Circle Representation for Medical Instance Object Segmentation
Mar 18, 2024Juming Xiong, Ethan H. Nguyen, Yilin Liu, Ruining Deng, Regina N Tyree, Hernan Correa, Girish Hiremath, Yaohong Wang, Haichun Yang, Agnes B. Fogo, Yuankai Huo
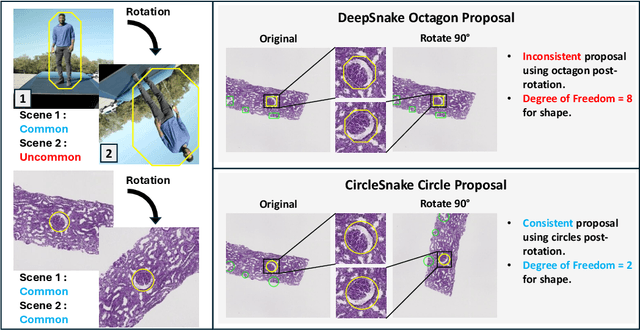
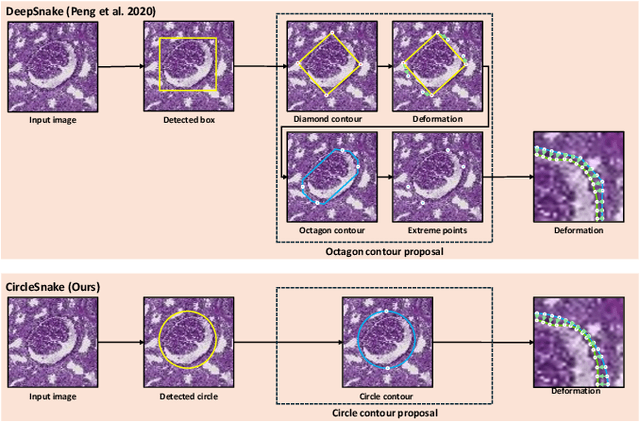
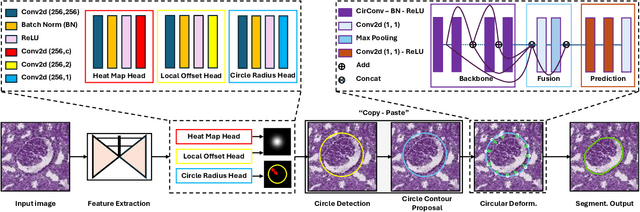
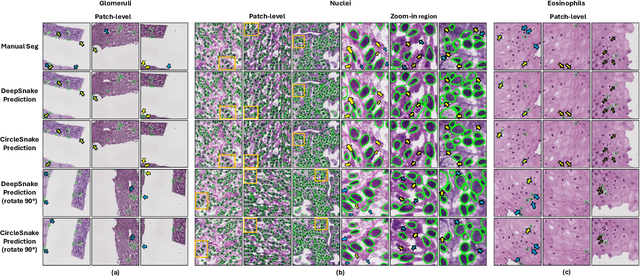
Recently, circle representation has been introduced for medical imaging, designed specifically to enhance the detection of instance objects that are spherically shaped (e.g., cells, glomeruli, and nuclei). Given its outstanding effectiveness in instance detection, it is compelling to consider the application of circle representation for segmenting instance medical objects. In this study, we introduce CircleSnake, a simple end-to-end segmentation approach that utilizes circle contour deformation for segmenting ball-shaped medical objects at the instance level. The innovation of CircleSnake lies in these three areas: (1) It substitutes the complex bounding box-to-octagon contour transformation with a more consistent and rotation-invariant bounding circle-to-circle contour adaptation. This adaptation specifically targets ball-shaped medical objects. (2) The circle representation employed in CircleSnake significantly reduces the degrees of freedom to two, compared to eight in the octagon representation. This reduction enhances both the robustness of the segmentation performance and the rotational consistency of the method. (3) CircleSnake is the first end-to-end deep instance segmentation pipeline to incorporate circle representation, encompassing consistent circle detection, circle contour proposal, and circular convolution in a unified framework. This integration is achieved through the novel application of circular graph convolution within the context of circle detection and instance segmentation. In practical applications, such as the detection of glomeruli, nuclei, and eosinophils in pathological images, CircleSnake has demonstrated superior performance and greater rotation invariance when compared to benchmarks. The code has been made publicly available: https://github.com/hrlblab/CircleSnake.
Towards Architecture-Insensitive Untrained Network Priors for Accelerated MRI Reconstruction
Dec 15, 2023Yilin Liu, Yunkui Pang, Jiang Li, Yong Chen, Pew-Thian Yap
Untrained neural networks pioneered by Deep Image Prior (DIP) have recently enabled MRI reconstruction without requiring fully-sampled measurements for training. Their success is widely attributed to the implicit regularization induced by suitable network architectures. However, the lack of understanding of such architectural priors results in superfluous design choices and sub-optimal outcomes. This work aims to simplify the architectural design decisions for DIP-MRI to facilitate its practical deployment. We observe that certain architectural components are more prone to causing overfitting regardless of the number of parameters, incurring severe reconstruction artifacts by hindering accurate extrapolation on the un-acquired measurements. We interpret this phenomenon from a frequency perspective and find that the architectural characteristics favoring low frequencies, i.e., deep and narrow with unlearnt upsampling, can lead to enhanced generalization and hence better reconstruction. Building on this insight, we propose two architecture-agnostic remedies: one to constrain the frequency range of the white-noise input and the other to penalize the Lipschitz constants of the network. We demonstrate that even with just one extra line of code on the input, the performance gap between the ill-designed models and the high-performing ones can be closed. These results signify that for the first time, architectural biases on untrained MRI reconstruction can be mitigated without architectural modifications.
Eosinophils Instance Object Segmentation on Whole Slide Imaging Using Multi-label Circle Representation
Aug 17, 2023Yilin Liu, Ruining Deng, Juming Xiong, Regina N Tyree, Hernan Correa, Girish Hiremath, Yaohong Wang, Yuankai Huo




Eosinophilic esophagitis (EoE) is a chronic and relapsing disease characterized by esophageal inflammation. Symptoms of EoE include difficulty swallowing, food impaction, and chest pain which significantly impact the quality of life, resulting in nutritional impairments, social limitations, and psychological distress. The diagnosis of EoE is typically performed with a threshold (15 to 20) of eosinophils (Eos) per high-power field (HPF). Since the current counting process of Eos is a resource-intensive process for human pathologists, automatic methods are desired. Circle representation has been shown as a more precise, yet less complicated, representation for automatic instance cell segmentation such as CircleSnake approach. However, the CircleSnake was designed as a single-label model, which is not able to deal with multi-label scenarios. In this paper, we propose the multi-label CircleSnake model for instance segmentation on Eos. It extends the original CircleSnake model from a single-label design to a multi-label model, allowing segmentation of multiple object types. Experimental results illustrate the CircleSnake model's superiority over the traditional Mask R-CNN model and DeepSnake model in terms of average precision (AP) in identifying and segmenting eosinophils, thereby enabling enhanced characterization of EoE. This automated approach holds promise for streamlining the assessment process and improving diagnostic accuracy in EoE analysis. The source code has been made publicly available at https://github.com/yilinliu610730/EoE.
Deep Learning-Based Open Source Toolkit for Eosinophil Detection in Pediatric Eosinophilic Esophagitis
Aug 11, 2023Juming Xiong, Yilin Liu, Ruining Deng, Regina N Tyree, Hernan Correa, Girish Hiremath, Yaohong Wang, Yuankai Huo
Eosinophilic Esophagitis (EoE) is a chronic, immune/antigen-mediated esophageal disease, characterized by symptoms related to esophageal dysfunction and histological evidence of eosinophil-dominant inflammation. Owing to the intricate microscopic representation of EoE in imaging, current methodologies which depend on manual identification are not only labor-intensive but also prone to inaccuracies. In this study, we develop an open-source toolkit, named Open-EoE, to perform end-to-end whole slide image (WSI) level eosinophil (Eos) detection using one line of command via Docker. Specifically, the toolkit supports three state-of-the-art deep learning-based object detection models. Furthermore, Open-EoE further optimizes the performance by implementing an ensemble learning strategy, and enhancing the precision and reliability of our results. The experimental results demonstrated that the Open-EoE toolkit can efficiently detect Eos on a testing set with 289 WSIs. At the widely accepted threshold of >= 15 Eos per high power field (HPF) for diagnosing EoE, the Open-EoE achieved an accuracy of 91%, showing decent consistency with pathologist evaluations. This suggests a promising avenue for integrating machine learning methodologies into the diagnostic process for EoE. The docker and source code has been made publicly available at https://github.com/hrlblab/Open-EoE.
Spatial Pathomics Toolkit for Quantitative Analysis of Podocyte Nuclei with Histology and Spatial Transcriptomics Data in Renal Pathology
Aug 10, 2023Jiayuan Chen, Yu Wang, Ruining Deng, Quan Liu, Can Cui, Tianyuan Yao, Yilin Liu, Jianyong Zhong, Agnes B. Fogo, Haichun Yang, Shilin Zhao, Yuankai Huo
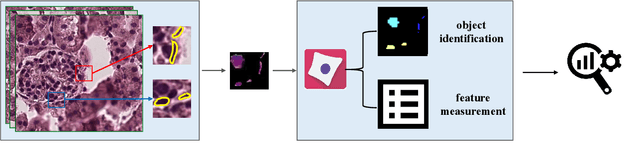

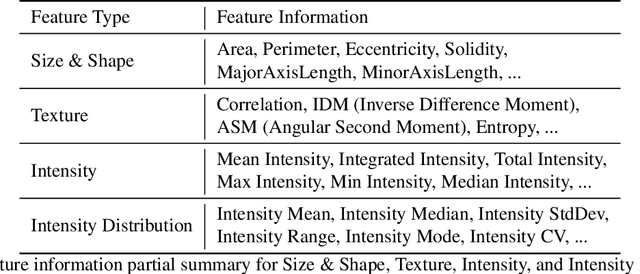
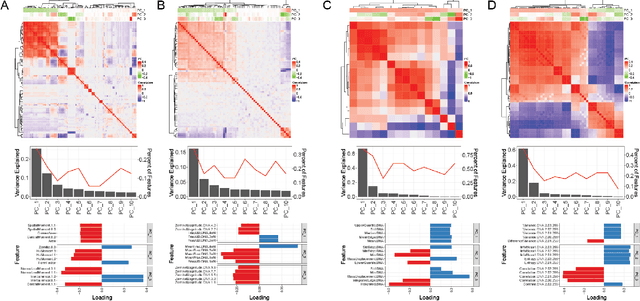
Podocytes, specialized epithelial cells that envelop the glomerular capillaries, play a pivotal role in maintaining renal health. The current description and quantification of features on pathology slides are limited, prompting the need for innovative solutions to comprehensively assess diverse phenotypic attributes within Whole Slide Images (WSIs). In particular, understanding the morphological characteristics of podocytes, terminally differentiated glomerular epithelial cells, is crucial for studying glomerular injury. This paper introduces the Spatial Pathomics Toolkit (SPT) and applies it to podocyte pathomics. The SPT consists of three main components: (1) instance object segmentation, enabling precise identification of podocyte nuclei; (2) pathomics feature generation, extracting a comprehensive array of quantitative features from the identified nuclei; and (3) robust statistical analyses, facilitating a comprehensive exploration of spatial relationships between morphological and spatial transcriptomics features.The SPT successfully extracted and analyzed morphological and textural features from podocyte nuclei, revealing a multitude of podocyte morphomic features through statistical analysis. Additionally, we demonstrated the SPT's ability to unravel spatial information inherent to podocyte distribution, shedding light on spatial patterns associated with glomerular injury. By disseminating the SPT, our goal is to provide the research community with a powerful and user-friendly resource that advances cellular spatial pathomics in renal pathology. The implementation and its complete source code of the toolkit are made openly accessible at https://github.com/hrlblab/spatial_pathomics.
The Devil is in the Upsampling: Architectural Decisions Made Simpler for Denoising with Deep Image Prior
Apr 22, 2023Yilin Liu, Jiang Li, Yunkui Pang, Dong Nie, Pew-thian Yap




Deep Image Prior (DIP) shows that some network architectures naturally bias towards smooth images and resist noises, a phenomenon known as spectral bias. Image denoising is an immediate application of this property. Although DIP has removed the requirement of large training sets, it still presents two practical challenges for denoising: architectural design and noise-fitting, which are often intertwined. Existing methods mostly handcraft or search for the architecture from a large design space, due to the lack of understanding on how the architectural choice corresponds to the image. In this study, we analyze from a frequency perspective to demonstrate that the unlearnt upsampling is the main driving force behind the denoising phenomenon in DIP. This finding then leads to strategies for estimating a suitable architecture for every image without a laborious search. Extensive experiments show that the estimated architectures denoise and preserve the textural details better than current methods with up to 95% fewer parameters. The under-parameterized nature also makes them especially robust to a higher level of noise.
Learning Reconstructability for Drone Aerial Path Planning
Sep 21, 2022Yilin Liu, Liqiang Lin, Yue Hu, Ke Xie, Chi-Wing Fu, Hao Zhang, Hui Huang




We introduce the first learning-based reconstructability predictor to improve view and path planning for large-scale 3D urban scene acquisition using unmanned drones. In contrast to previous heuristic approaches, our method learns a model that explicitly predicts how well a 3D urban scene will be reconstructed from a set of viewpoints. To make such a model trainable and simultaneously applicable to drone path planning, we simulate the proxy-based 3D scene reconstruction during training to set up the prediction. Specifically, the neural network we design is trained to predict the scene reconstructability as a function of the proxy geometry, a set of viewpoints, and optionally a series of scene images acquired in flight. To reconstruct a new urban scene, we first build the 3D scene proxy, then rely on the predicted reconstruction quality and uncertainty measures by our network, based off of the proxy geometry, to guide the drone path planning. We demonstrate that our data-driven reconstructability predictions are more closely correlated to the true reconstruction quality than prior heuristic measures. Further, our learned predictor can be easily integrated into existing path planners to yield improvements. Finally, we devise a new iterative view planning framework, based on the learned reconstructability, and show superior performance of the new planner when reconstructing both synthetic and real scenes.
Real-Time Mapping of Tissue Properties for Magnetic Resonance Fingerprinting
Jul 16, 2021Yilin Liu, Yong Chen, Pew-Thian Yap




Magnetic resonance Fingerprinting (MRF) is a relatively new multi-parametric quantitative imaging method that involves a two-step process: (i) reconstructing a series of time frames from highly-undersampled non-Cartesian spiral k-space data and (ii) pattern matching using the time frames to infer tissue properties (e.g., T1 and T2 relaxation times). In this paper, we introduce a novel end-to-end deep learning framework to seamlessly map the tissue properties directly from spiral k-space MRF data, thereby avoiding time-consuming processing such as the nonuniform fast Fourier transform (NUFFT) and the dictionary-based Fingerprint matching. Our method directly consumes the non-Cartesian k- space data, performs adaptive density compensation, and predicts multiple tissue property maps in one forward pass. Experiments on both 2D and 3D MRF data demonstrate that quantification accuracy comparable to state-of-the-art methods can be accomplished within 0.5 second, which is 1100 to 7700 times faster than the original MRF framework. The proposed method is thus promising for facilitating the adoption of MRF in clinical settings.
UrbanScene3D: A Large Scale Urban Scene Dataset and Simulator
Jul 09, 2021Yilin Liu, Fuyou Xue, Hui Huang



The ability to perceive the environments in different ways is essential to robotic research. This involves the analysis of both 2D and 3D data sources. We present a large scale urban scene dataset associated with a handy simulator based on Unreal Engine 4 and AirSim, which consists of both man-made and real-world reconstruction scenes in different scales, referred to as UrbanScene3D. Unlike previous works that purely based on 2D information or man-made 3D CAD models, UrbanScene3D contains both compact man-made models and detailed real-world models reconstructed by aerial images. Each building has been manually extracted from the entire scene model and then has been assigned with a unique label, forming an instance segmentation map. The provided 3D ground-truth textured models with instance segmentation labels in UrbanScene3D allow users to obtain all kinds of data they would like to have: instance segmentation map, depth map in arbitrary resolution, 3D point cloud/mesh in both visible and invisible places, etc. In addition, with the help of AirSim, users can also simulate the robots (cars/drones)to test a variety of autonomous tasks in the proposed city environment. Please refer to our paper and website(https://vcc.tech/UrbanScene3D/) for further details and applications.
VGF-Net: Visual-Geometric Fusion Learning for Simultaneous Drone Navigation and Height Mapping
Apr 07, 2021Yilin Liu, Ke Xie, Hui Huang




The drone navigation requires the comprehensive understanding of both visual and geometric information in the 3D world. In this paper, we present a Visual-Geometric Fusion Network(VGF-Net), a deep network for the fusion analysis of visual/geometric data and the construction of 2.5D height maps for simultaneous drone navigation in novel environments. Given an initial rough height map and a sequence of RGB images, our VGF-Net extracts the visual information of the scene, along with a sparse set of 3D keypoints that capture the geometric relationship between objects in the scene. Driven by the data, VGF-Net adaptively fuses visual and geometric information, forming a unified Visual-Geometric Representation. This representation is fed to a new Directional Attention Model(DAM), which helps enhance the visual-geometric object relationship and propagates the informative data to dynamically refine the height map and the corresponding keypoints. An entire end-to-end information fusion and mapping system is formed, demonstrating remarkable robustness and high accuracy on the autonomous drone navigation across complex indoor and large-scale outdoor scenes. The dataset can be found in http://vcc.szu.edu.cn/research/2021/VGFNet.
* Accepted by CVM 2021
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge