion Model for Epileptic Focus Localization
Feb 02, 2024Wentao Chen, Jiwei Li, Xichen Xu, Hui Huang, Siyu Yuan, Miao Zhang, Tianming Xu, Jie Luo, Weimin Zhou
[$^{18}$F]fluorodeoxyglucose (FDG) positron emission tomography (PET) has emerged as a crucial tool in identifying the epileptic focus, especially in cases where magnetic resonance imaging (MRI) diagnosis yields indeterminate results. FDG PET can provide the metabolic information of glucose and help identify abnormal areas that are not easily found through MRI. However, the effectiveness of FDG PET-based assessment and diagnosis depends on the selection of a healthy control group. The healthy control group typically consists of healthy individuals similar to epilepsy patients in terms of age, gender, and other aspects for providing normal FDG PET data, which will be used as a reference for enhancing the accuracy and reliability of the epilepsy diagnosis. However, significant challenges arise when a healthy PET control group is unattainable. Yaakub \emph{et al.} have previously introduced a Pix2PixGAN-based method for MRI to PET translation. This method used paired MRI and FDG PET scans from healthy individuals for training, and produced pseudo normal FDG PET images from patient MRIs that are subsequently used for lesion detection. However, this approach requires a large amount of high-quality, paired MRI and PET images from healthy control subjects, which may not always be available. In this study, we investigated unsupervised learning methods for unpaired MRI to PET translation for generating pseudo normal FDG PET for epileptic focus localization. Two deep learning methods, CycleGAN and SynDiff, were employed, and we found that diffusion-based method achieved improved performance in accurately localizing the epileptic focus.
Ambient-Pix2PixGAN for Translating Medical Images from Noisy Data
Feb 02, 2024Wentao Chen, Xichen Xu, Jie Luo, Weimin Zhou
Image-to-image translation is a common task in computer vision and has been rapidly increasing the impact on the field of medical imaging. Deep learning-based methods that employ conditional generative adversarial networks (cGANs), such as Pix2PixGAN, have been extensively explored to perform image-to-image translation tasks. However, when noisy medical image data are considered, such methods cannot be directly applied to produce clean images. Recently, an augmented GAN architecture named AmbientGAN has been proposed that can be trained on noisy measurement data to synthesize high-quality clean medical images. Inspired by AmbientGAN, in this work, we propose a new cGAN architecture, Ambient-Pix2PixGAN, for performing medical image-to-image translation tasks by use of noisy measurement data. Numerical studies that consider MRI-to-PET translation are conducted. Both traditional image quality metrics and task-based image quality metrics are employed to assess the proposed Ambient-Pix2PixGAN. It is demonstrated that our proposed Ambient-Pix2PixGAN can be successfully trained on noisy measurement data to produce high-quality translated images in target imaging modality.
A comparison of controller architectures and learning mechanisms for arbitrary robot morphologies
Sep 25, 2023Jie Luo, Jakub Tomczak, Karine Miras, Agoston E. Eiben


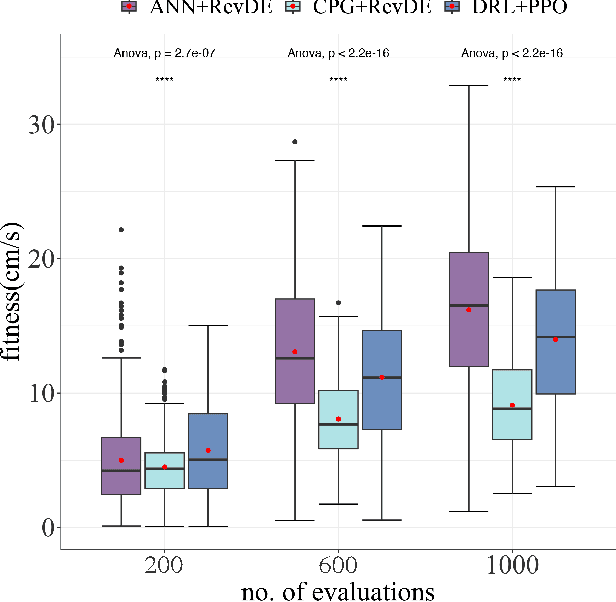
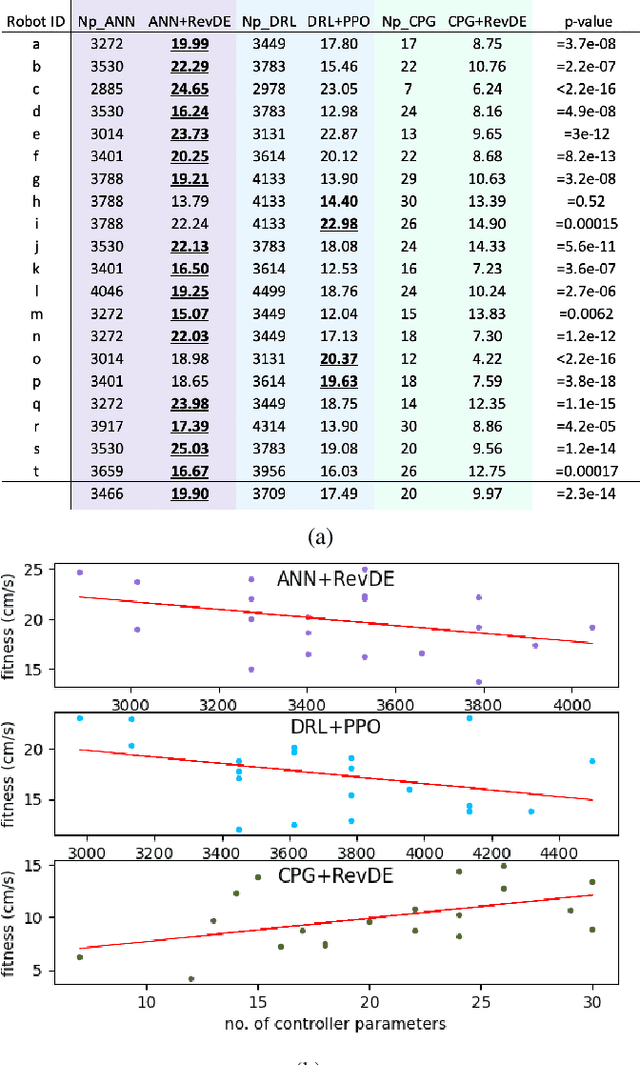
The main question this paper addresses is: What combination of a robot controller and a learning method should be used, if the morphology of the learning robot is not known in advance? Our interest is rooted in the context of morphologically evolving modular robots, but the question is also relevant in general, for system designers interested in widely applicable solutions. We perform an experimental comparison of three controller-and-learner combinations: one approach where controllers are based on modelling animal locomotion (Central Pattern Generators, CPG) and the learner is an evolutionary algorithm, a completely different method using Reinforcement Learning (RL) with a neural network controller architecture, and a combination `in-between' where controllers are neural networks and the learner is an evolutionary algorithm. We apply these three combinations to a test suite of modular robots and compare their efficacy, efficiency, and robustness. Surprisingly, the usual CPG-based and RL-based options are outperformed by the in-between combination that is more robust and efficient than the other two setups.
Exploring Robot Morphology Spaces through Breadth-First Search and Random Query
Sep 25, 2023Jie Luo
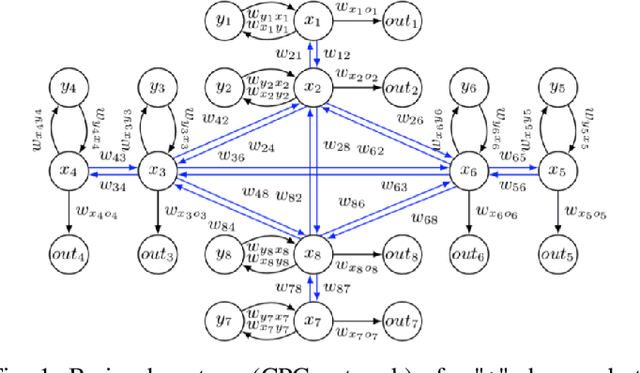
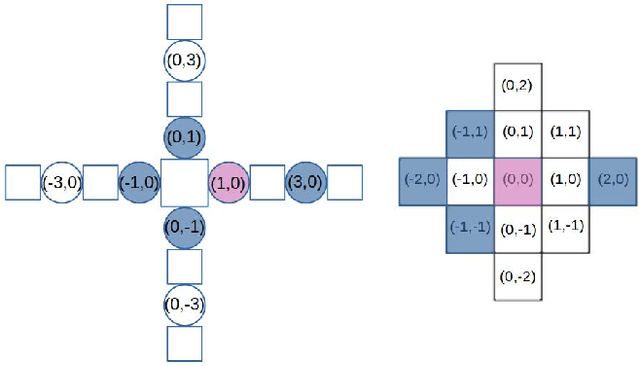

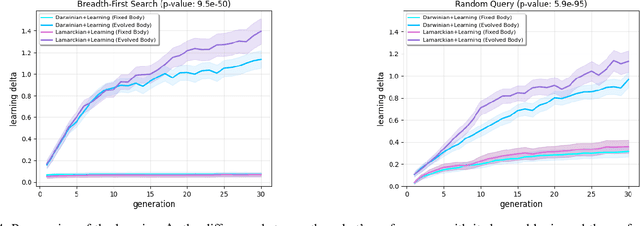
Evolutionary robotics offers a powerful framework for designing and evolving robot morphologies, particularly in the context of modular robots. However, the role of query mechanisms during the genotype-to-phenotype mapping process has been largely overlooked. This research addresses this gap by conducting a comparative analysis of query mechanisms in the brain-body co-evolution of modular robots. Using two different query mechanisms, Breadth-First Search (BFS) and Random Query, within the context of evolving robot morphologies using CPPNs and robot controllers using tensors, and testing them in two evolutionary frameworks, Lamarckian and Darwinian systems, this study investigates their influence on evolutionary outcomes and performance. The findings demonstrate the impact of the two query mechanisms on the evolution and performance of modular robot bodies, including morphological intelligence, diversity, and morphological traits. This study suggests that BFS is both more effective and efficient in producing highly performing robots. It also reveals that initially, robot diversity was higher with BFS compared to Random Query, but in the Lamarckian system, it declines faster, converging to superior designs, while in the Darwinian system, BFS led to higher end-process diversity.
Lamarck's Revenge: Inheritance of Learned Traits Can Make Robot Evolution Better
Sep 22, 2023Jie Luo, Karine Miras, Jakub Tomczak, Agoston E. Eiben
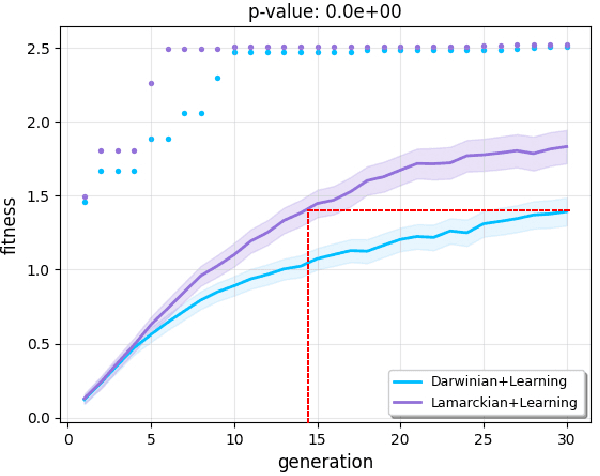
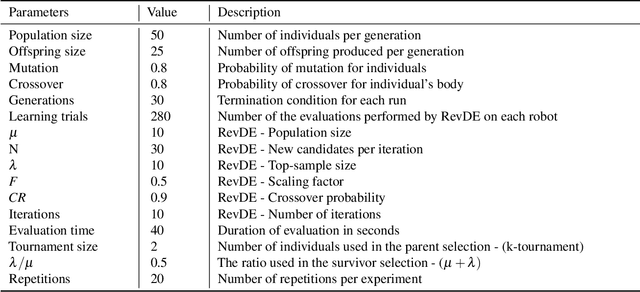
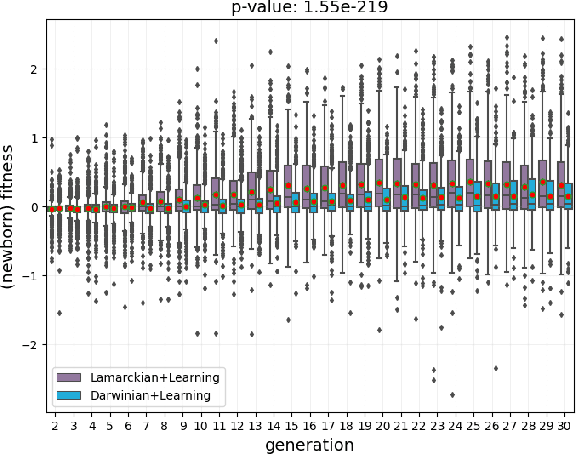

Evolutionary robot systems offer two principal advantages: an advanced way of developing robots through evolutionary optimization and a special research platform to conduct what-if experiments regarding questions about evolution. Our study sits at the intersection of these. We investigate the question ``What if the 18th-century biologist Lamarck was not completely wrong and individual traits learned during a lifetime could be passed on to offspring through inheritance?'' We research this issue through simulations with an evolutionary robot framework where morphologies (bodies) and controllers (brains) of robots are evolvable and robots also can improve their controllers through learning during their lifetime. Within this framework, we compare a Lamarckian system, where learned bits of the brain are inheritable, with a Darwinian system, where they are not. Analyzing simulations based on these systems, we obtain new insights about Lamarckian evolution dynamics and the interaction between evolution and learning. Specifically, we show that Lamarckism amplifies the emergence of `morphological intelligence', the ability of a given robot body to acquire a good brain by learning, and identify the source of this success: `newborn' robots have a higher fitness because their inherited brains match their bodies better than those in a Darwinian system.
Radiology-Llama2: Best-in-Class Large Language Model for Radiology
Aug 29, 2023Zhengliang Liu, Yiwei Li, Peng Shu, Aoxiao Zhong, Longtao Yang, Chao Ju, Zihao Wu, Chong Ma, Jie Luo, Cheng Chen, Sekeun Kim, Jiang Hu, Haixing Dai, Lin Zhao, Dajiang Zhu, Jun Liu, Wei Liu, Dinggang Shen, Tianming Liu, Quanzheng Li, Xiang Li
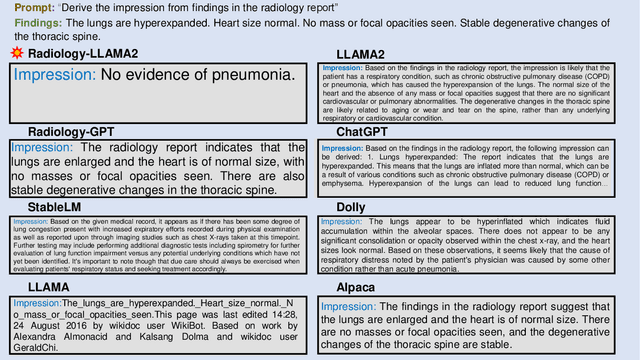
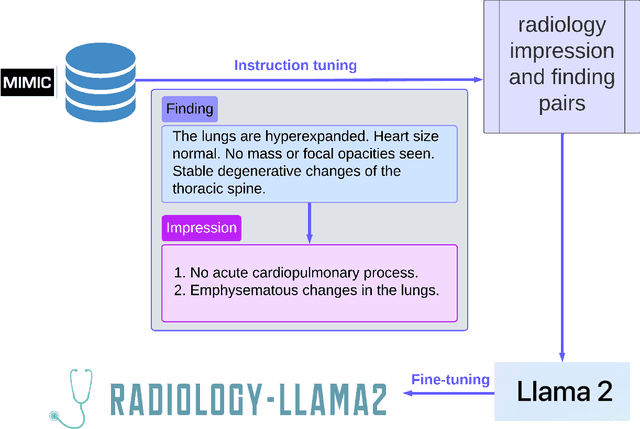
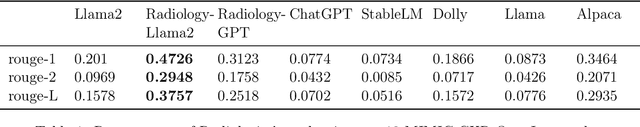

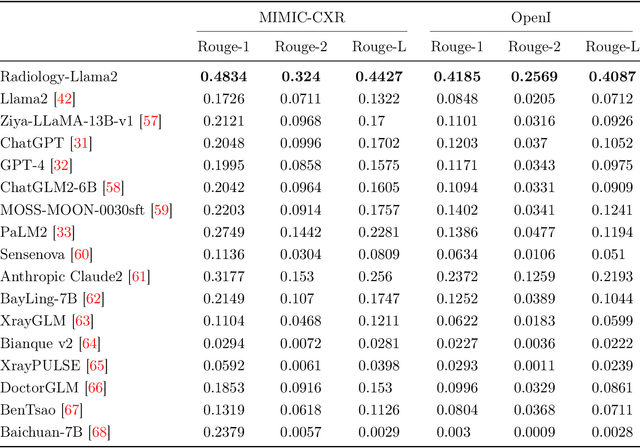
This paper introduces Radiology-Llama2, a large language model specialized for radiology through a process known as instruction tuning. Radiology-Llama2 is based on the Llama2 architecture and further trained on a large dataset of radiology reports to generate coherent and clinically useful impressions from radiological findings. Quantitative evaluations using ROUGE metrics on the MIMIC-CXR and OpenI datasets demonstrate that Radiology-Llama2 achieves state-of-the-art performance compared to other generative language models, with a Rouge-1 score of 0.4834 on MIMIC-CXR and 0.4185 on OpenI. Additional assessments by radiology experts highlight the model's strengths in understandability, coherence, relevance, conciseness, and clinical utility. The work illustrates the potential of localized language models designed and tuned for specialized domains like radiology. When properly evaluated and deployed, such models can transform fields like radiology by automating rote tasks and enhancing human expertise.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge