Model-based causal feature selection for general response types
Sep 22, 2023Lucas Kook, Sorawit Saengkyongam, Anton Rask Lundborg, Torsten Hothorn, Jonas Peters
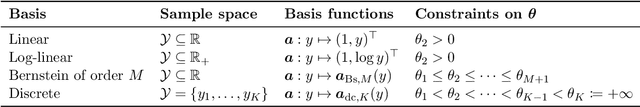
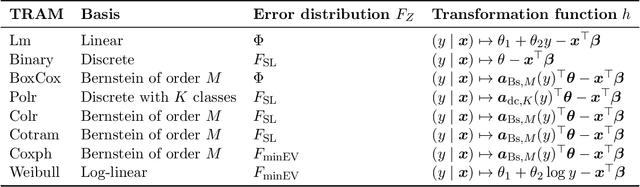
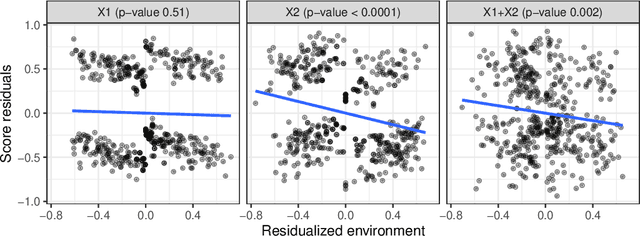


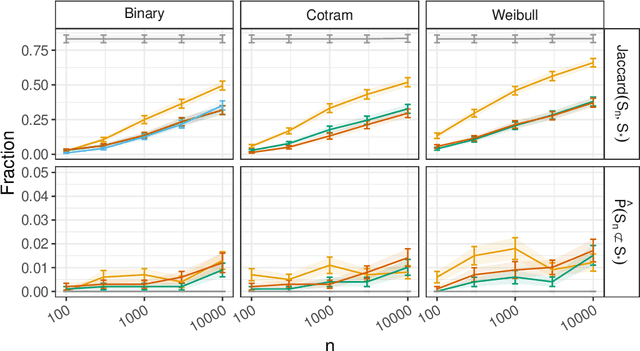
Discovering causal relationships from observational data is a fundamental yet challenging task. In some applications, it may suffice to learn the causal features of a given response variable, instead of learning the entire underlying causal structure. Invariant causal prediction (ICP, Peters et al., 2016) is a method for causal feature selection which requires data from heterogeneous settings. ICP assumes that the mechanism for generating the response from its direct causes is the same in all settings and exploits this invariance to output a subset of the causal features. The framework of ICP has been extended to general additive noise models and to nonparametric settings using conditional independence testing. However, nonparametric conditional independence testing often suffers from low power (or poor type I error control) and the aforementioned parametric models are not suitable for applications in which the response is not measured on a continuous scale, but rather reflects categories or counts. To bridge this gap, we develop ICP in the context of transformation models (TRAMs), allowing for continuous, categorical, count-type, and uninformatively censored responses (we show that, in general, these model classes do not allow for identifiability when there is no exogenous heterogeneity). We propose TRAM-GCM, a test for invariance of a subset of covariates, based on the expected conditional covariance between environments and score residuals which satisfies uniform asymptotic level guarantees. For the special case of linear shift TRAMs, we propose an additional invariance test, TRAM-Wald, based on the Wald statistic. We implement both proposed methods in the open-source R package "tramicp" and show in simulations that under the correct model specification, our approach empirically yields higher power than nonparametric ICP based on conditional independence testing.
Deep conditional transformation models for survival analysis
Oct 20, 2022Gabriele Campanella, Lucas Kook, Ida Häggström, Torsten Hothorn, Thomas J. Fuchs
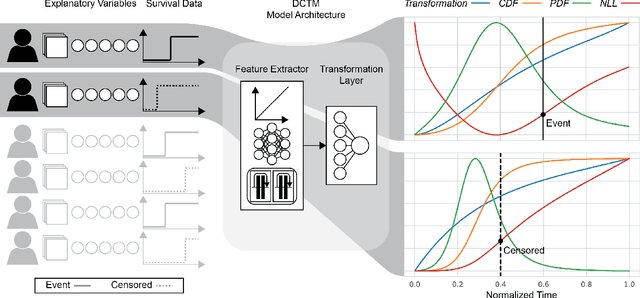
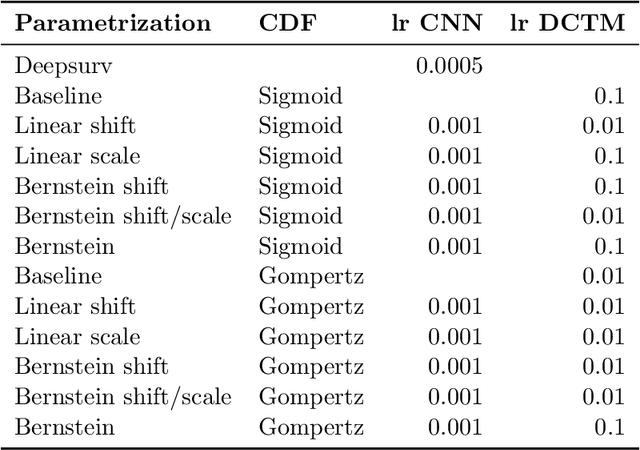
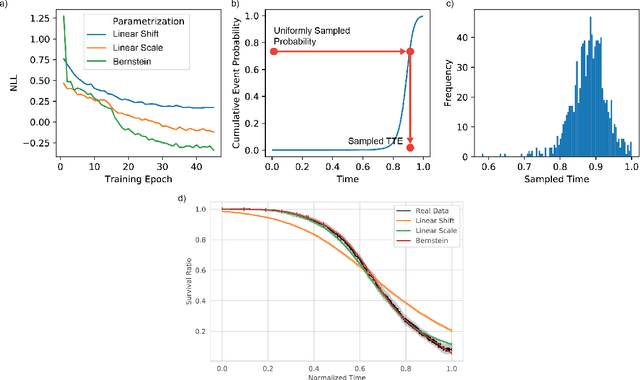
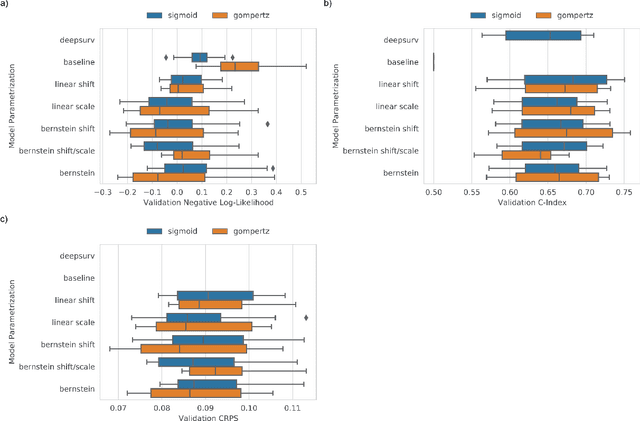
An every increasing number of clinical trials features a time-to-event outcome and records non-tabular patient data, such as magnetic resonance imaging or text data in the form of electronic health records. Recently, several neural-network based solutions have been proposed, some of which are binary classifiers. Parametric, distribution-free approaches which make full use of survival time and censoring status have not received much attention. We present deep conditional transformation models (DCTMs) for survival outcomes as a unifying approach to parametric and semiparametric survival analysis. DCTMs allow the specification of non-linear and non-proportional hazards for both tabular and non-tabular data and extend to all types of censoring and truncation. On real and semi-synthetic data, we show that DCTMs compete with state-of-the-art DL approaches to survival analysis.
Deep interpretable ensembles
May 25, 2022Lucas Kook, Andrea Götschi, Philipp FM Baumann, Torsten Hothorn, Beate Sick
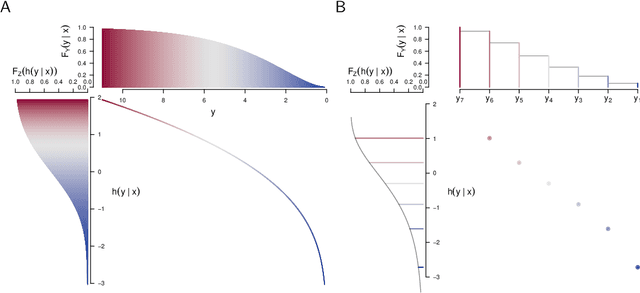
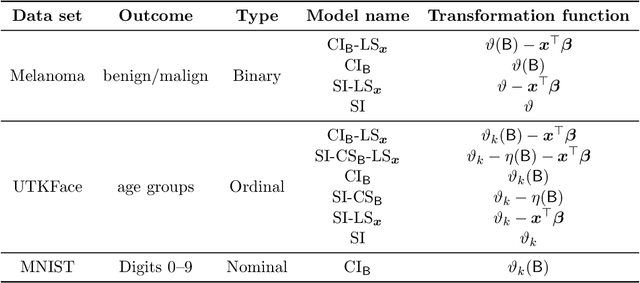
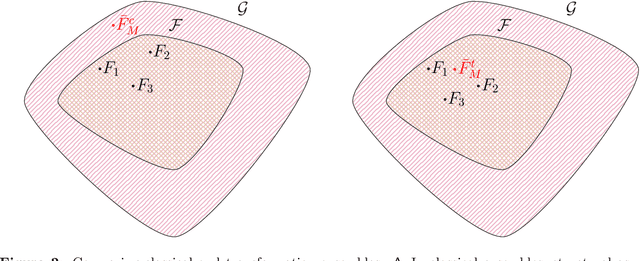
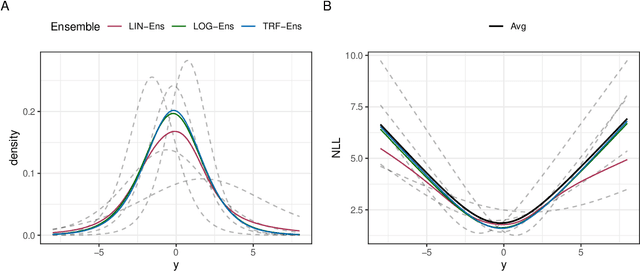
Ensembles improve prediction performance and allow uncertainty quantification by aggregating predictions from multiple models. In deep ensembling, the individual models are usually black box neural networks, or recently, partially interpretable semi-structured deep transformation models. However, interpretability of the ensemble members is generally lost upon aggregation. This is a crucial drawback of deep ensembles in high-stake decision fields, in which interpretable models are desired. We propose a novel transformation ensemble which aggregates probabilistic predictions with the guarantee to preserve interpretability and yield uniformly better predictions than the ensemble members on average. Transformation ensembles are tailored towards interpretable deep transformation models but are applicable to a wider range of probabilistic neural networks. In experiments on several publicly available data sets, we demonstrate that transformation ensembles perform on par with classical deep ensembles in terms of prediction performance, discrimination, and calibration. In addition, we demonstrate how transformation ensembles quantify both aleatoric and epistemic uncertainty, and produce minimax optimal predictions under certain conditions.
Ordinal Neural Network Transformation Models: Deep and interpretable regression models for ordinal outcomes
Oct 26, 2020Lucas Kook, Lisa Herzog, Torsten Hothorn, Oliver Dürr, Beate Sick
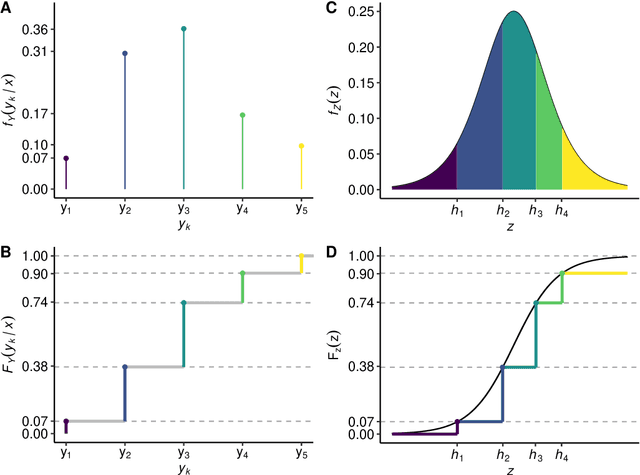
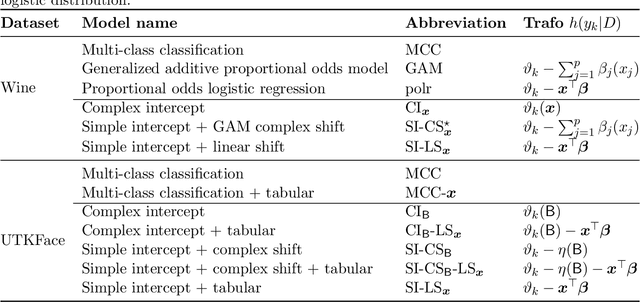
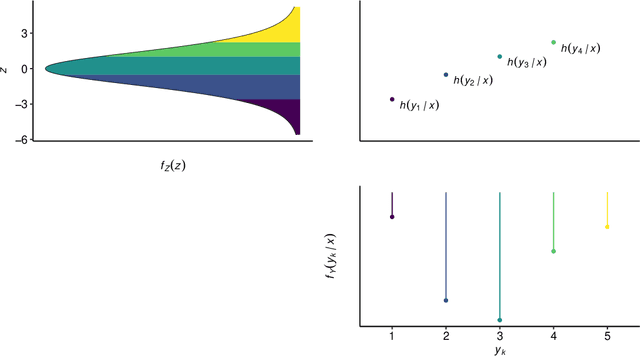
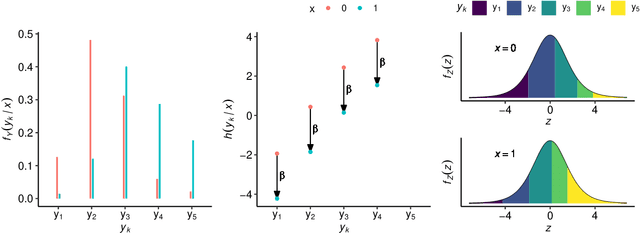
Outcomes with a natural order commonly occur in prediction tasks and oftentimes the available input data are a mixture of complex data, like images, and tabular predictors. Although deep Learning (DL) methods have shown outstanding performance on image classification, most models treat ordered outcomes as unordered and lack interpretability. In contrast, classical ordinal regression models yield interpretable predictor effects but are limited to tabular input data. Here, we present the highly modular class of ordinal neural network transformation models (ONTRAMs). Transformation models use a parametric transformation function and a simple distribution to trade off flexibility and interpretability of individual model components. In ONTRAMs, this trade-off is achieved by additively decomposing the transformation function into terms for the tabular and image data using a set of jointly trained neural networks. We show that the most flexible ONTRAMs achieve on-par performance with DL classifiers while outperforming them in training speed. We discuss how to interpret components of ONTRAMs in general and in the case of correlated tabular and image data. Taken together, ONTRAMs join benefits of DL and distributional regression to create interpretable prediction models for ordinal outcomes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge