Towards SAMBA: Segment Anything Model for Brain Tumor Segmentation in Sub-Sharan African Populations
Dec 19, 2023Mohannad Barakat, Noha Magdy, Jjuuko George William, Ethel Phiri, Raymond Confidence, Dong Zhang, Udunna C Anazodo
Gliomas, the most prevalent primary brain tumors, require precise segmentation for diagnosis and treatment planning. However, this task poses significant challenges, particularly in the African population, were limited access to high-quality imaging data hampers algorithm performance. In this study, we propose an innovative approach combining the Segment Anything Model (SAM) and a voting network for multi-modal glioma segmentation. By fine-tuning SAM with bounding box-guided prompts (SAMBA), we adapt the model to the complexities of African datasets. Our ensemble strategy, utilizing multiple modalities and views, produces a robust consensus segmentation, addressing intra-tumoral heterogeneity. Although the low quality of scans presents difficulties, our methodology has the potential to profoundly impact clinical practice in resource-limited settings such as Africa, improving treatment decisions and advancing neuro-oncology research. Furthermore, successful application to other brain tumor types and lesions in the future holds promise for a broader transformation in neurological imaging, improving healthcare outcomes across all settings. This study was conducted on the Brain Tumor Segmentation (BraTS) Challenge Africa (BraTS-Africa) dataset, which provides a valuable resource for addressing challenges specific to resource-limited settings, particularly the African population, and facilitating the development of effective and more generalizable segmentation algorithms. To illustrate our approach's potential, our experiments on the BraTS-Africa dataset yielded compelling results, with SAM attaining a Dice coefficient of 86.6 for binary segmentation and 60.4 for multi-class segmentation.
The Brain Tumor Segmentation (BraTS) Challenge 2023: Glioma Segmentation in Sub-Saharan Africa Patient Population (BraTS-Africa)
May 30, 2023Maruf Adewole, Jeffrey D. Rudie, Anu Gbadamosi, Oluyemisi Toyobo, Confidence Raymond, Dong Zhang, Olubukola Omidiji, Rachel Akinola, Mohammad Abba Suwaid, Adaobi Emegoakor, Nancy Ojo, Kenneth Aguh, Chinasa Kalaiwo, Gabriel Babatunde, Afolabi Ogunleye, Yewande Gbadamosi, Kator Iorpagher, Evan Calabrese, Mariam Aboian, Marius Linguraru, Jake Albrecht, Benedikt Wiestler, Florian Kofler, Anastasia Janas, Dominic LaBella, Anahita Fathi Kzerooni, Hongwei Bran Li, Juan Eugenio Iglesias, Keyvan Farahani, James Eddy, Timothy Bergquist, Verena Chung, Russell Takeshi Shinohara, Walter Wiggins, Zachary Reitman, Chunhao Wang, Xinyang Liu, Zhifan Jiang, Ariana Familiar, Koen Van Leemput, Christina Bukas, Maire Piraud, Gian-Marco Conte, Elaine Johansson, Zeke Meier, Bjoern H Menze, Ujjwal Baid, Spyridon Bakas, Farouk Dako, Abiodun Fatade, Udunna C Anazodo
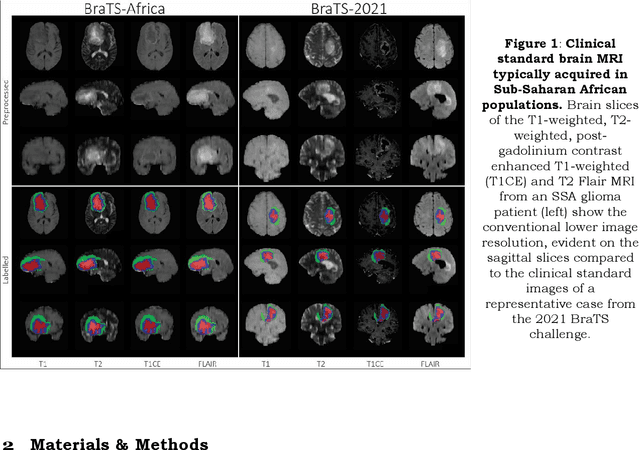
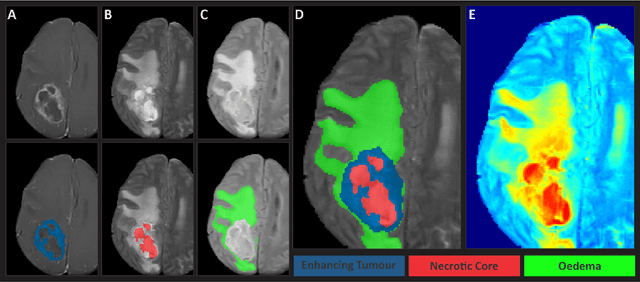
Gliomas are the most common type of primary brain tumors. Although gliomas are relatively rare, they are among the deadliest types of cancer, with a survival rate of less than 2 years after diagnosis. Gliomas are challenging to diagnose, hard to treat and inherently resistant to conventional therapy. Years of extensive research to improve diagnosis and treatment of gliomas have decreased mortality rates across the Global North, while chances of survival among individuals in low- and middle-income countries (LMICs) remain unchanged and are significantly worse in Sub-Saharan Africa (SSA) populations. Long-term survival with glioma is associated with the identification of appropriate pathological features on brain MRI and confirmation by histopathology. Since 2012, the Brain Tumor Segmentation (BraTS) Challenge have evaluated state-of-the-art machine learning methods to detect, characterize, and classify gliomas. However, it is unclear if the state-of-the-art methods can be widely implemented in SSA given the extensive use of lower-quality MRI technology, which produces poor image contrast and resolution and more importantly, the propensity for late presentation of disease at advanced stages as well as the unique characteristics of gliomas in SSA (i.e., suspected higher rates of gliomatosis cerebri). Thus, the BraTS-Africa Challenge provides a unique opportunity to include brain MRI glioma cases from SSA in global efforts through the BraTS Challenge to develop and evaluate computer-aided-diagnostic (CAD) methods for the detection and characterization of glioma in resource-limited settings, where the potential for CAD tools to transform healthcare are more likely.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge