End-to-end Supervised Prediction of Arbitrary-size Graphs with Partially-Masked Fused Gromov-Wasserstein Matching
Feb 23, 2024Paul Krzakala, Junjie Yang, Rémi Flamary, Florence d'Alché-Buc, Charlotte Laclau, Matthieu Labeau
We present a novel end-to-end deep learning-based approach for Supervised Graph Prediction (SGP). We introduce an original Optimal Transport (OT)-based loss, the Partially-Masked Fused Gromov-Wasserstein loss (PM-FGW), that allows to directly leverage graph representations such as adjacency and feature matrices. PM-FGW exhibits all the desirable properties for SGP: it is node permutation invariant, sub-differentiable and handles graphs of different sizes by comparing their padded representations as well as their masking vectors. Moreover, we present a flexible transformer-based architecture that easily adapts to different types of input data. In the experimental section, three different tasks, a novel and challenging synthetic dataset (image2graph) and two real-world tasks, image2map and fingerprint2molecule - showcase the efficiency and versatility of the approach compared to competitors.
Distributional Reduction: Unifying Dimensionality Reduction and Clustering with Gromov-Wasserstein Projection
Feb 03, 2024Hugues Van Assel, Cédric Vincent-Cuaz, Nicolas Courty, Rémi Flamary, Pascal Frossard, Titouan Vayer
Unsupervised learning aims to capture the underlying structure of potentially large and high-dimensional datasets. Traditionally, this involves using dimensionality reduction methods to project data onto interpretable spaces or organizing points into meaningful clusters. In practice, these methods are used sequentially, without guaranteeing that the clustering aligns well with the conducted dimensionality reduction. In this work, we offer a fresh perspective: that of distributions. Leveraging tools from optimal transport, particularly the Gromov-Wasserstein distance, we unify clustering and dimensionality reduction into a single framework called distributional reduction. This allows us to jointly address clustering and dimensionality reduction with a single optimization problem. Through comprehensive experiments, we highlight the versatility and interpretability of our method and show that it outperforms existing approaches across a variety of image and genomics datasets.
Weakly supervised covariance matrices alignment through Stiefel matrices estimation for MEG applications
Jan 24, 2024Antoine Collas, Rémi Flamary, Alexandre Gramfort
This paper introduces a novel domain adaptation technique for time series data, called Mixing model Stiefel Adaptation (MSA), specifically addressing the challenge of limited labeled signals in the target dataset. Leveraging a domain-dependent mixing model and the optimal transport domain adaptation assumption, we exploit abundant unlabeled data in the target domain to ensure effective prediction by establishing pairwise correspondence with equivalent signal variances between domains. Theoretical foundations are laid for identifying crucial Stiefel matrices, essential for recovering underlying signal variances from a Riemannian representation of observed signal covariances. We propose an integrated cost function that simultaneously learns these matrices, pairwise domain relationships, and a predictor, classifier, or regressor, depending on the task. Applied to neuroscience problems, MSA outperforms recent methods in brain-age regression with task variations using magnetoencephalography (MEG) signals from the Cam-CAN dataset.
Interpolating between Clustering and Dimensionality Reduction with Gromov-Wasserstein
Oct 05, 2023Hugues Van Assel, Cédric Vincent-Cuaz, Titouan Vayer, Rémi Flamary, Nicolas Courty
We present a versatile adaptation of existing dimensionality reduction (DR) objectives, enabling the simultaneous reduction of both sample and feature sizes. Correspondances between input and embedding samples are computed through a semi-relaxed Gromov-Wasserstein optimal transport (OT) problem. When the embedding sample size matches that of the input, our model recovers classical popular DR models. When the embedding's dimensionality is unconstrained, we show that the OT plan delivers a competitive hard clustering. We emphasize the importance of intermediate stages that blend DR and clustering for summarizing real data and apply our method to visualize datasets of images.
Properties of Discrete Sliced Wasserstein Losses
Jul 19, 2023Eloi Tanguy, Rémi Flamary, Julie Delon
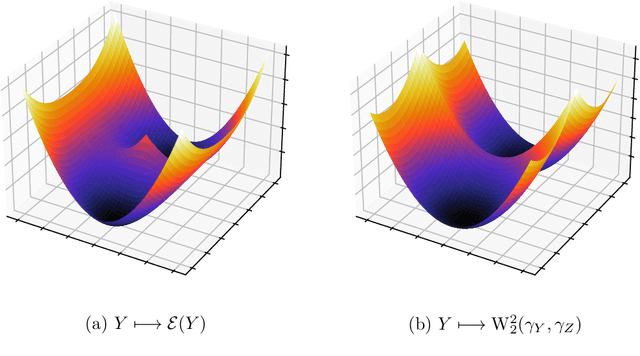
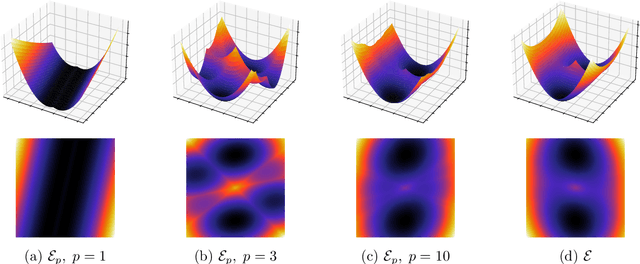
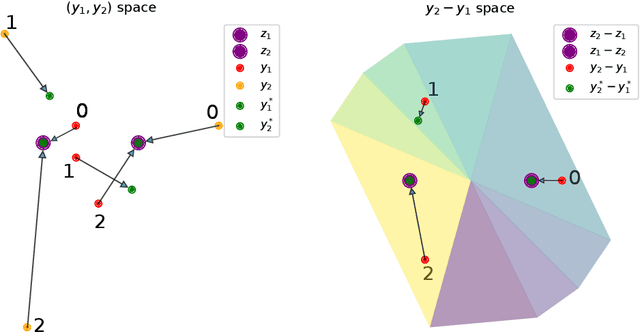
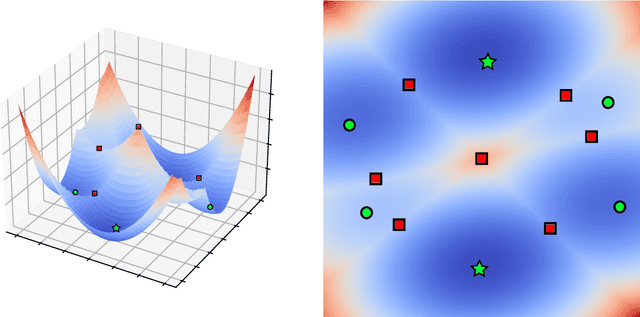
The Sliced Wasserstein (SW) distance has become a popular alternative to the Wasserstein distance for comparing probability measures. Widespread applications include image processing, domain adaptation and generative modelling, where it is common to optimise some parameters in order to minimise SW, which serves as a loss function between discrete probability measures (since measures admitting densities are numerically unattainable). All these optimisation problems bear the same sub-problem, which is minimising the Sliced Wasserstein energy. In this paper we study the properties of $\mathcal{E}: Y \longmapsto \mathrm{SW}_2^2(\gamma_Y, \gamma_Z)$, i.e. the SW distance between two uniform discrete measures with the same amount of points as a function of the support $Y \in \mathbb{R}^{n \times d}$ of one of the measures. We investigate the regularity and optimisation properties of this energy, as well as its Monte-Carlo approximation $\mathcal{E}_p$ (estimating the expectation in SW using only $p$ samples) and show convergence results on the critical points of $\mathcal{E}_p$ to those of $\mathcal{E}$, as well as an almost-sure uniform convergence. Finally, we show that in a certain sense, Stochastic Gradient Descent methods minimising $\mathcal{E}$ and $\mathcal{E}_p$ converge towards (Clarke) critical points of these energies.
Convolutional Monge Mapping Normalization for learning on biosignals
May 31, 2023Théo Gnassounou, Rémi Flamary, Alexandre Gramfort
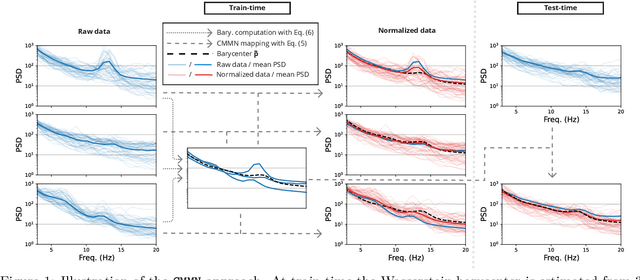
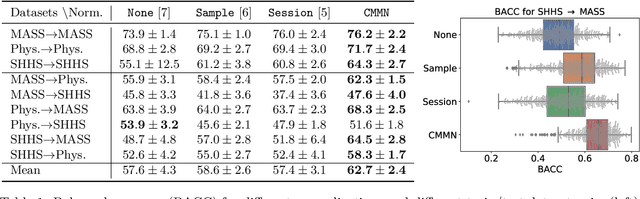
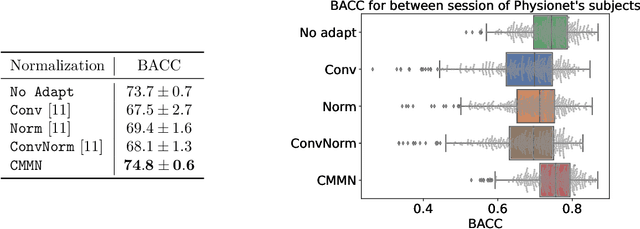
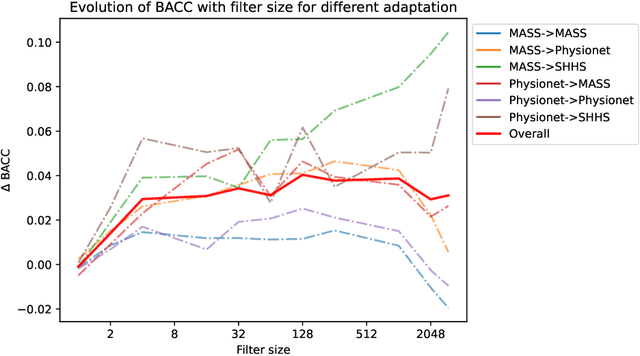
In many machine learning applications on signals and biomedical data, especially electroencephalogram (EEG), one major challenge is the variability of the data across subjects, sessions, and hardware devices. In this work, we propose a new method called Convolutional Monge Mapping Normalization (CMMN), which consists in filtering the signals in order to adapt their power spectrum density (PSD) to a Wasserstein barycenter estimated on training data. CMMN relies on novel closed-form solutions for optimal transport mappings and barycenters and provides individual test time adaptation to new data without needing to retrain a prediction model. Numerical experiments on sleep EEG data show that CMMN leads to significant and consistent performance gains independent from the neural network architecture when adapting between subjects, sessions, and even datasets collected with different hardware. Notably our performance gain is on par with much more numerically intensive Domain Adaptation (DA) methods and can be used in conjunction with those for even better performances.
SNEkhorn: Dimension Reduction with Symmetric Entropic Affinities
May 23, 2023Hugues Van Assel, Titouan Vayer, Rémi Flamary, Nicolas Courty
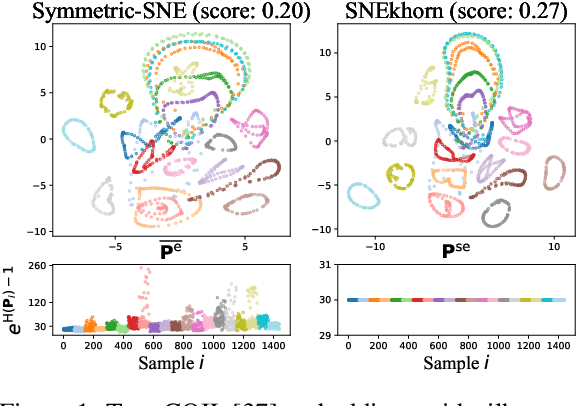
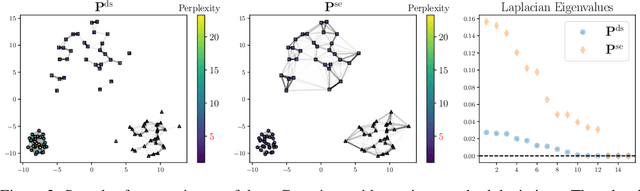
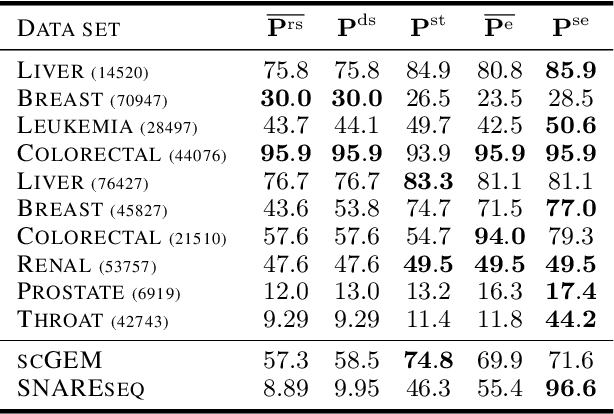
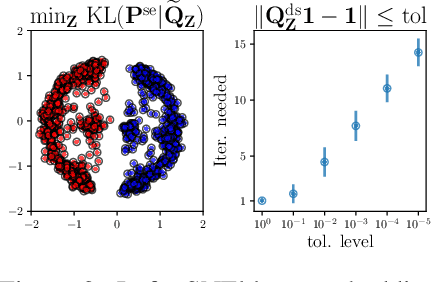
Many approaches in machine learning rely on a weighted graph to encode the similarities between samples in a dataset. Entropic affinities (EAs), which are notably used in the popular Dimensionality Reduction (DR) algorithm t-SNE, are particular instances of such graphs. To ensure robustness to heterogeneous sampling densities, EAs assign a kernel bandwidth parameter to every sample in such a way that the entropy of each row in the affinity matrix is kept constant at a specific value, whose exponential is known as perplexity. EAs are inherently asymmetric and row-wise stochastic, but they are used in DR approaches after undergoing heuristic symmetrization methods that violate both the row-wise constant entropy and stochasticity properties. In this work, we uncover a novel characterization of EA as an optimal transport problem, allowing a natural symmetrization that can be computed efficiently using dual ascent. The corresponding novel affinity matrix derives advantages from symmetric doubly stochastic normalization in terms of clustering performance, while also effectively controlling the entropy of each row thus making it particularly robust to varying noise levels. Following, we present a new DR algorithm, SNEkhorn, that leverages this new affinity matrix. We show its clear superiority to state-of-the-art approaches with several indicators on both synthetic and real-world datasets.
Entropic Wasserstein Component Analysis
Mar 09, 2023Antoine Collas, Titouan Vayer, Rémi Flamary, Arnaud Breloy
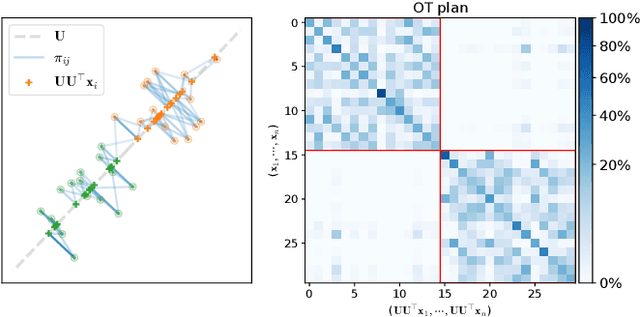

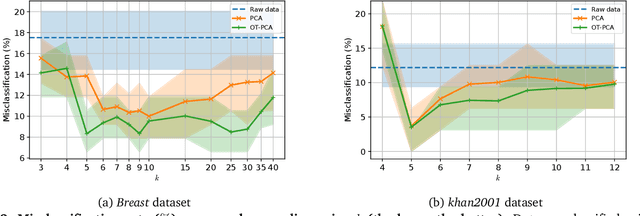
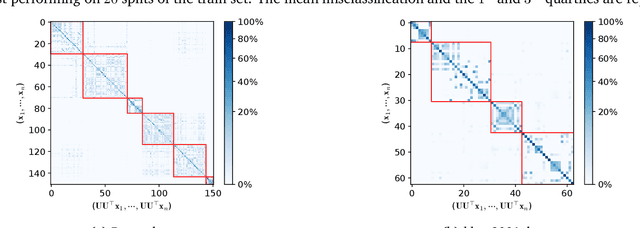
Dimension reduction (DR) methods provide systematic approaches for analyzing high-dimensional data. A key requirement for DR is to incorporate global dependencies among original and embedded samples while preserving clusters in the embedding space. To achieve this, we combine the principles of optimal transport (OT) and principal component analysis (PCA). Our method seeks the best linear subspace that minimizes reconstruction error using entropic OT, which naturally encodes the neighborhood information of the samples. From an algorithmic standpoint, we propose an efficient block-majorization-minimization solver over the Stiefel manifold. Our experimental results demonstrate that our approach can effectively preserve high-dimensional clusters, leading to more interpretable and effective embeddings. Python code of the algorithms and experiments is available online.
Aligning individual brains with Fused Unbalanced Gromov-Wasserstein
Jun 19, 2022Alexis Thual, Huy Tran, Tatiana Zemskova, Nicolas Courty, Rémi Flamary, Stanislas Dehaene, Bertrand Thirion
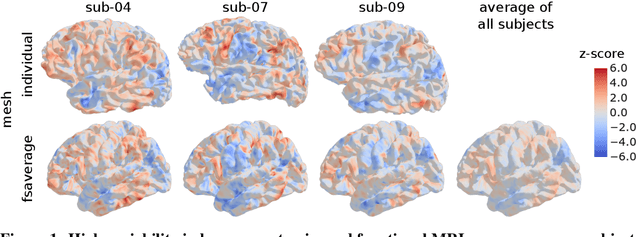
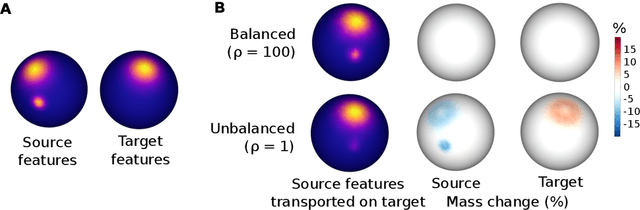
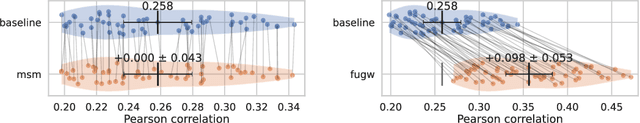
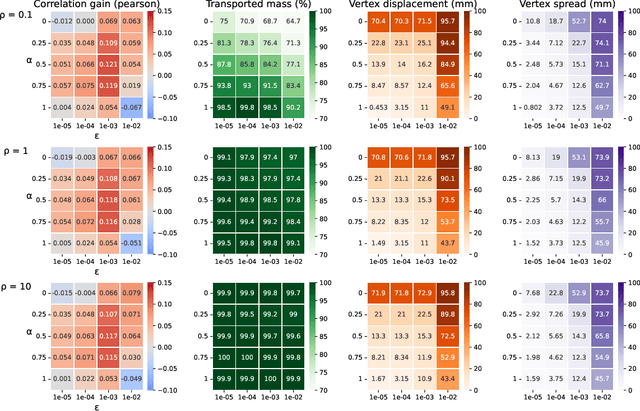
Individual brains vary in both anatomy and functional organization, even within a given species. Inter-individual variability is a major impediment when trying to draw generalizable conclusions from neuroimaging data collected on groups of subjects. Current co-registration procedures rely on limited data, and thus lead to very coarse inter-subject alignments. In this work, we present a novel method for inter-subject alignment based on Optimal Transport, denoted as Fused Unbalanced Gromov Wasserstein (FUGW). The method aligns cortical surfaces based on the similarity of their functional signatures in response to a variety of stimulation settings, while penalizing large deformations of individual topographic organization. We demonstrate that FUGW is well-suited for whole-brain landmark-free alignment. The unbalanced feature allows to deal with the fact that functional areas vary in size across subjects. Our results show that FUGW alignment significantly increases between-subject correlation of activity for independent functional data, and leads to more precise mapping at the group level.
Unbalanced CO-Optimal Transport
May 31, 2022Quang Huy Tran, Hicham Janati, Nicolas Courty, Rémi Flamary, Ievgen Redko, Pinar Demetci, Ritambhara Singh
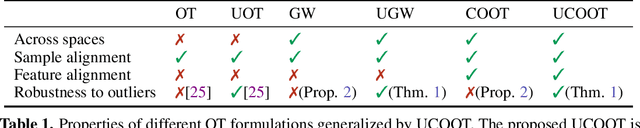
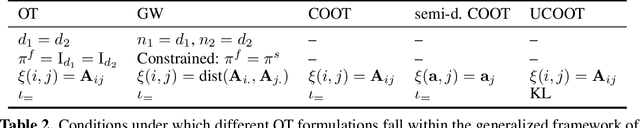

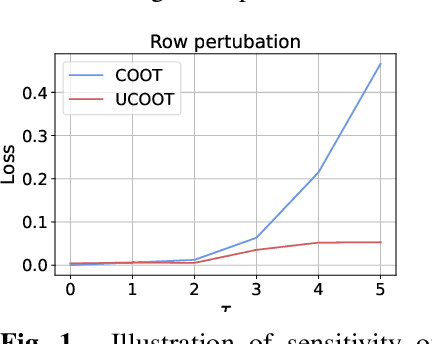


Optimal transport (OT) compares probability distributions by computing a meaningful alignment between their samples. CO-optimal transport (COOT) takes this comparison further by inferring an alignment between features as well. While this approach leads to better alignments and generalizes both OT and Gromov-Wasserstein distances, we provide a theoretical result showing that it is sensitive to outliers that are omnipresent in real-world data. This prompts us to propose unbalanced COOT for which we provably show its robustness to noise in the compared datasets. To the best of our knowledge, this is the first such result for OT methods in incomparable spaces. With this result in hand, we provide empirical evidence of this robustness for the challenging tasks of heterogeneous domain adaptation with and without varying proportions of classes and simultaneous alignment of samples and features across single-cell measurements.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge