NDGGNET-A Node Independent Gate based Graph Neural Networks
May 11, 2022Ye Tang, Xuesong Yang, Xinrui Liu, Xiwei Zhao, Zhangang Lin, Changping Peng


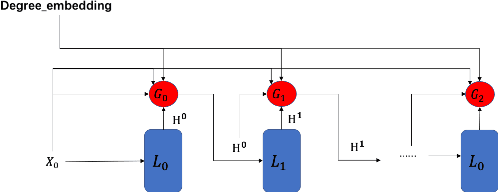

Graph Neural Networks (GNNs) is an architecture for structural data, and has been adopted in a mass of tasks and achieved fabulous results, such as link prediction, node classification, graph classification and so on. Generally, for a certain node in a given graph, a traditional GNN layer can be regarded as an aggregation from one-hop neighbors, thus a set of stacked layers are able to fetch and update node status within multi-hops. For nodes with sparse connectivity, it is difficult to obtain enough information through a single GNN layer as not only there are only few nodes directly connected to them but also can not propagate the high-order neighbor information. However, as the number of layer increases, the GNN model is prone to over-smooth for nodes with the dense connectivity, which resulting in the decrease of accuracy. To tackle this issue, in this thesis, we define a novel framework that allows the normal GNN model to accommodate more layers. Specifically, a node-degree based gate is employed to adjust weight of layers dynamically, that try to enhance the information aggregation ability and reduce the probability of over-smoothing. Experimental results show that our proposed model can effectively increase the model depth and perform well on several datasets.
LIAF-Net: Leaky Integrate and Analog Fire Network for Lightweight and Efficient Spatiotemporal Information Processing
Nov 12, 2020Zhenzhi Wu, Hehui Zhang, Yihan Lin, Guoqi Li, Meng Wang, Ye Tang
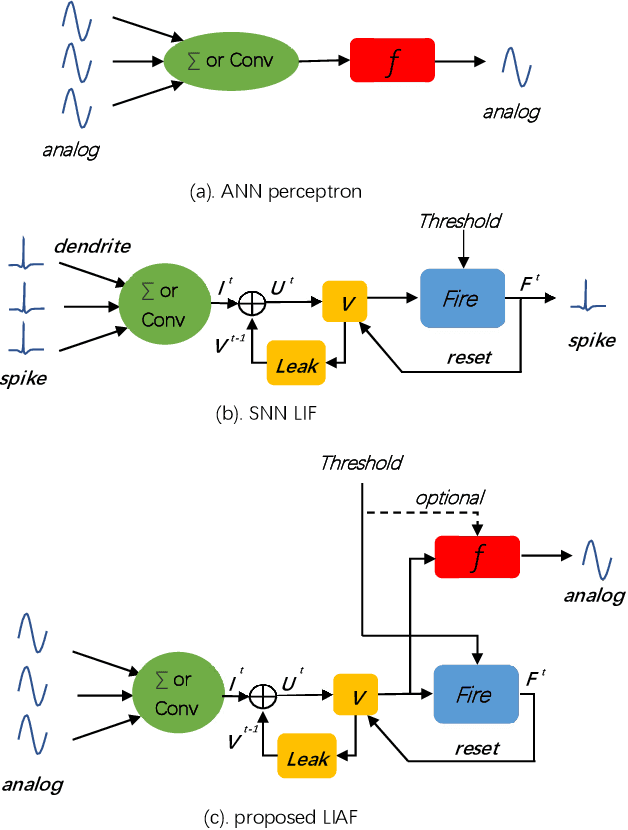
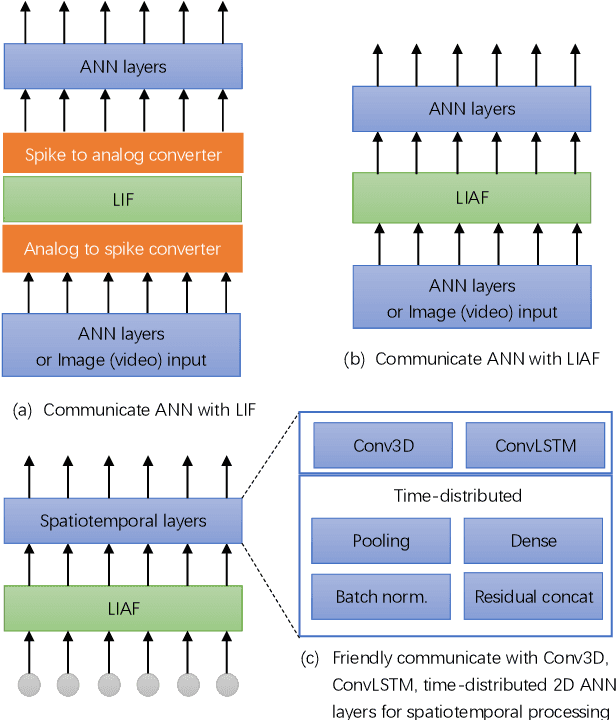

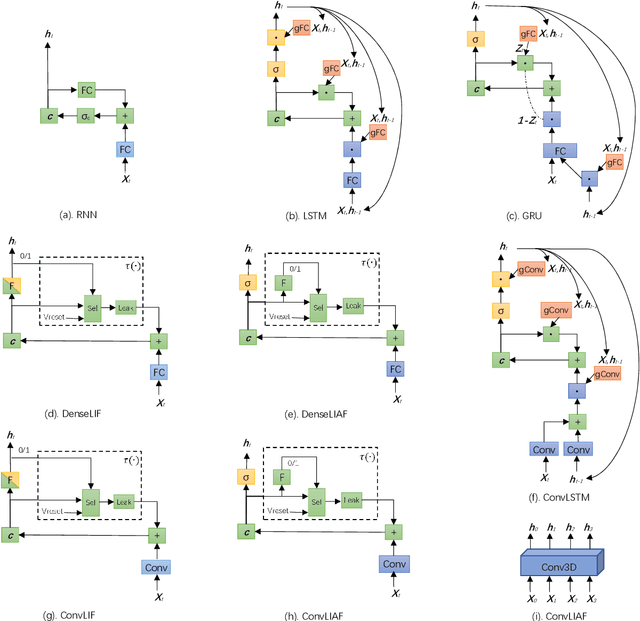
Spiking neural networks (SNNs) based on Leaky Integrate and Fire (LIF) model have been applied to energy-efficient temporal and spatiotemporal processing tasks. Thanks to the bio-plausible neuronal dynamics and simplicity, LIF-SNN benefits from event-driven processing, however, usually faces the embarrassment of reduced performance. This may because in LIF-SNN the neurons transmit information via spikes. To address this issue, in this work, we propose a Leaky Integrate and Analog Fire (LIAF) neuron model, so that analog values can be transmitted among neurons, and a deep network termed as LIAF-Net is built on it for efficient spatiotemporal processing. In the temporal domain, LIAF follows the traditional LIF dynamics to maintain its temporal processing capability. In the spatial domain, LIAF is able to integrate spatial information through convolutional integration or fully-connected integration. As a spatiotemporal layer, LIAF can also be used with traditional artificial neural network (ANN) layers jointly. Experiment results indicate that LIAF-Net achieves comparable performance to Gated Recurrent Unit (GRU) and Long short-term memory (LSTM) on bAbI Question Answering (QA) tasks, and achieves state-of-the-art performance on spatiotemporal Dynamic Vision Sensor (DVS) datasets, including MNIST-DVS, CIFAR10-DVS and DVS128 Gesture, with much less number of synaptic weights and computational overhead compared with traditional networks built by LSTM, GRU, Convolutional LSTM (ConvLSTM) or 3D convolution (Conv3D). Compared with traditional LIF-SNN, LIAF-Net also shows dramatic accuracy gain on all these experiments. In conclusion, LIAF-Net provides a framework combining the advantages of both ANNs and SNNs for lightweight and efficient spatiotemporal information processing.
Unsupervised Learning for Cell-level Visual Representation in Histopathology Images with Generative Adversarial Networks
Jul 07, 2018Bo Hu, Ye Tang, Eric I-Chao Chang, Yubo Fan, Maode Lai, Yan Xu




The visual attributes of cells, such as the nuclear morphology and chromatin openness, are critical for histopathology image analysis. By learning cell-level visual representation, we can obtain a rich mix of features that are highly reusable for various tasks, such as cell-level classification, nuclei segmentation, and cell counting. In this paper, we propose a unified generative adversarial networks architecture with a new formulation of loss to perform robust cell-level visual representation learning in an unsupervised setting. Our model is not only label-free and easily trained but also capable of cell-level unsupervised classification with interpretable visualization, which achieves promising results in the unsupervised classification of bone marrow cellular components. Based on the proposed cell-level visual representation learning, we further develop a pipeline that exploits the varieties of cellular elements to perform histopathology image classification, the advantages of which are demonstrated on bone marrow datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge