MEDVOC: Vocabulary Adaptation for Fine-tuning Pre-trained Language Models on Medical Text Summarization
May 07, 2024Gunjan Balde, Soumyadeep Roy, Mainack Mondal, Niloy Ganguly
This work presents a dynamic vocabulary adaptation strategy, MEDVOC, for fine-tuning pre-trained language models (PLMs) like BertSumAbs, BART, and PEGASUS for improved medical text summarization. In contrast to existing domain adaptation approaches in summarization, MEDVOC treats vocabulary as an optimizable parameter and optimizes the PLM vocabulary based on fragment score conditioned only on the downstream task's reference summaries. Unlike previous works on vocabulary adaptation (limited only to classification tasks), optimizing vocabulary based on summarization tasks requires an extremely costly intermediate fine-tuning step on large summarization datasets. To that end, our novel fragment score-based hyperparameter search very significantly reduces this fine-tuning time -- from 450 days to less than 2 days on average. Furthermore, while previous works on vocabulary adaptation are often primarily tied to single PLMs, MEDVOC is designed to be deployable across multiple PLMs (with varying model vocabulary sizes, pre-training objectives, and model sizes) -- bridging the limited vocabulary overlap between the biomedical literature domain and PLMs. MEDVOC outperforms baselines by 15.74% in terms of Rouge-L in zero-shot setting and shows gains of 17.29% in high Out-Of-Vocabulary (OOV) concentrations. Our human evaluation shows MEDVOC generates more faithful medical summaries (88% compared to 59% in baselines). We make the codebase publicly available at https://github.com/gb-kgp/MEDVOC.
TIGQA:An Expert Annotated Question Answering Dataset in Tigrinya
Apr 26, 2024Hailay Teklehaymanot, Dren Fazlija, Niloy Ganguly, Gourab K. Patro, Wolfgang Nejdl
The absence of explicitly tailored, accessible annotated datasets for educational purposes presents a notable obstacle for NLP tasks in languages with limited resources.This study initially explores the feasibility of using machine translation (MT) to convert an existing dataset into a Tigrinya dataset in SQuAD format. As a result, we present TIGQA, an expert annotated educational dataset consisting of 2.68K question-answer pairs covering 122 diverse topics such as climate, water, and traffic. These pairs are from 537 context paragraphs in publicly accessible Tigrinya and Biology books. Through comprehensive analyses, we demonstrate that the TIGQA dataset requires skills beyond simple word matching, requiring both single-sentence and multiple-sentence inference abilities. We conduct experiments using state-of-the art MRC methods, marking the first exploration of such models on TIGQA. Additionally, we estimate human performance on the dataset and juxtapose it with the results obtained from pretrained models.The notable disparities between human performance and best model performance underscore the potential for further enhancements to TIGQA through continued research. Our dataset is freely accessible via the provided link to encourage the research community to address the challenges in the Tigrinya MRC.
* 9 pages,3 figures, 7 tables,2 listings
Beyond Accuracy: Investigating Error Types in GPT-4 Responses to USMLE Questions
Apr 20, 2024Soumyadeep Roy, Aparup Khatua, Fatemeh Ghoochani, Uwe Hadler, Wolfgang Nejdl, Niloy Ganguly
GPT-4 demonstrates high accuracy in medical QA tasks, leading with an accuracy of 86.70%, followed by Med-PaLM 2 at 86.50%. However, around 14% of errors remain. Additionally, current works use GPT-4 to only predict the correct option without providing any explanation and thus do not provide any insight into the thinking process and reasoning used by GPT-4 or other LLMs. Therefore, we introduce a new domain-specific error taxonomy derived from collaboration with medical students. Our GPT-4 USMLE Error (G4UE) dataset comprises 4153 GPT-4 correct responses and 919 incorrect responses to the United States Medical Licensing Examination (USMLE) respectively. These responses are quite long (258 words on average), containing detailed explanations from GPT-4 justifying the selected option. We then launch a large-scale annotation study using the Potato annotation platform and recruit 44 medical experts through Prolific, a well-known crowdsourcing platform. We annotated 300 out of these 919 incorrect data points at a granular level for different classes and created a multi-label span to identify the reasons behind the error. In our annotated dataset, a substantial portion of GPT-4's incorrect responses is categorized as a "Reasonable response by GPT-4," by annotators. This sheds light on the challenge of discerning explanations that may lead to incorrect options, even among trained medical professionals. We also provide medical concepts and medical semantic predications extracted using the SemRep tool for every data point. We believe that it will aid in evaluating the ability of LLMs to answer complex medical questions. We make the resources available at https://github.com/roysoumya/usmle-gpt4-error-taxonomy .
Order-Based Pre-training Strategies for Procedural Text Understanding
Apr 06, 2024Abhilash Nandy, Yash Kulkarni, Pawan Goyal, Niloy Ganguly
In this paper, we propose sequence-based pretraining methods to enhance procedural understanding in natural language processing. Procedural text, containing sequential instructions to accomplish a task, is difficult to understand due to the changing attributes of entities in the context. We focus on recipes, which are commonly represented as ordered instructions, and use this order as a supervision signal. Our work is one of the first to compare several 'order as-supervision' transformer pre-training methods, including Permutation Classification, Embedding Regression, and Skip-Clip, and shows that these methods give improved results compared to the baselines and SoTA LLMs on two downstream Entity-Tracking datasets: NPN-Cooking dataset in recipe domain and ProPara dataset in open domain. Our proposed methods address the non-trivial Entity Tracking Task that requires prediction of entity states across procedure steps, which requires understanding the order of steps. These methods show an improvement over the best baseline by 1.6% and 7-9% on NPN-Cooking and ProPara Datasets respectively across metrics.
How Robust are the Tabular QA Models for Scientific Tables? A Study using Customized Dataset
Mar 30, 2024Akash Ghosh, B Venkata Sahith, Niloy Ganguly, Pawan Goyal, Mayank Singh
Question-answering (QA) on hybrid scientific tabular and textual data deals with scientific information, and relies on complex numerical reasoning. In recent years, while tabular QA has seen rapid progress, understanding their robustness on scientific information is lacking due to absence of any benchmark dataset. To investigate the robustness of the existing state-of-the-art QA models on scientific hybrid tabular data, we propose a new dataset, "SciTabQA", consisting of 822 question-answer pairs from scientific tables and their descriptions. With the help of this dataset, we assess the state-of-the-art Tabular QA models based on their ability (i) to use heterogeneous information requiring both structured data (table) and unstructured data (text) and (ii) to perform complex scientific reasoning tasks. In essence, we check the capability of the models to interpret scientific tables and text. Our experiments show that "SciTabQA" is an innovative dataset to study question-answering over scientific heterogeneous data. We benchmark three state-of-the-art Tabular QA models, and find that the best F1 score is only 0.462.
Cost-Performance Optimization for Processing Low-Resource Language Tasks Using Commercial LLMs
Mar 08, 2024Arijit Nag, Animesh Mukherjee, Niloy Ganguly, Soumen Chakrabarti
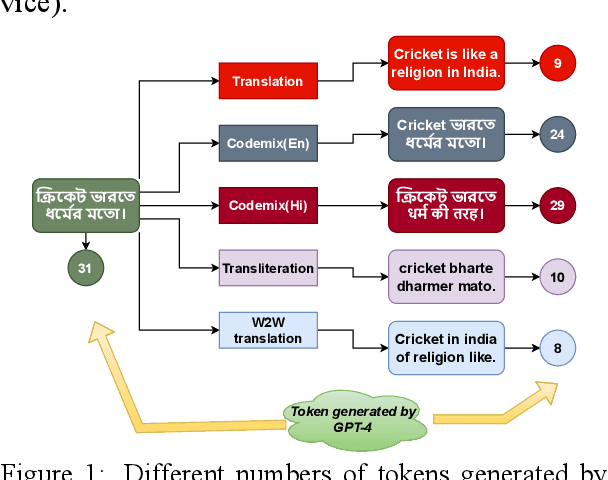
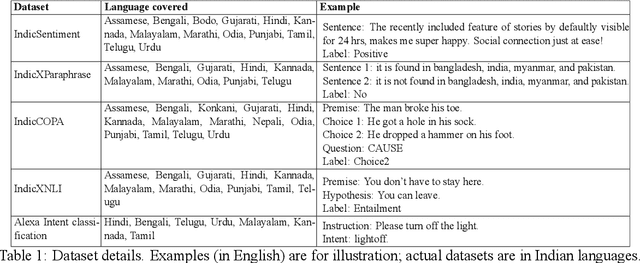
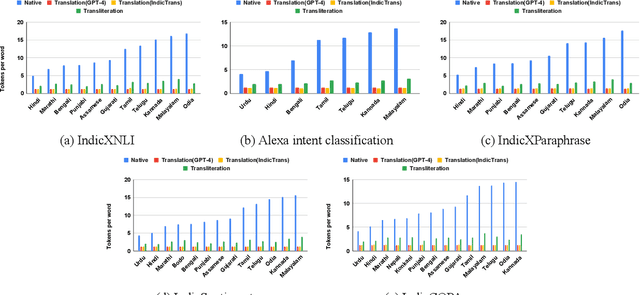
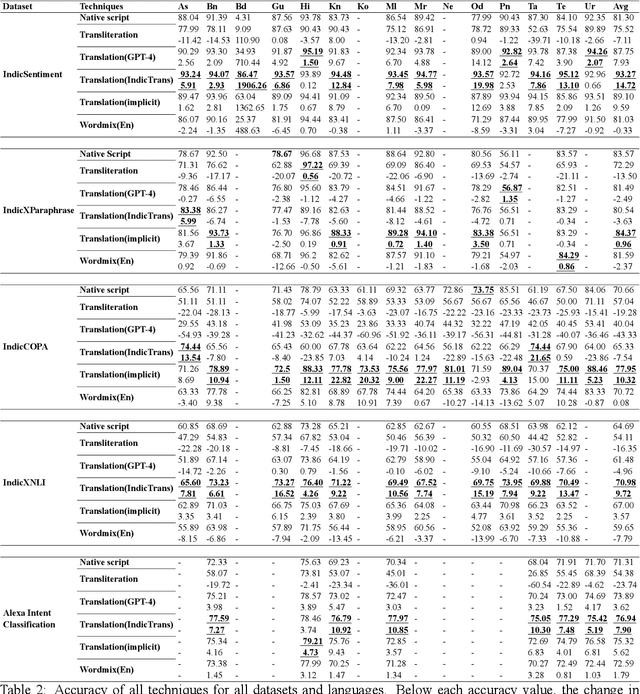
Large Language Models (LLMs) exhibit impressive zero/few-shot inference and generation quality for high-resource languages(HRLs). A few of them have been trained in low-resource languages (LRLs) and give decent performance. Owing to the prohibitive costs of training LLMs, they are usually used as a network service, with the client charged by the count of input and output tokens. The number of tokens strongly depends on the script and language, as well as the LLM's sub-word vocabulary. We show that LRLs are at a pricing disadvantage, because the well-known LLMs produce more tokens for LRLs than HRLs. This is because most currently popular LLMs are optimized for HRL vocabularies. Our objective is to level the playing field: reduce the cost of processing LRLs in contemporary LLMs while ensuring that predictive and generative qualities are not compromised. As means to reduce the number of tokens processed by the LLM, we consider code-mixing, translation, and transliteration of LRLs to HRLs. We perform an extensive study using the IndicXTREME dataset, covering 15 Indian languages, while using GPT-4 (one of the costliest LLM services released so far) as a commercial LLM. We observe and analyze interesting patterns involving token count, cost,and quality across a multitude of languages and tasks. We show that choosing the best policy to interact with the LLM can reduce cost by 90% while giving better or comparable performance, compared to communicating with the LLM in the original LRL.
Long Dialog Summarization: An Analysis
Feb 26, 2024Ankan Mullick, Ayan Kumar Bhowmick, Raghav R, Ravi Kokku, Prasenjit Dey, Pawan Goyal, Niloy Ganguly
Dialog summarization has become increasingly important in managing and comprehending large-scale conversations across various domains. This task presents unique challenges in capturing the key points, context, and nuances of multi-turn long conversations for summarization. It is worth noting that the summarization techniques may vary based on specific requirements such as in a shopping-chatbot scenario, the dialog summary helps to learn user preferences, whereas in the case of a customer call center, the summary may involve the problem attributes that a user specified, and the final resolution provided. This work emphasizes the significance of creating coherent and contextually rich summaries for effective communication in various applications. We explore current state-of-the-art approaches for long dialog summarization in different domains and benchmark metrics based evaluations show that one single model does not perform well across various areas for distinct summarization tasks.
BOXREC: Recommending a Box of Preferred Outfits in Online Shopping
Feb 26, 2024Debopriyo Banerjee, Krothapalli Sreenivasa Rao, Shamik Sural, Niloy Ganguly
Over the past few years, automation of outfit composition has gained much attention from the research community. Most of the existing outfit recommendation systems focus on pairwise item compatibility prediction (using visual and text features) to score an outfit combination having several items, followed by recommendation of top-n outfits or a capsule wardrobe having a collection of outfits based on user's fashion taste. However, none of these consider user's preference of price-range for individual clothing types or an overall shopping budget for a set of items. In this paper, we propose a box recommendation framework - BOXREC - which at first, collects user preferences across different item types (namely, top-wear, bottom-wear and foot-wear) including price-range of each type and a maximum shopping budget for a particular shopping session. It then generates a set of preferred outfits by retrieving all types of preferred items from the database (according to user specified preferences including price-ranges), creates all possible combinations of three preferred items (belonging to distinct item types) and verifies each combination using an outfit scoring framework - BOXREC-OSF. Finally, it provides a box full of fashion items, such that different combinations of the items maximize the number of outfits suitable for an occasion while satisfying maximum shopping budget. Empirical results show superior performance of BOXREC-OSF over the baseline methods.
CLMSM: A Multi-Task Learning Framework for Pre-training on Procedural Text
Oct 22, 2023Abhilash Nandy, Manav Nitin Kapadnis, Pawan Goyal, Niloy Ganguly
In this paper, we propose CLMSM, a domain-specific, continual pre-training framework, that learns from a large set of procedural recipes. CLMSM uses a Multi-Task Learning Framework to optimize two objectives - a) Contrastive Learning using hard triplets to learn fine-grained differences across entities in the procedures, and b) a novel Mask-Step Modelling objective to learn step-wise context of a procedure. We test the performance of CLMSM on the downstream tasks of tracking entities and aligning actions between two procedures on three datasets, one of which is an open-domain dataset not conforming with the pre-training dataset. We show that CLMSM not only outperforms baselines on recipes (in-domain) but is also able to generalize to open-domain procedural NLP tasks.
GeneMask: Fast Pretraining of Gene Sequences to Enable Few-Shot Learning
Jul 29, 2023Soumyadeep Roy, Jonas Wallat, Sowmya S Sundaram, Wolfgang Nejdl, Niloy Ganguly




Large-scale language models such as DNABert and LOGO aim to learn optimal gene representations and are trained on the entire Human Reference Genome. However, standard tokenization schemes involve a simple sliding window of tokens like k-mers that do not leverage any gene-based semantics and thus may lead to (trivial) masking of easily predictable sequences and subsequently inefficient Masked Language Modeling (MLM) training. Therefore, we propose a novel masking algorithm, GeneMask, for MLM training of gene sequences, where we randomly identify positions in a gene sequence as mask centers and locally select the span around the mask center with the highest Normalized Pointwise Mutual Information (NPMI) to mask. We observe that in the absence of human-understandable semantics in the genomics domain (in contrast, semantic units like words and phrases are inherently available in NLP), GeneMask-based models substantially outperform the SOTA models (DNABert and LOGO) over four benchmark gene sequence classification datasets in five few-shot settings (10 to 1000-shot). More significantly, the GeneMask-based DNABert model is trained for less than one-tenth of the number of epochs of the original SOTA model. We also observe a strong correlation between top-ranked PMI tokens and conserved DNA sequence motifs, which may indicate the incorporation of latent genomic information. The codes (including trained models) and datasets are made publicly available at https://github.com/roysoumya/GeneMask.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge