Estimating Causal Effects with Double Machine Learning -- A Method Evaluation
Mar 21, 2024Jonathan Fuhr, Philipp Berens, Dominik Papies
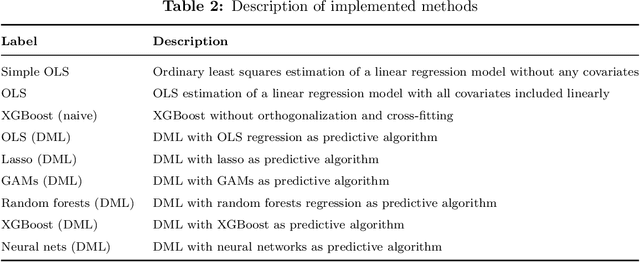
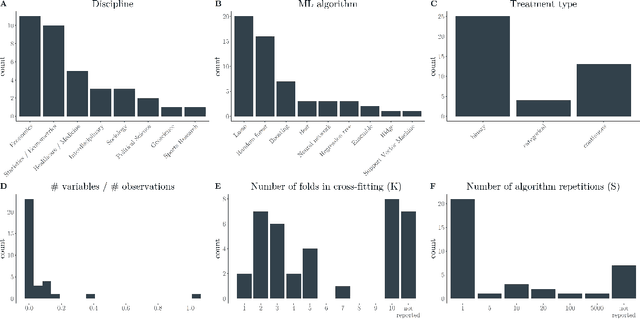
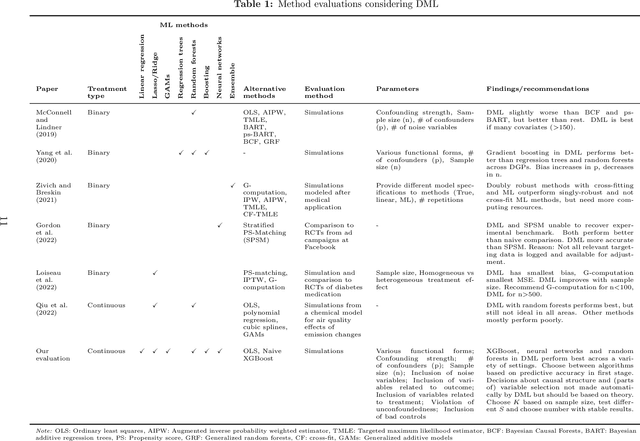

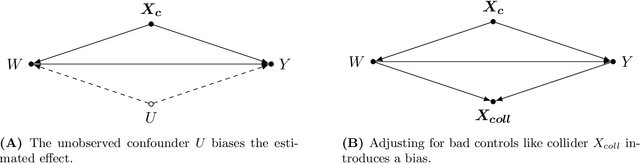
The estimation of causal effects with observational data continues to be a very active research area. In recent years, researchers have developed new frameworks which use machine learning to relax classical assumptions necessary for the estimation of causal effects. In this paper, we review one of the most prominent methods - "double/debiased machine learning" (DML) - and empirically evaluate it by comparing its performance on simulated data relative to more traditional statistical methods, before applying it to real-world data. Our findings indicate that the application of a suitably flexible machine learning algorithm within DML improves the adjustment for various nonlinear confounding relationships. This advantage enables a departure from traditional functional form assumptions typically necessary in causal effect estimation. However, we demonstrate that the method continues to critically depend on standard assumptions about causal structure and identification. When estimating the effects of air pollution on housing prices in our application, we find that DML estimates are consistently larger than estimates of less flexible methods. From our overall results, we provide actionable recommendations for specific choices researchers must make when applying DML in practice.
Disentangling representations of retinal images with generative models
Feb 29, 2024Sarah Müller, Lisa M. Koch, Hendrik P. A. Lensch, Philipp Berens
Retinal fundus images play a crucial role in the early detection of eye diseases and, using deep learning approaches, recent studies have even demonstrated their potential for detecting cardiovascular risk factors and neurological disorders. However, the impact of technical factors on these images can pose challenges for reliable AI applications in ophthalmology. For example, large fundus cohorts are often confounded by factors like camera type, image quality or illumination level, bearing the risk of learning shortcuts rather than the causal relationships behind the image generation process. Here, we introduce a novel population model for retinal fundus images that effectively disentangles patient attributes from camera effects, thus enabling controllable and highly realistic image generation. To achieve this, we propose a novel disentanglement loss based on distance correlation. Through qualitative and quantitative analyses, we demonstrate the effectiveness of this novel loss function in disentangling the learned subspaces. Our results show that our model provides a new perspective on the complex relationship between patient attributes and technical confounders in retinal fundus image generation.
Self-supervised Visualisation of Medical Image Datasets
Feb 22, 2024Ifeoma Veronica Nwabufo, Jan Niklas Böhm, Philipp Berens, Dmitry Kobak
Self-supervised learning methods based on data augmentations, such as SimCLR, BYOL, or DINO, allow obtaining semantically meaningful representations of image datasets and are widely used prior to supervised fine-tuning. A recent self-supervised learning method, $t$-SimCNE, uses contrastive learning to directly train a 2D representation suitable for visualisation. When applied to natural image datasets, $t$-SimCNE yields 2D visualisations with semantically meaningful clusters. In this work, we used $t$-SimCNE to visualise medical image datasets, including examples from dermatology, histology, and blood microscopy. We found that increasing the set of data augmentations to include arbitrary rotations improved the results in terms of class separability, compared to data augmentations used for natural images. Our 2D representations show medically relevant structures and can be used to aid data exploration and annotation, improving on common approaches for data visualisation.
Diffusion Tempering Improves Parameter Estimation with Probabilistic Integrators for Ordinary Differential Equations
Feb 19, 2024Jonas Beck, Nathanael Bosch, Michael Deistler, Kyra L. Kadhim, Jakob H. Macke, Philipp Hennig, Philipp Berens
Ordinary differential equations (ODEs) are widely used to describe dynamical systems in science, but identifying parameters that explain experimental measurements is challenging. In particular, although ODEs are differentiable and would allow for gradient-based parameter optimization, the nonlinear dynamics of ODEs often lead to many local minima and extreme sensitivity to initial conditions. We therefore propose diffusion tempering, a novel regularization technique for probabilistic numerical methods which improves convergence of gradient-based parameter optimization in ODEs. By iteratively reducing a noise parameter of the probabilistic integrator, the proposed method converges more reliably to the true parameters. We demonstrate that our method is effective for dynamical systems of different complexity and show that it obtains reliable parameter estimates for a Hodgkin-Huxley model with a practically relevant number of parameters.
A computational approach to visual ecology with deep reinforcement learning
Feb 07, 2024Sacha Sokoloski, Jure Majnik, Philipp Berens
Animal vision is thought to optimize various objectives from metabolic efficiency to discrimination performance, yet its ultimate objective is to facilitate the survival of the animal within its ecological niche. However, modeling animal behavior in complex environments has been challenging. To study how environments shape and constrain visual processing, we developed a deep reinforcement learning framework in which an agent moves through a 3-d environment that it perceives through a vision model, where its only goal is to survive. Within this framework we developed a foraging task where the agent must gather food that sustains it, and avoid food that harms it. We first established that the complexity of the vision model required for survival on this task scaled with the variety and visual complexity of the food in the environment. Moreover, we showed that a recurrent network architecture was necessary to fully exploit complex vision models on the most visually demanding tasks. Finally, we showed how different network architectures learned distinct representations of the environment and task, and lead the agent to exhibit distinct behavioural strategies. In summary, this paper lays the foundation for a computational approach to visual ecology, provides extensive benchmarks for future work, and demonstrates how representations and behaviour emerge from an agent's drive for survival.
Generating Realistic Counterfactuals for Retinal Fundus and OCT Images using Diffusion Models
Dec 04, 2023Indu Ilanchezian, Valentyn Boreiko, Laura Kühlewein, Ziwei Huang, Murat Seçkin Ayhan, Matthias Hein, Lisa Koch, Philipp Berens
Counterfactual reasoning is often used in clinical settings to explain decisions or weigh alternatives. Therefore, for imaging based specialties such as ophthalmology, it would be beneficial to be able to create counterfactual images, illustrating answers to questions like "If the subject had had diabetic retinopathy, how would the fundus image have looked?". Here, we demonstrate that using a diffusion model in combination with an adversarially robust classifier trained on retinal disease classification tasks enables the generation of highly realistic counterfactuals of retinal fundus images and optical coherence tomography (OCT) B-scans. The key to the realism of counterfactuals is that these classifiers encode salient features indicative for each disease class and can steer the diffusion model to depict disease signs or remove disease-related lesions in a realistic way. In a user study, domain experts also found the counterfactuals generated using our method significantly more realistic than counterfactuals generated from a previous method, and even indistinguishable from real images.
Persistent homology for high-dimensional data based on spectral methods
Nov 06, 2023Sebastian Damrich, Philipp Berens, Dmitry Kobak
Persistent homology is a popular computational tool for detecting non-trivial topology of point clouds, such as the presence of loops or voids. However, many real-world datasets with low intrinsic dimensionality reside in an ambient space of much higher dimensionality. We show that in this case vanilla persistent homology becomes very sensitive to noise and fails to detect the correct topology. The same holds true for most existing refinements of persistent homology. As a remedy, we find that spectral distances on the $k$-nearest-neighbor graph of the data, such as diffusion distance and effective resistance, allow persistent homology to detect the correct topology even in the presence of high-dimensional noise. Furthermore, we derive a novel closed-form expression for effective resistance in terms of the eigendecomposition of the graph Laplacian, and describe its relation to diffusion distances. Finally, we apply these methods to several high-dimensional single-cell RNA-sequencing datasets and show that spectral distances on the $k$-nearest-neighbor graph allow robust detection of cell cycle loops.
Deep Hypothesis Tests Detect Clinically Relevant Subgroup Shifts in Medical Images
Mar 08, 2023Lisa M. Koch, Christian M. Schürch, Christian F. Baumgartner, Arthur Gretton, Philipp Berens
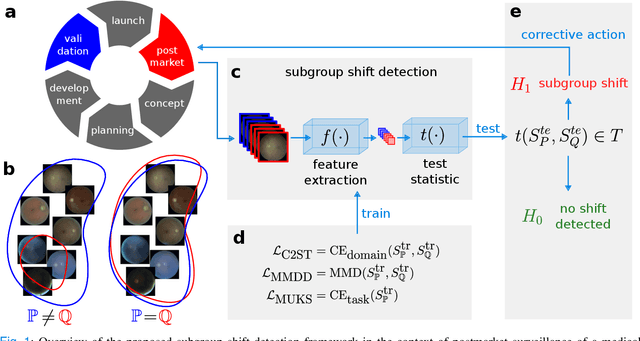

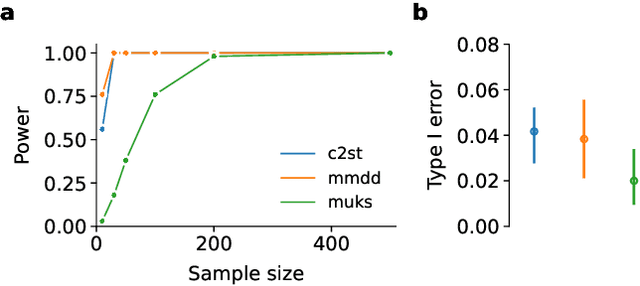
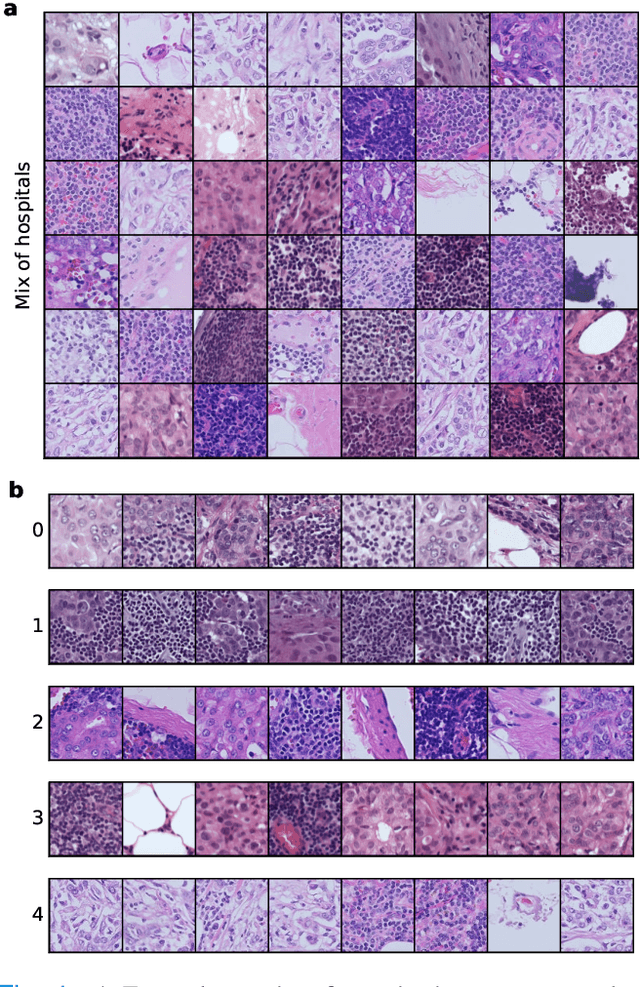
Distribution shifts remain a fundamental problem for the safe application of machine learning systems. If undetected, they may impact the real-world performance of such systems or will at least render original performance claims invalid. In this paper, we focus on the detection of subgroup shifts, a type of distribution shift that can occur when subgroups have a different prevalence during validation compared to the deployment setting. For example, algorithms developed on data from various acquisition settings may be predominantly applied in hospitals with lower quality data acquisition, leading to an inadvertent performance drop. We formulate subgroup shift detection in the framework of statistical hypothesis testing and show that recent state-of-the-art statistical tests can be effectively applied to subgroup shift detection on medical imaging data. We provide synthetic experiments as well as extensive evaluation on clinically meaningful subgroup shifts on histopathology as well as retinal fundus images. We conclude that classifier-based subgroup shift detection tests could be a particularly useful tool for post-market surveillance of deployed ML systems.
AI for Science: An Emerging Agenda
Mar 07, 2023Philipp Berens, Kyle Cranmer, Neil D. Lawrence, Ulrike von Luxburg, Jessica Montgomery
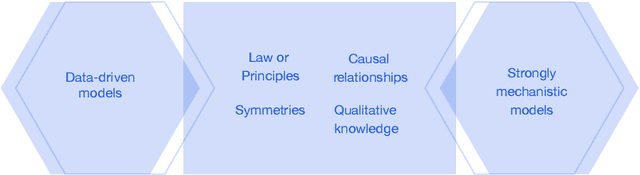
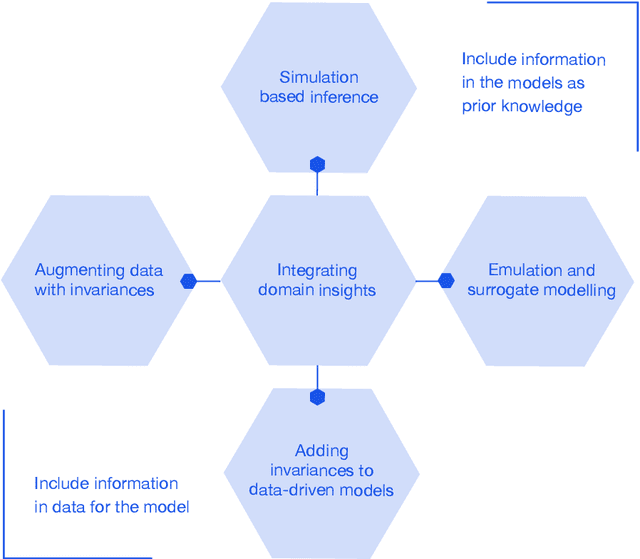
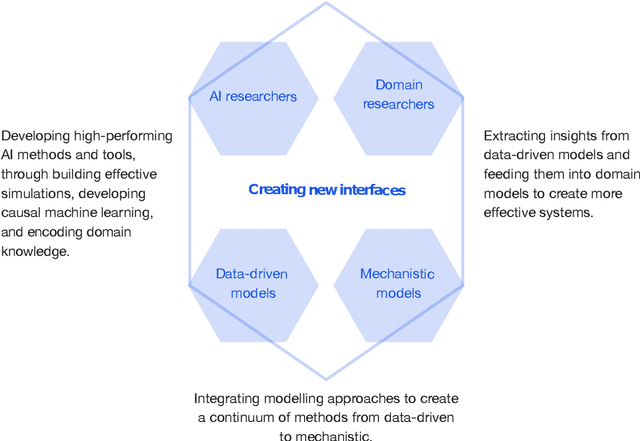
This report documents the programme and the outcomes of Dagstuhl Seminar 22382 "Machine Learning for Science: Bridging Data-Driven and Mechanistic Modelling". Today's scientific challenges are characterised by complexity. Interconnected natural, technological, and human systems are influenced by forces acting across time- and spatial-scales, resulting in complex interactions and emergent behaviours. Understanding these phenomena -- and leveraging scientific advances to deliver innovative solutions to improve society's health, wealth, and well-being -- requires new ways of analysing complex systems. The transformative potential of AI stems from its widespread applicability across disciplines, and will only be achieved through integration across research domains. AI for science is a rendezvous point. It brings together expertise from $\mathrm{AI}$ and application domains; combines modelling knowledge with engineering know-how; and relies on collaboration across disciplines and between humans and machines. Alongside technical advances, the next wave of progress in the field will come from building a community of machine learning researchers, domain experts, citizen scientists, and engineers working together to design and deploy effective AI tools. This report summarises the discussions from the seminar and provides a roadmap to suggest how different communities can collaborate to deliver a new wave of progress in AI and its application for scientific discovery.
Unsupervised visualization of image datasets using contrastive learning
Oct 18, 2022Jan Niklas Böhm, Philipp Berens, Dmitry Kobak
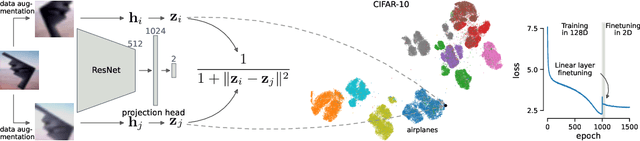
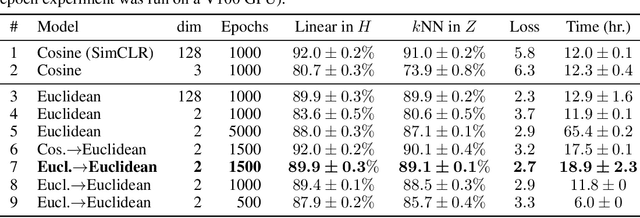
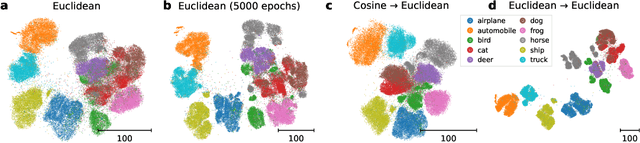
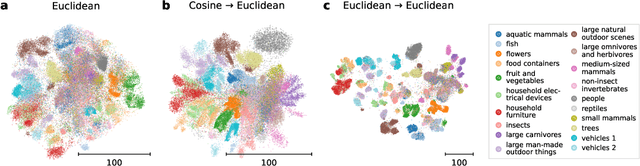
Visualization methods based on the nearest neighbor graph, such as t-SNE or UMAP, are widely used for visualizing high-dimensional data. Yet, these approaches only produce meaningful results if the nearest neighbors themselves are meaningful. For images represented in pixel space this is not the case, as distances in pixel space are often not capturing our sense of similarity and therefore neighbors are not semantically close. This problem can be circumvented by self-supervised approaches based on contrastive learning, such as SimCLR, relying on data augmentation to generate implicit neighbors, but these methods do not produce two-dimensional embeddings suitable for visualization. Here, we present a new method, called t-SimCNE, for unsupervised visualization of image data. T-SimCNE combines ideas from contrastive learning and neighbor embeddings, and trains a parametric mapping from the high-dimensional pixel space into two dimensions. We show that the resulting 2D embeddings achieve classification accuracy comparable to the state-of-the-art high-dimensional SimCLR representations, thus faithfully capturing semantic relationships. Using t-SimCNE, we obtain informative visualizations of the CIFAR-10 and CIFAR-100 datasets, showing rich cluster structure and highlighting artifacts and outliers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge