Causal Coupled Mechanisms: A Control Method with Cooperation and Competition for Complex System
Sep 15, 2022Xuehui Yu, Jingchi Jiang, Xinmiao Yu, Yi Guan, Xue Li
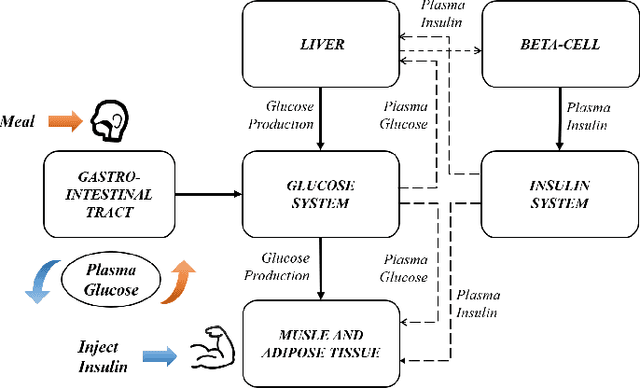


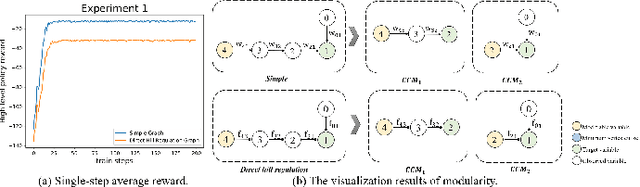
Complex systems are ubiquitous in the real world and tend to have complicated and poorly understood dynamics. For their control issues, the challenge is to guarantee accuracy, robustness, and generalization in such bloated and troubled environments. Fortunately, a complex system can be divided into multiple modular structures that human cognition appears to exploit. Inspired by this cognition, a novel control method, Causal Coupled Mechanisms (CCMs), is proposed that explores the cooperation in division and competition in combination. Our method employs the theory of hierarchical reinforcement learning (HRL), in which 1) the high-level policy with competitive awareness divides the whole complex system into multiple functional mechanisms, and 2) the low-level policy finishes the control task of each mechanism. Specifically for cooperation, a cascade control module helps the series operation of CCMs, and a forward coupled reasoning module is used to recover the coupling information lost in the division process. On both synthetic systems and a real-world biological regulatory system, the CCM method achieves robust and state-of-the-art control results even with unpredictable random noise. Moreover, generalization results show that reusing prepared specialized CCMs helps to perform well in environments with different confounders and dynamics.
Medical Knowledge Embedding Based on Recursive Neural Network for Multi-Disease Diagnosis
Sep 22, 2018Jingchi Jiang, Huanzheng Wang, Jing Xie, Xitong Guo, Yi Guan, Qiubin Yu
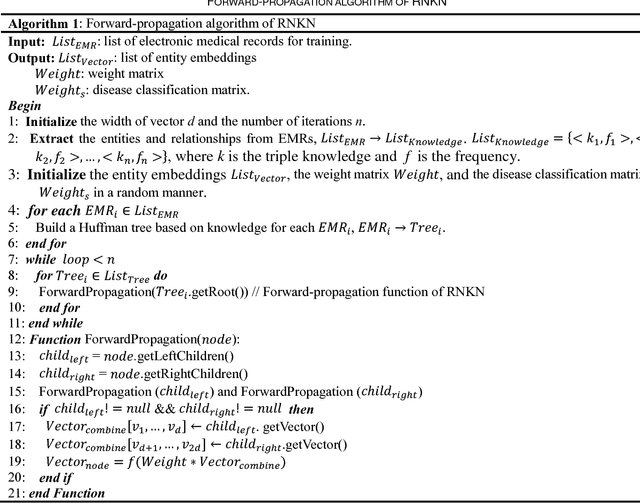

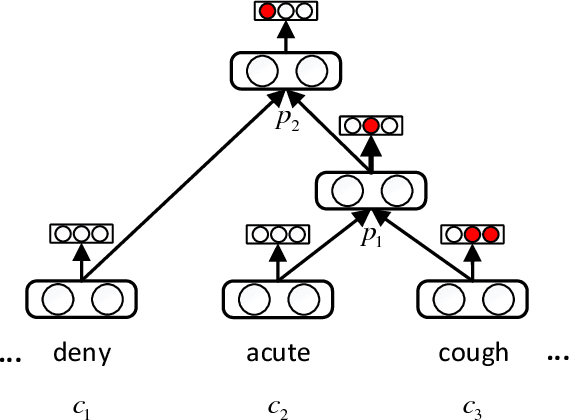


The representation of knowledge based on first-order logic captures the richness of natural language and supports multiple probabilistic inference models. Although symbolic representation enables quantitative reasoning with statistical probability, it is difficult to utilize with machine learning models as they perform numerical operations. In contrast, knowledge embedding (i.e., high-dimensional and continuous vectors) is a feasible approach to complex reasoning that can not only retain the semantic information of knowledge but also establish the quantifiable relationship among them. In this paper, we propose recursive neural knowledge network (RNKN), which combines medical knowledge based on first-order logic with recursive neural network for multi-disease diagnosis. After RNKN is efficiently trained from manually annotated Chinese Electronic Medical Records (CEMRs), diagnosis-oriented knowledge embeddings and weight matrixes are learned. Experimental results verify that the diagnostic accuracy of RNKN is superior to that of some classical machine learning models and Markov logic network (MLN). The results also demonstrate that the more explicit the evidence extracted from CEMRs is, the better is the performance achieved. RNKN gradually exhibits the interpretation of knowledge embeddings as the number of training epochs increases.
De-identification of medical records using conditional random fields and long short-term memory networks
Sep 29, 2017Zhipeng Jiang, Chao Zhao, Bin He, Yi Guan, Jingchi Jiang

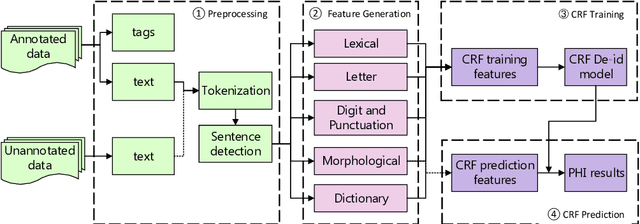

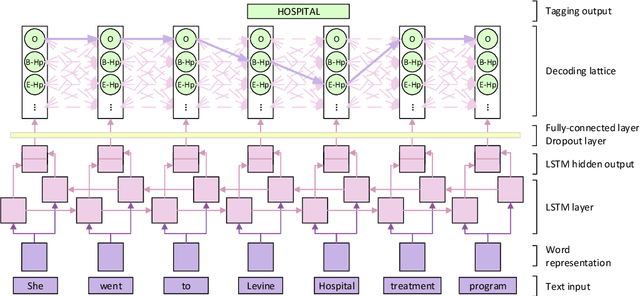
The CEGS N-GRID 2016 Shared Task 1 in Clinical Natural Language Processing focuses on the de-identification of psychiatric evaluation records. This paper describes two participating systems of our team, based on conditional random fields (CRFs) and long short-term memory networks (LSTMs). A pre-processing module was introduced for sentence detection and tokenization before de-identification. For CRFs, manually extracted rich features were utilized to train the model. For LSTMs, a character-level bi-directional LSTM network was applied to represent tokens and classify tags for each token, following which a decoding layer was stacked to decode the most probable protected health information (PHI) terms. The LSTM-based system attained an i2b2 strict micro-F_1 measure of 89.86%, which was higher than that of the CRF-based system.
EMR-based medical knowledge representation and inference via Markov random fields and distributed representation learning
Sep 20, 2017Chao Zhao, Jingchi Jiang, Yi Guan
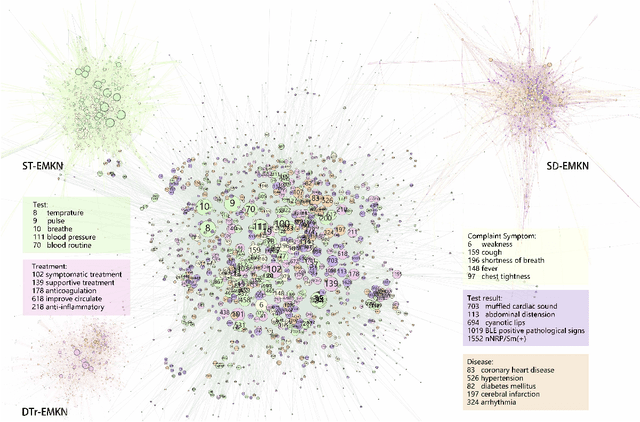
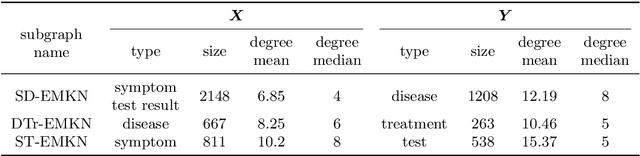
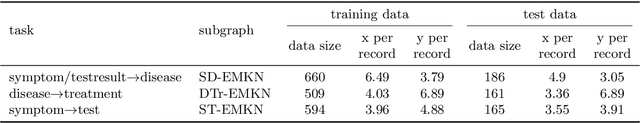
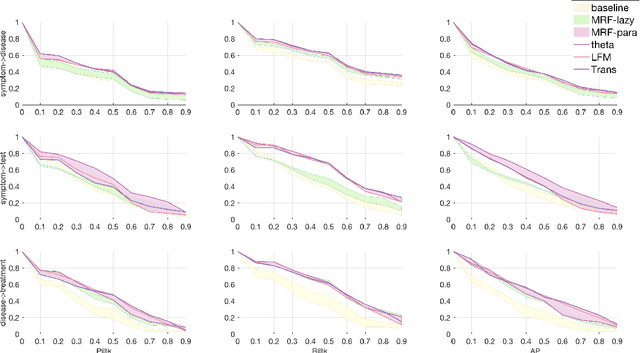
Objective: Electronic medical records (EMRs) contain an amount of medical knowledge which can be used for clinical decision support (CDS). Our objective is a general system that can extract and represent these knowledge contained in EMRs to support three CDS tasks: test recommendation, initial diagnosis, and treatment plan recommendation, with the given condition of one patient. Methods: We extracted four kinds of medical entities from records and constructed an EMR-based medical knowledge network (EMKN), in which nodes are entities and edges reflect their co-occurrence in a single record. Three bipartite subgraphs (bi-graphs) were extracted from the EMKN to support each task. One part of the bi-graph was the given condition (e.g., symptoms), and the other was the condition to be inferred (e.g., diseases). Each bi-graph was regarded as a Markov random field to support the inference. Three lazy energy functions and one parameter-based energy function were proposed, as well as two knowledge representation learning-based energy functions, which can provide a distributed representation of medical entities. Three measures were utilized for performance evaluation. Results: On the initial diagnosis task, 80.11% of the test records identified at least one correct disease from top 10 candidates. Test and treatment recommendation results were 87.88% and 92.55%, respectively. These results altogether indicate that the proposed system outperformed the baseline methods. The distributed representation of medical entities does reflect similarity relationships in regards to knowledge level. Conclusion: Combining EMKN and MRF is an effective approach for general medical knowledge representation and inference. Different tasks, however, require designing their energy functions individually.
Learning and inference in knowledge-based probabilistic model for medical diagnosis
Mar 28, 2017Jingchi Jiang, Chao Zhao, Yi Guan, Qiubin Yu


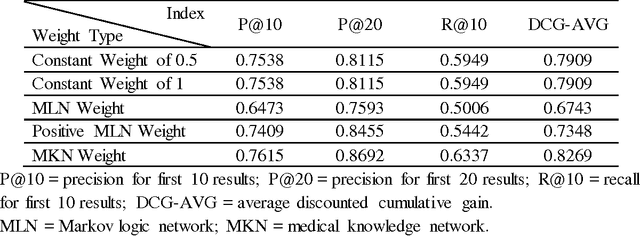
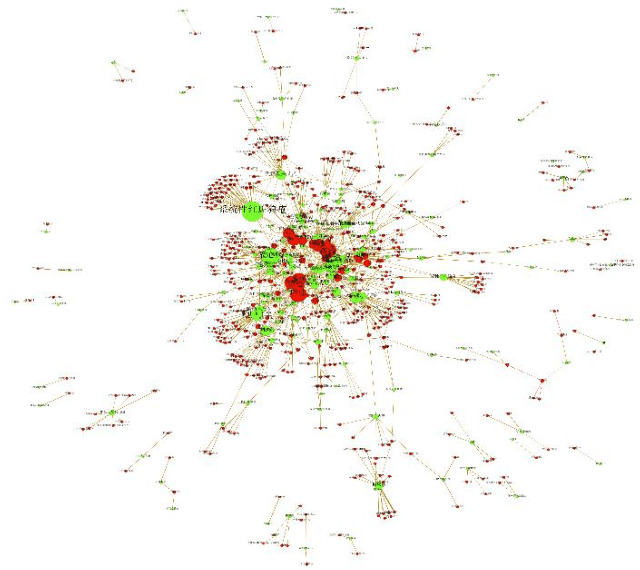
Based on a weighted knowledge graph to represent first-order knowledge and combining it with a probabilistic model, we propose a methodology for the creation of a medical knowledge network (MKN) in medical diagnosis. When a set of symptoms is activated for a specific patient, we can generate a ground medical knowledge network composed of symptom nodes and potential disease nodes. By Incorporating a Boltzmann machine into the potential function of a Markov network, we investigated the joint probability distribution of the MKN. In order to deal with numerical symptoms, a multivariate inference model is presented that uses conditional probability. In addition, the weights for the knowledge graph were efficiently learned from manually annotated Chinese Electronic Medical Records (CEMRs). In our experiments, we found numerically that the optimum choice of the quality of disease node and the expression of symptom variable can improve the effectiveness of medical diagnosis. Our experimental results comparing a Markov logic network and the logistic regression algorithm on an actual CEMR database indicate that our method holds promise and that MKN can facilitate studies of intelligent diagnosis.
Developing a cardiovascular disease risk factor annotated corpus of Chinese electronic medical records
Mar 03, 2017Jia Su, Bin He, Yi Guan, Jingchi Jiang, Jinfeng Yang
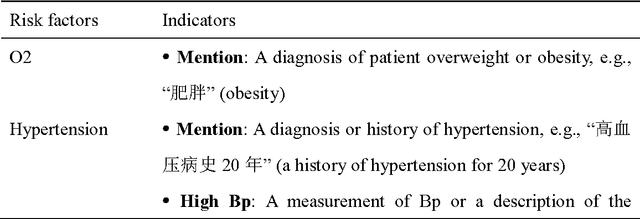

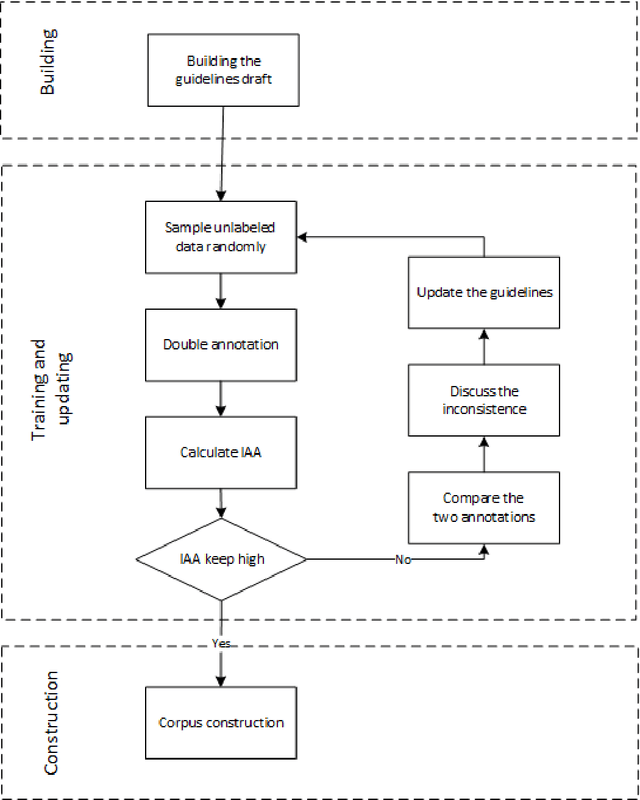
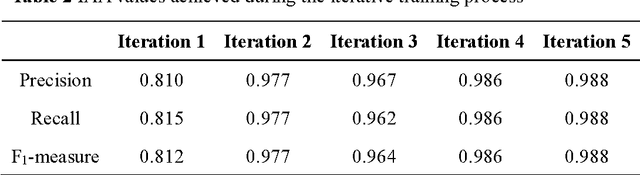
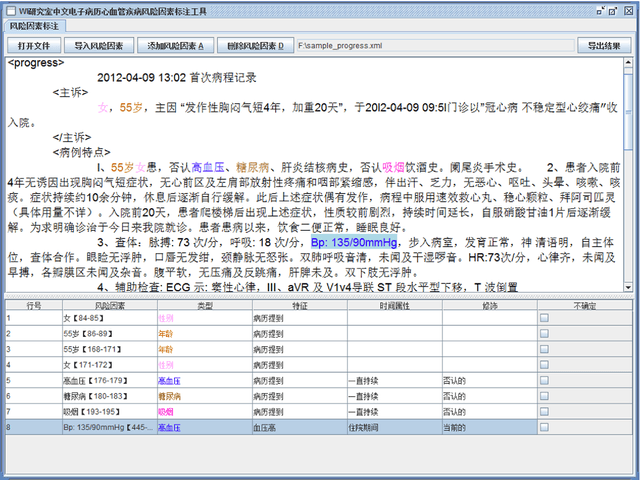
Cardiovascular disease (CVD) has become the leading cause of death in China, and most of the cases can be prevented by controlling risk factors. The goal of this study was to build a corpus of CVD risk factor annotations based on Chinese electronic medical records (CEMRs). This corpus is intended to be used to develop a risk factor information extraction system that, in turn, can be applied as a foundation for the further study of the progress of risk factors and CVD. We designed a light annotation task to capture CVD risk factors with indicators, temporal attributes and assertions that were explicitly or implicitly displayed in the records. The task included: 1) preparing data; 2) creating guidelines for capturing annotations (these were created with the help of clinicians); 3) proposing an annotation method including building the guidelines draft, training the annotators and updating the guidelines, and corpus construction. Then, a risk factor annotated corpus based on de-identified discharge summaries and progress notes from 600 patients was developed. Built with the help of clinicians, this corpus has an inter-annotator agreement (IAA) F1-measure of 0.968, indicating a high reliability. To the best of our knowledge, this is the first annotated corpus concerning CVD risk factors in CEMRs and the guidelines for capturing CVD risk factor annotations from CEMRs were proposed. The obtained document-level annotations can be applied in future studies to monitor risk factors and CVD over the long term.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge