CATS: Enhancing Multivariate Time Series Forecasting by Constructing Auxiliary Time Series as Exogenous Variables
Mar 04, 2024Jiecheng Lu, Xu Han, Yan Sun, Shihao Yang
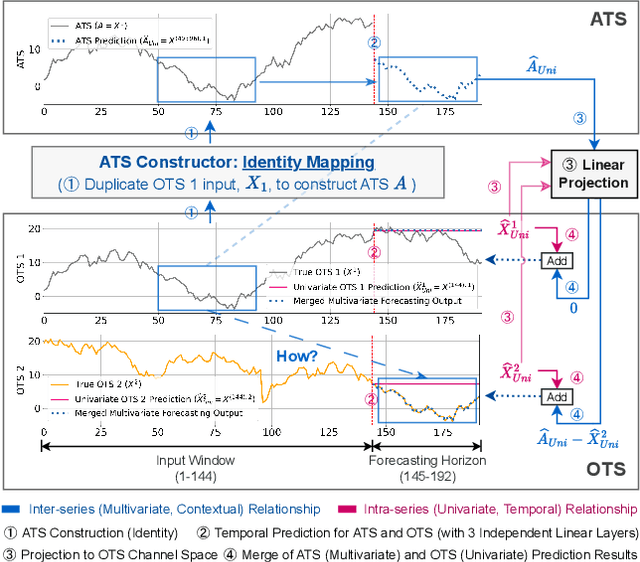
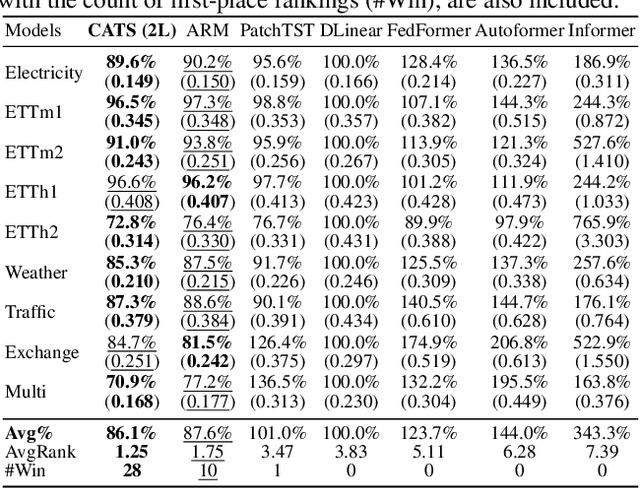
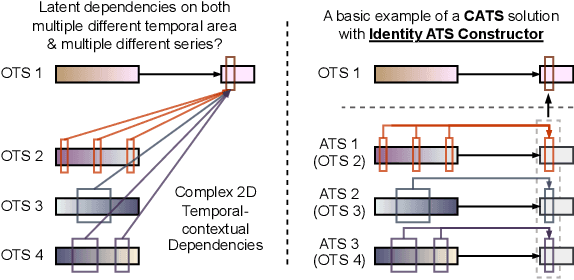

For Multivariate Time Series Forecasting (MTSF), recent deep learning applications show that univariate models frequently outperform multivariate ones. To address the difficiency in multivariate models, we introduce a method to Construct Auxiliary Time Series (CATS) that functions like a 2D temporal-contextual attention mechanism, which generates Auxiliary Time Series (ATS) from Original Time Series (OTS) to effectively represent and incorporate inter-series relationships for forecasting. Key principles of ATS - continuity, sparsity, and variability - are identified and implemented through different modules. Even with a basic 2-layer MLP as core predictor, CATS achieves state-of-the-art, significantly reducing complexity and parameters compared to previous multivariate models, marking it an efficient and transferable MTSF solution.
Identification of Craving Maps among Marijuana Users via Analysis of Functional Brain Networks with High-Order Attention Graph Neural Networks
Mar 04, 2024Jun-En Ding, Shihao Yang, Anna Zilverstand, Feng Liu




The consumption of high doses of marijuana can have significant psychological and social impacts. In this study, we propose an interpretable novel framework called the HOGAB (High-Order Graph Attention Neural Networks) model for addictive Marijuana classification and analysis of the localized network clusters that demonstrated abnormal brain activities among chronic marijuana users. The HOGAB integrates dynamic intrinsic functional networks with LSTM technology to capture temporal patterns in fMRI time series of marijuana users. We employed the high-order attention module in neighborhood nodes for information fusion and message passing, enhancing community clustering analysis for long-term marijuana users. Furthermore, we improve the overall classification ability of the model by incorporating attention mechanisms, achieving an AUC of 85.1% and an accuracy of 80.7% in classification, higher than the comparison algoirthms. Specifically, we identified the most relevant subnetworks and cognitive regions that are influenced by persistent marijuana usage, revealing that chronic marijuana consumption adversely affects cognitive control, particularly within the Dorsal Attention and Frontoparietal networks, which are essential for attentional, cognitive and higher cognitive functions. The results show that our proposed model is capable of accurately predicting craving bahavior and identifying brain maps associated with long-term cravings, and thus pinpointing brain regions that are important for analysis.
ARM: Refining Multivariate Forecasting with Adaptive Temporal-Contextual Learning
Oct 14, 2023Jiecheng Lu, Xu Han, Shihao Yang
Long-term time series forecasting (LTSF) is important for various domains but is confronted by challenges in handling the complex temporal-contextual relationships. As multivariate input models underperforming some recent univariate counterparts, we posit that the issue lies in the inefficiency of existing multivariate LTSF Transformers to model series-wise relationships: the characteristic differences between series are often captured incorrectly. To address this, we introduce ARM: a multivariate temporal-contextual adaptive learning method, which is an enhanced architecture specifically designed for multivariate LTSF modelling. ARM employs Adaptive Univariate Effect Learning (AUEL), Random Dropping (RD) training strategy, and Multi-kernel Local Smoothing (MKLS), to better handle individual series temporal patterns and correctly learn inter-series dependencies. ARM demonstrates superior performance on multiple benchmarks without significantly increasing computational costs compared to vanilla Transformer, thereby advancing the state-of-the-art in LTSF. ARM is also generally applicable to other LTSF architecture beyond vanilla Transformer.
Deep Attention Q-Network for Personalized Treatment Recommendation
Jul 04, 2023Simin Ma, Junghwan Lee, Nicoleta Serban, Shihao Yang




Tailoring treatment for individual patients is crucial yet challenging in order to achieve optimal healthcare outcomes. Recent advances in reinforcement learning offer promising personalized treatment recommendations; however, they rely solely on current patient observations (vital signs, demographics) as the patient's state, which may not accurately represent the true health status of the patient. This limitation hampers policy learning and evaluation, ultimately limiting treatment effectiveness. In this study, we propose the Deep Attention Q-Network for personalized treatment recommendations, utilizing the Transformer architecture within a deep reinforcement learning framework to efficiently incorporate all past patient observations. We evaluated the model on real-world sepsis and acute hypotension cohorts, demonstrating its superiority to state-of-the-art models. The source code for our model is available at https://github.com/stevenmsm/RL-ICU-DAQN.
Inference of Nonlinear Partial Differential Equations via Constrained Gaussian Processes
Dec 22, 2022Zhaohui Li, Shihao Yang, Jeff Wu




Partial differential equations (PDEs) are widely used for description of physical and engineering phenomena. Some key parameters involved in PDEs, which represents certain physical properties with important scientific interpretations, are difficult or even impossible to be measured directly. Estimation of these parameters from noisy and sparse experimental data of related physical quantities is an important task. Many methods for PDE parameter inference involve a large number of evaluations of numerical solution of PDE through algorithms such as finite element method, which can be time-consuming especially for nonlinear PDEs. In this paper, we propose a novel method for estimating unknown parameters in PDEs, called PDE-Informed Gaussian Process Inference (PIGPI). Through modeling the PDE solution as a Gaussian process (GP), we derive the manifold constraints induced by the (linear) PDE structure such that under the constraints, the GP satisfies the PDE. For nonlinear PDEs, we propose an augmentation method that transfers the nonlinear PDE into an equivalent PDE system linear in all derivatives that our PIGPI can handle. PIGPI can be applied to multi-dimensional PDE systems and PDE systems with unobserved components. The method completely bypasses the numerical solver for PDE, thus achieving drastic savings in computation time, especially for nonlinear PDEs. Moreover, the PIGPI method can give the uncertainty quantification for both the unknown parameters and the PDE solution. The proposed method is demonstrated by several application examples from different areas.
COVID-19 Hospitalizations Forecasts Using Internet Search Data
Feb 03, 2022Tao Wang, Simin Ma, Soobin Baek, Shihao Yang




As the COVID-19 spread over the globe and new variants of COVID-19 keep occurring, reliable real-time forecasts of COVID-19 hospitalizations are critical for public health decision on medical resources allocations such as ICU beds, ventilators, and personnel to prepare for the surge of COVID-19 pandemics. Inspired by the strong association between public search behavior and hospitalization admission, we extended previously-proposed influenza tracking model, ARGO (AutoRegression with GOogle search data), to predict future 2-week national and state-level COVID-19 new hospital admissions. Leveraging the COVID-19 related time series information and Google search data, our method is able to robustly capture new COVID-19 variants' surges, and self-correct at both national and state level. Based on our retrospective out-of-sample evaluation over 12-month comparison period, our method achieves on average 15\% error reduction over the best alternative models collected from COVID-19 forecast hub. Overall, we showed that our method is flexible, self-correcting, robust, accurate, and interpretable, making it a potentially powerful tool to assist health-care officials and decision making for the current and future infectious disease outbreak.
Early Detection of COVID-19 Hotspots Using Spatio-Temporal Data
May 31, 2021Shixiang Zhu, Alexander Bukharin, Liyan Xie, Shihao Yang, Pinar Keskinocak, Yao Xie




Recently, the Centers for Disease Control and Prevention (CDC) has worked with other federal agencies to identify counties with increasing coronavirus disease 2019 (COVID-19) incidence (hotspots) and offers support to local health departments to limit the spread of the disease. Understanding the spatio-temporal dynamics of hotspot events is of great importance to support policy decisions and prevent large-scale outbreaks. This paper presents a spatio-temporal Bayesian framework for early detection of COVID-19 hotspots (at the county level) in the United States. We assume both the observed number of cases and hotspots depend on a class of latent random variables, which encode the underlying spatio-temporal dynamics of the transmission of COVID-19. Such latent variables follow a zero-mean Gaussian process, whose covariance is specified by a non-stationary kernel function. The most salient feature of our kernel function is that deep neural networks are introduced to enhance the model's representative power while still enjoying the interpretability of the kernel. We derive a sparse model and fit the model using a variational learning strategy to circumvent the computational intractability for large data sets. Our model demonstrates better interpretability and superior hotspot-detection performance compared to other baseline methods.
MAGI-X: Manifold-Constrained Gaussian Process Inference for Unknown System Dynamics
May 28, 2021Chaofan Huang, Simin Ma, Shihao Yang




Ordinary differential equations (ODEs), commonly used to characterize the dynamic systems, are difficult to propose in closed-form for many complicated scientific applications, even with the help of domain expert. We propose a fast and accurate data-driven method, MAGI-X, to learn the unknown dynamic from the observation data in a non-parametric fashion, without the need of any domain knowledge. Unlike the existing methods that mainly rely on the costly numerical integration, MAGI-X utilizes the powerful functional approximator of neural network to learn the unknown nonlinear dynamic within the MAnifold-constrained Gaussian process Inference (MAGI) framework that completely circumvents the numerical integration. Comparing against the state-of-the-art methods on three realistic examples, MAGI-X achieves competitive accuracy in both fitting and forecasting while only taking a fraction of computational time. Moreover, MAGI-X provides practical solution for the inference of partial observed systems, which no previous method is able to handle.
COUnty aggRegation mixup AuGmEntation (COURAGE) COVID-19 Prediction
May 03, 2021Siawpeng Er, Shihao Yang, Tuo Zhao




The global spread of COVID-19, the disease caused by the novel coronavirus SARS-CoV-2, has cast a significant threat to mankind. As the COVID-19 situation continues to evolve, predicting localized disease severity is crucial for advanced resource allocation. This paper proposes a method named COURAGE (COUnty aggRegation mixup AuGmEntation) to generate a short-term prediction of 2-week-ahead COVID-19 related deaths for each county in the United States, leveraging modern deep learning techniques. Specifically, our method adopts a self-attention model from Natural Language Processing, known as the transformer model, to capture both short-term and long-term dependencies within the time series while enjoying computational efficiency. Our model fully utilizes publicly available information of COVID-19 related confirmed cases, deaths, community mobility trends and demographic information, and can produce state-level prediction as an aggregation of the corresponding county-level predictions. Our numerical experiments demonstrate that our model achieves the state-of-the-art performance among the publicly available benchmark models.
Accurate estimation of influenza epidemics using Google search data via ARGO
Nov 16, 2015Shihao Yang, Mauricio Santillana, S. C. Kou



Accurate real-time tracking of influenza outbreaks helps public health officials make timely and meaningful decisions that could save lives. We propose an influenza tracking model, ARGO (AutoRegression with GOogle search data), that uses publicly available online search data. In addition to having a rigorous statistical foundation, ARGO outperforms all previously available Google-search-based tracking models, including the latest version of Google Flu Trends, even though it uses only low-quality search data as input from publicly available Google Trends and Google Correlate websites. ARGO not only incorporates the seasonality in influenza epidemics but also captures changes in people's online search behavior over time. ARGO is also flexible, self-correcting, robust, and scalable, making it a potentially powerful tool that can be used for real-time tracking of other social events at multiple temporal and spatial resolutions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge