Lancet: Accelerating Mixture-of-Experts Training via Whole Graph Computation-Communication Overlapping
Apr 30, 2024Chenyu Jiang, Ye Tian, Zhen Jia, Shuai Zheng, Chuan Wu, Yida Wang
The Mixture-of-Expert (MoE) technique plays a crucial role in expanding the size of DNN model parameters. However, it faces the challenge of extended all-to-all communication latency during the training process. Existing methods attempt to mitigate this issue by overlapping all-to-all with expert computation. Yet, these methods frequently fall short of achieving sufficient overlap, consequently restricting the potential for performance enhancements. In our study, we extend the scope of this challenge by considering overlap at the broader training graph level. During the forward pass, we enable non-MoE computations to overlap with all-to-all through careful partitioning and pipelining. In the backward pass, we achieve overlap with all-to-all by scheduling gradient weight computations. We implement these techniques in Lancet, a system using compiler-based optimization to automatically enhance MoE model training. Our extensive evaluation reveals that Lancet significantly reduces the time devoted to non-overlapping communication, by as much as 77%. Moreover, it achieves a notable end-to-end speedup of up to 1.3 times when compared to the state-of-the-art solutions.
High-fidelity Endoscopic Image Synthesis by Utilizing Depth-guided Neural Surfaces
Apr 20, 2024Baoru Huang, Yida Wang, Anh Nguyen, Daniel Elson, Francisco Vasconcelos, Danail Stoyanov
In surgical oncology, screening colonoscopy plays a pivotal role in providing diagnostic assistance, such as biopsy, and facilitating surgical navigation, particularly in polyp detection. Computer-assisted endoscopic surgery has recently gained attention and amalgamated various 3D computer vision techniques, including camera localization, depth estimation, surface reconstruction, etc. Neural Radiance Fields (NeRFs) and Neural Implicit Surfaces (NeuS) have emerged as promising methodologies for deriving accurate 3D surface models from sets of registered images, addressing the limitations of existing colon reconstruction approaches stemming from constrained camera movement. However, the inadequate tissue texture representation and confused scale problem in monocular colonoscopic image reconstruction still impede the progress of the final rendering results. In this paper, we introduce a novel method for colon section reconstruction by leveraging NeuS applied to endoscopic images, supplemented by a single frame of depth map. Notably, we pioneered the exploration of utilizing only one frame depth map in photorealistic reconstruction and neural rendering applications while this single depth map can be easily obtainable from other monocular depth estimation networks with an object scale. Through rigorous experimentation and validation on phantom imagery, our approach demonstrates exceptional accuracy in completely rendering colon sections, even capturing unseen portions of the surface. This breakthrough opens avenues for achieving stable and consistently scaled reconstructions, promising enhanced quality in cancer screening procedures and treatment interventions.
HLAT: High-quality Large Language Model Pre-trained on AWS Trainium
Apr 16, 2024Haozheng Fan, Hao Zhou, Guangtai Huang, Parameswaran Raman, Xinwei Fu, Gaurav Gupta, Dhananjay Ram, Yida Wang, Jun Huan
Getting large language models (LLMs) to perform well on the downstream tasks requires pre-training over trillions of tokens. This typically demands a large number of powerful computational devices in addition to a stable distributed training framework to accelerate the training. The growing number of applications leveraging AI/ML had led to a scarcity of the expensive conventional accelerators (such as GPUs), which begs the need for the alternative specialized-accelerators that are scalable and cost-efficient. AWS Trainium is the second-generation machine learning accelerator that has been purposely built for training large deep learning models. Its corresponding instance, Amazon EC2 trn1, is an alternative to GPU instances for LLM training. However, training LLMs with billions of parameters on trn1 is challenging due to its relatively nascent software ecosystem. In this paper, we showcase HLAT: a 7 billion parameter decoder-only LLM pre-trained using trn1 instances over 1.8 trillion tokens. The performance of HLAT is benchmarked against popular open source baseline models including LLaMA and OpenLLaMA, which have been trained on NVIDIA GPUs and Google TPUs, respectively. On various evaluation tasks, we show that HLAT achieves model quality on par with the baselines. We also share the best practice of using the Neuron Distributed Training Library (NDTL), a customized distributed training library for AWS Trainium to achieve efficient training. Our work demonstrates that AWS Trainium powered by the NDTL is able to successfully pre-train state-of-the-art LLM models with high performance and cost-effectiveness.
ContrastDiagnosis: Enhancing Interpretability in Lung Nodule Diagnosis Using Contrastive Learning
Mar 08, 2024Chenglong Wang, Yinqiao Yi, Yida Wang, Chengxiu Zhang, Yun Liu, Kensaku Mori, Mei Yuan, Guang Yang
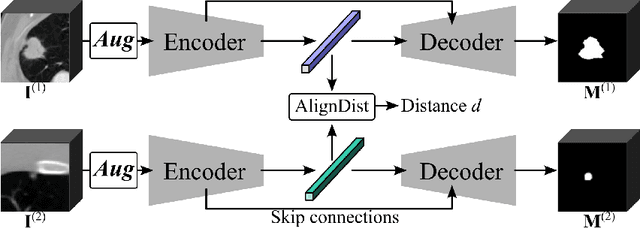
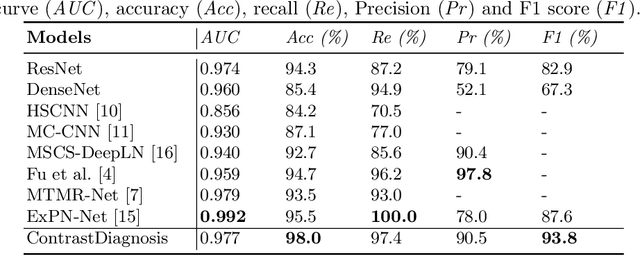
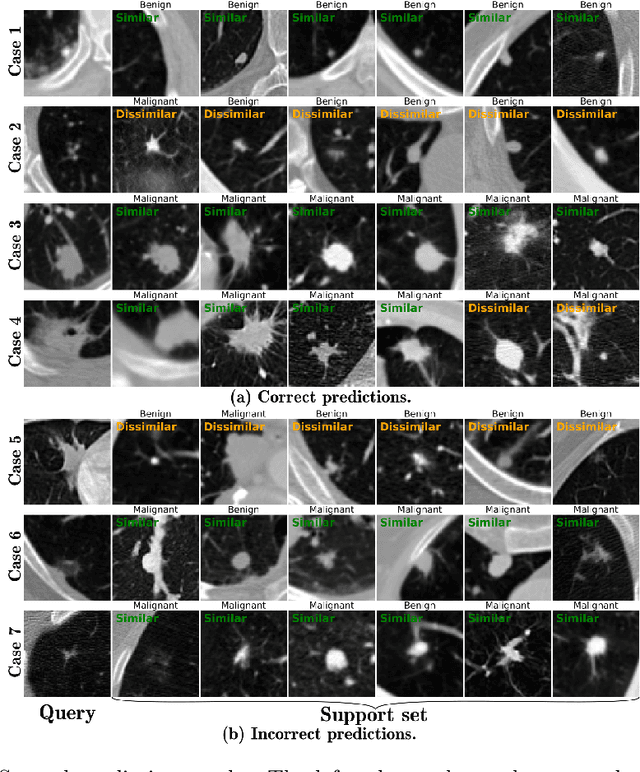
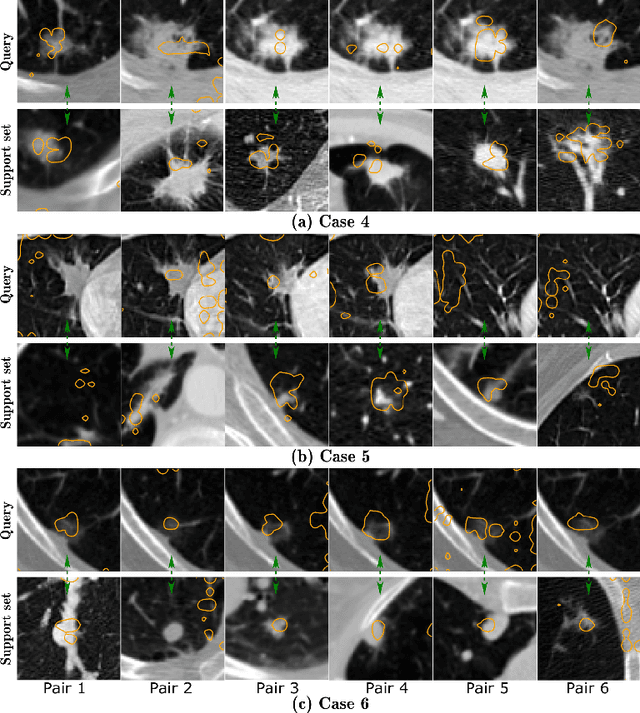
With the ongoing development of deep learning, an increasing number of AI models have surpassed the performance levels of human clinical practitioners. However, the prevalence of AI diagnostic products in actual clinical practice remains significantly lower than desired. One crucial reason for this gap is the so-called `black box' nature of AI models. Clinicians' distrust of black box models has directly hindered the clinical deployment of AI products. To address this challenge, we propose ContrastDiagnosis, a straightforward yet effective interpretable diagnosis framework. This framework is designed to introduce inherent transparency and provide extensive post-hoc explainability for deep learning model, making them more suitable for clinical medical diagnosis. ContrastDiagnosis incorporates a contrastive learning mechanism to provide a case-based reasoning diagnostic rationale, enhancing the model's transparency and also offers post-hoc interpretability by highlighting similar areas. High diagnostic accuracy was achieved with AUC of 0.977 while maintain a high transparency and explainability.
DynaPipe: Optimizing Multi-task Training through Dynamic Pipelines
Nov 17, 2023Chenyu Jiang, Zhen Jia, Shuai Zheng, Yida Wang, Chuan Wu
Multi-task model training has been adopted to enable a single deep neural network model (often a large language model) to handle multiple tasks (e.g., question answering and text summarization). Multi-task training commonly receives input sequences of highly different lengths due to the diverse contexts of different tasks. Padding (to the same sequence length) or packing (short examples into long sequences of the same length) is usually adopted to prepare input samples for model training, which is nonetheless not space or computation efficient. This paper proposes a dynamic micro-batching approach to tackle sequence length variation and enable efficient multi-task model training. We advocate pipeline-parallel training of the large model with variable-length micro-batches, each of which potentially comprises a different number of samples. We optimize micro-batch construction using a dynamic programming-based approach, and handle micro-batch execution time variation through dynamic pipeline and communication scheduling, enabling highly efficient pipeline training. Extensive evaluation on the FLANv2 dataset demonstrates up to 4.39x higher training throughput when training T5, and 3.25x when training GPT, as compared with packing-based baselines. DynaPipe's source code is publicly available at https://github.com/awslabs/optimizing-multitask-training-through-dynamic-pipelines.
Serving Deep Learning Model in Relational Databases
Oct 10, 2023Alexandre Eichenberger, Qi Lin, Saif Masood, Hong Min, Alexander Sim, Jie Wang, Yida Wang, Kesheng Wu, Binhang Yuan, Lixi Zhou, Jia Zou
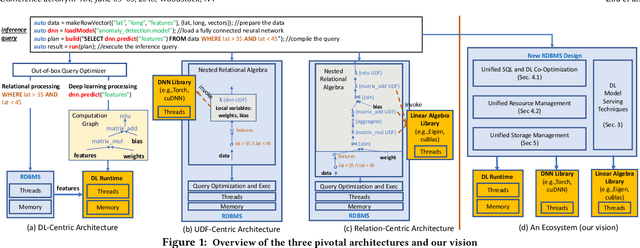
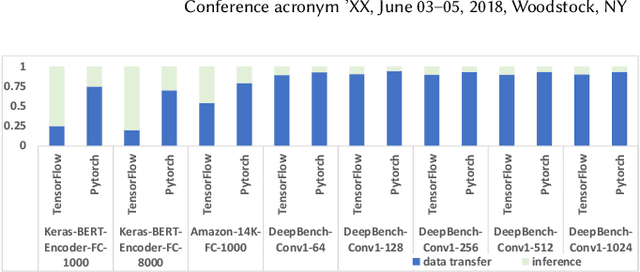

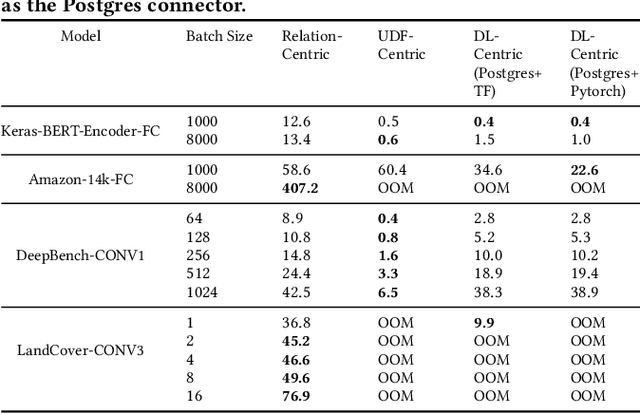
Serving deep learning (DL) models on relational data has become a critical requirement across diverse commercial and scientific domains, sparking growing interest recently. In this visionary paper, we embark on a comprehensive exploration of representative architectures to address the requirement. We highlight three pivotal paradigms: The state-of-the-artDL-Centricarchitecture offloadsDL computations to dedicated DL frameworks. The potential UDF-Centric architecture encapsulates one or more tensor computations into User Defined Functions (UDFs) within the database system. The potentialRelation-Centricarchitecture aims to represent a large-scale tensor computation through relational operators. While each of these architectures demonstrates promise in specific use scenarios, we identify urgent requirements for seamless integration of these architectures and the middle ground between these architectures. We delve into the gaps that impede the integration and explore innovative strategies to close them. We present a pathway to establish a novel database system for enabling a broad class of data-intensive DL inference applications.
Target-independent XLA optimization using Reinforcement Learning
Aug 28, 2023Milan Ganai, Haichen Li, Theodore Enns, Yida Wang, Randy Huang



An important challenge in Machine Learning compilers like XLA is multi-pass optimization and analysis. There has been recent interest chiefly in XLA target-dependent optimization on the graph-level, subgraph-level, and kernel-level phases. We specifically focus on target-independent optimization XLA HLO pass ordering: our approach aims at finding the optimal sequence of compiler optimization passes, which is decoupled from target-dependent optimization. However, there is little domain specific study in pass ordering for XLA HLO. To this end, we propose introducing deep Reinforcement Learning (RL) based search for optimal XLA HLO pass ordering. We also propose enhancements to the deep RL algorithms to further improve optimal search performance and open the research direction for domain-specific guidance for RL. We create an XLA Gym experimentation framework as a tool to enable RL algorithms to interact with the compiler for passing optimizations and thereby train agents. Overall, in our experimentation we observe an average of $13.3\%$ improvement in operation count reduction on a benchmark of GPT-2 training graphs and $10.4\%$ improvement on a diverse benchmark including GPT-2, BERT, and ResNet graphs using the proposed approach over the compiler's default phase ordering.
$\mathrm{SAM^{Med}}$: A medical image annotation framework based on large vision model
Jul 11, 2023Chenglong Wang, Dexuan Li, Sucheng Wang, Chengxiu Zhang, Yida Wang, Yun Liu, Guang Yang

Recently, large vision model, Segment Anything Model (SAM), has revolutionized the computer vision field, especially for image segmentation. SAM presented a new promptable segmentation paradigm that exhibit its remarkable zero-shot generalization ability. An extensive researches have explore the potential and limits of SAM in various downstream tasks. In this study, we presents $\mathrm{SAM^{Med}}$, an enhanced framework for medical image annotation that leverages the capabilities of SAM. $\mathrm{SAM^{Med}}$ framework consisted of two submodules, namely $\mathrm{SAM^{assist}}$ and $\mathrm{SAM^{auto}}$. The $\mathrm{SAM^{assist}}$ demonstrates the generalization ability of SAM to the downstream medical segmentation task using the prompt-learning approach. Results show a significant improvement in segmentation accuracy with only approximately 5 input points. The $\mathrm{SAM^{auto}}$ model aims to accelerate the annotation process by automatically generating input prompts. The proposed SAP-Net model achieves superior segmentation performance with only five annotated slices, achieving an average Dice coefficient of 0.80 and 0.82 for kidney and liver segmentation, respectively. Overall, $\mathrm{SAM^{Med}}$ demonstrates promising results in medical image annotation. These findings highlight the potential of leveraging large-scale vision models in medical image annotation tasks.
RAF: Holistic Compilation for Deep Learning Model Training
Mar 08, 2023Cody Hao Yu, Haozheng Fan, Guangtai Huang, Zhen Jia, Yizhi Liu, Jie Wang, Zach Zheng, Yuan Zhou, Haichen Shen, Junru Shao, Mu Li, Yida Wang
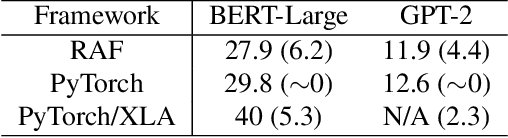

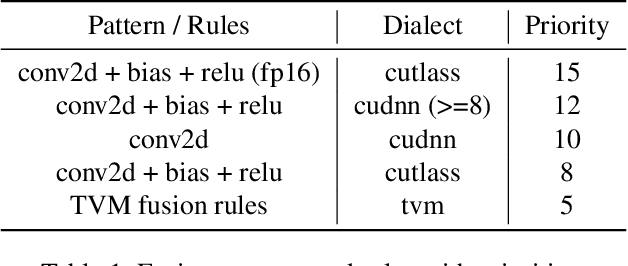


As deep learning is pervasive in modern applications, many deep learning frameworks are presented for deep learning practitioners to develop and train DNN models rapidly. Meanwhile, as training large deep learning models becomes a trend in recent years, the training throughput and memory footprint are getting crucial. Accordingly, optimizing training workloads with compiler optimizations is inevitable and getting more and more attentions. However, existing deep learning compilers (DLCs) mainly target inference and do not incorporate holistic optimizations, such as automatic differentiation and automatic mixed precision, in training workloads. In this paper, we present RAF, a deep learning compiler for training. Unlike existing DLCs, RAF accepts a forward model and in-house generates a training graph. Accordingly, RAF is able to systematically consolidate graph optimizations for performance, memory and distributed training. In addition, to catch up to the state-of-the-art performance with hand-crafted kernel libraries as well as tensor compilers, RAF proposes an operator dialect mechanism to seamlessly integrate all possible kernel implementations. We demonstrate that by in-house training graph generation and operator dialect mechanism, we are able to perform holistic optimizations and achieve either better training throughput or larger batch size against PyTorch (eager and torchscript mode), XLA, and DeepSpeed for popular transformer models on GPUs.
Decoupled Model Schedule for Deep Learning Training
Feb 16, 2023Hongzheng Chen, Cody Hao Yu, Shuai Zheng, Zhen Zhang, Zhiru Zhang, Yida Wang
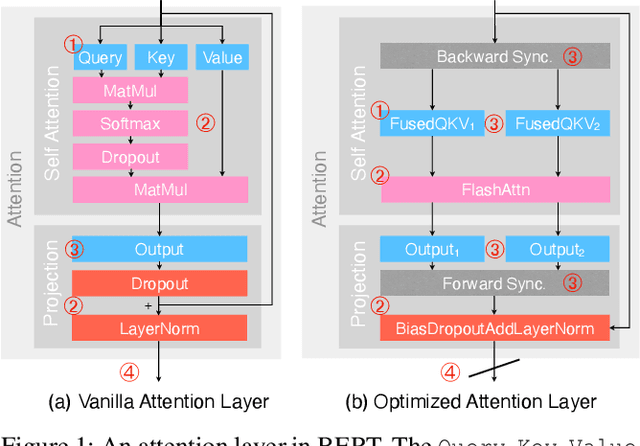

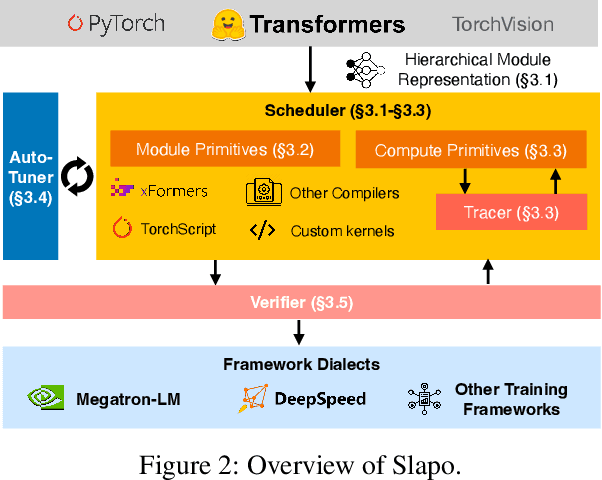
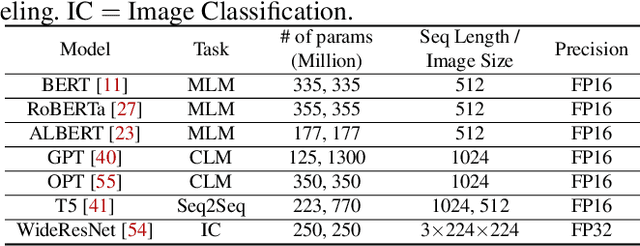
Recent years have seen an increase in the development of large deep learning (DL) models, which makes training efficiency crucial. Common practice is struggling with the trade-off between usability and performance. On one hand, DL frameworks such as PyTorch use dynamic graphs to facilitate model developers at a price of sub-optimal model training performance. On the other hand, practitioners propose various approaches to improving the training efficiency by sacrificing some of the flexibility, ranging from making the graph static for more thorough optimization (e.g., XLA) to customizing optimization towards large-scale distributed training (e.g., DeepSpeed and Megatron-LM). In this paper, we aim to address the tension between usability and training efficiency through separation of concerns. Inspired by DL compilers that decouple the platform-specific optimizations of a tensor-level operator from its arithmetic definition, this paper proposes a schedule language to decouple model execution from definition. Specifically, the schedule works on a PyTorch model and uses a set of schedule primitives to convert the model for common model training optimizations such as high-performance kernels, effective 3D parallelism, and efficient activation checkpointing. Compared to existing optimization solutions, we optimize the model as-needed through high-level primitives, and thus preserving programmability and debuggability for users to a large extent. Our evaluation results show that by scheduling the existing hand-crafted optimizations in a systematic way, we are able to improve training throughput by up to 3.35x on a single machine with 8 NVIDIA V100 GPUs, and by up to 1.32x on multiple machines with up to 64 GPUs, when compared to the out-of-the-box performance of DeepSpeed and Megatron-LM.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge