EEG-MACS: Manifold Attention and Confidence Stratification for EEG-based Cross-Center Brain Disease Diagnosis under Unreliable Annotations
Apr 29, 2024Zhenxi Song, Ruihan Qin, Huixia Ren, Zhen Liang, Yi Guo, Min Zhang, Zhiguo Zhang
Cross-center data heterogeneity and annotation unreliability significantly challenge the intelligent diagnosis of diseases using brain signals. A notable example is the EEG-based diagnosis of neurodegenerative diseases, which features subtler abnormal neural dynamics typically observed in small-group settings. To advance this area, in this work, we introduce a transferable framework employing Manifold Attention and Confidence Stratification (MACS) to diagnose neurodegenerative disorders based on EEG signals sourced from four centers with unreliable annotations. The MACS framework's effectiveness stems from these features: 1) The Augmentor generates various EEG-represented brain variants to enrich the data space; 2) The Switcher enhances the feature space for trusted samples and reduces overfitting on incorrectly labeled samples; 3) The Encoder uses the Riemannian manifold and Euclidean metrics to capture spatiotemporal variations and dynamic synchronization in EEG; 4) The Projector, equipped with dual heads, monitors consistency across multiple brain variants and ensures diagnostic accuracy; 5) The Stratifier adaptively stratifies learned samples by confidence levels throughout the training process; 6) Forward and backpropagation in MACS are constrained by confidence stratification to stabilize the learning system amid unreliable annotations. Our subject-independent experiments, conducted on both neurocognitive and movement disorders using cross-center corpora, have demonstrated superior performance compared to existing related algorithms. This work not only improves EEG-based diagnostics for cross-center and small-setting brain diseases but also offers insights into extending MACS techniques to other data analyses, tackling data heterogeneity and annotation unreliability in multimedia and multimodal content understanding.
MDDD: Manifold-based Domain Adaptation with Dynamic Distribution for Non-Deep Transfer Learning in Cross-subject and Cross-session EEG-based Emotion Recognition
Apr 24, 2024Ting Luo, Jing Zhang, Yingwei Qiu, Li Zhang, Yaohua Hu, Zhuliang Yu, Zhen Liang
Emotion decoding using Electroencephalography (EEG)-based affective brain-computer interfaces represents a significant area within the field of affective computing. In the present study, we propose a novel non-deep transfer learning method, termed as Manifold-based Domain adaptation with Dynamic Distribution (MDDD). The proposed MDDD includes four main modules: manifold feature transformation, dynamic distribution alignment, classifier learning, and ensemble learning. The data undergoes a transformation onto an optimal Grassmann manifold space, enabling dynamic alignment of the source and target domains. This process prioritizes both marginal and conditional distributions according to their significance, ensuring enhanced adaptation efficiency across various types of data. In the classifier learning, the principle of structural risk minimization is integrated to develop robust classification models. This is complemented by dynamic distribution alignment, which refines the classifier iteratively. Additionally, the ensemble learning module aggregates the classifiers obtained at different stages of the optimization process, which leverages the diversity of the classifiers to enhance the overall prediction accuracy. The experimental results indicate that MDDD outperforms traditional non-deep learning methods, achieving an average improvement of 3.54%, and is comparable to deep learning methods. This suggests that MDDD could be a promising method for enhancing the utility and applicability of aBCIs in real-world scenarios.
Joint Contrastive Learning with Feature Alignment for Cross-Corpus EEG-based Emotion Recognition
Apr 15, 2024Qile Liu, Zhihao Zhou, Jiyuan Wang, Zhen Liang
The integration of human emotions into multimedia applications shows great potential for enriching user experiences and enhancing engagement across various digital platforms. Unlike traditional methods such as questionnaires, facial expressions, and voice analysis, brain signals offer a more direct and objective understanding of emotional states. However, in the field of electroencephalography (EEG)-based emotion recognition, previous studies have primarily concentrated on training and testing EEG models within a single dataset, overlooking the variability across different datasets. This oversight leads to significant performance degradation when applying EEG models to cross-corpus scenarios. In this study, we propose a novel Joint Contrastive learning framework with Feature Alignment (JCFA) to address cross-corpus EEG-based emotion recognition. The JCFA model operates in two main stages. In the pre-training stage, a joint domain contrastive learning strategy is introduced to characterize generalizable time-frequency representations of EEG signals, without the use of labeled data. It extracts robust time-based and frequency-based embeddings for each EEG sample, and then aligns them within a shared latent time-frequency space. In the fine-tuning stage, JCFA is refined in conjunction with downstream tasks, where the structural connections among brain electrodes are considered. The model capability could be further enhanced for the application in emotion detection and interpretation. Extensive experimental results on two well-recognized emotional datasets show that the proposed JCFA model achieves state-of-the-art (SOTA) performance, outperforming the second-best method by an average accuracy increase of 4.09% in cross-corpus EEG-based emotion recognition tasks.
UR4NNV: Neural Network Verification, Under-approximation Reachability Works!
Jan 23, 2024Zhen Liang, Taoran Wu, Ran Zhao, Bai Xue, Ji Wang, Wenjing Yang, Shaojun Deng, Wanwei Liu
Recently, formal verification of deep neural networks (DNNs) has garnered considerable attention, and over-approximation based methods have become popular due to their effectiveness and efficiency. However, these strategies face challenges in addressing the "unknown dilemma" concerning whether the exact output region or the introduced approximation error violates the property in question. To address this, this paper introduces the UR4NNV verification framework, which utilizes under-approximation reachability analysis for DNN verification for the first time. UR4NNV focuses on DNNs with Rectified Linear Unit (ReLU) activations and employs a binary tree branch-based under-approximation algorithm. In each epoch, UR4NNV under-approximates a sub-polytope of the reachable set and verifies this polytope against the given property. Through a trial-and-error approach, UR4NNV effectively falsifies DNN properties while providing confidence levels when reaching verification epoch bounds and failing falsifying properties. Experimental comparisons with existing verification methods demonstrate the effectiveness and efficiency of UR4NNV, significantly reducing the impact of the "unknown dilemma".
Semi-Supervised Dual-Stream Self-Attentive Adversarial Graph Contrastive Learning for Cross-Subject EEG-based Emotion Recognition
Aug 13, 2023Weishan Ye, Zhiguo Zhang, Min Zhang, Fei Teng, Li Zhang, Linling Li, Gan Huang, Jianhong Wang, Dong Ni, Zhen Liang
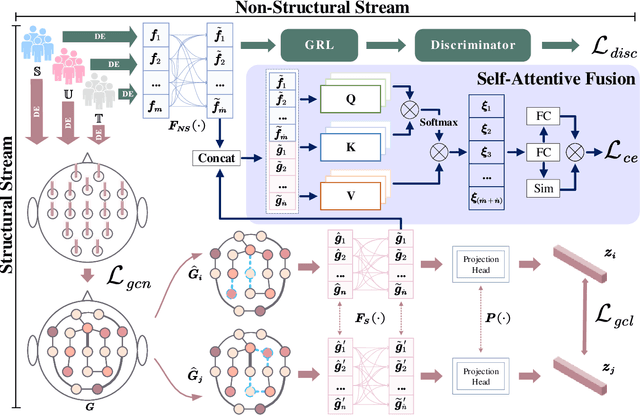
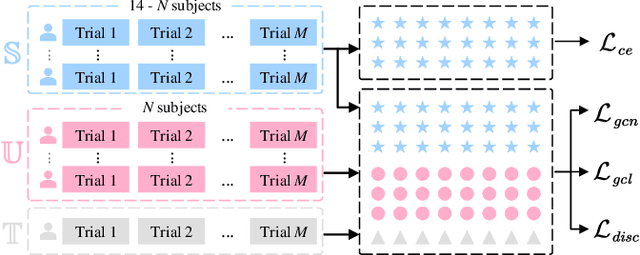
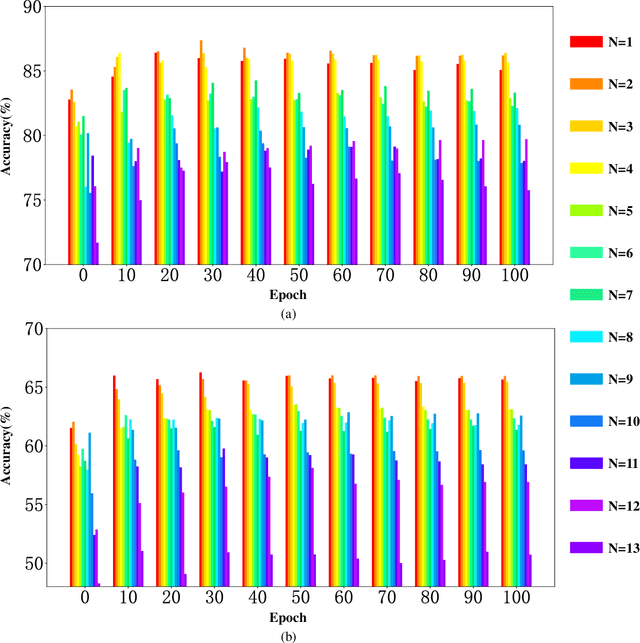
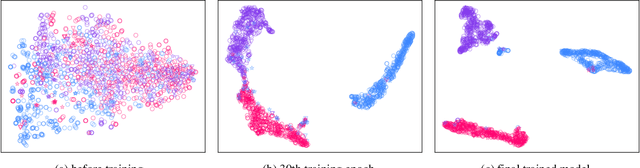
Electroencephalography (EEG) is an objective tool for emotion recognition with promising applications. However, the scarcity of labeled data remains a major challenge in this field, limiting the widespread use of EEG-based emotion recognition. In this paper, a semi-supervised Dual-stream Self-Attentive Adversarial Graph Contrastive learning framework (termed as DS-AGC) is proposed to tackle the challenge of limited labeled data in cross-subject EEG-based emotion recognition. The DS-AGC framework includes two parallel streams for extracting non-structural and structural EEG features. The non-structural stream incorporates a semi-supervised multi-domain adaptation method to alleviate distribution discrepancy among labeled source domain, unlabeled source domain, and unknown target domain. The structural stream develops a graph contrastive learning method to extract effective graph-based feature representation from multiple EEG channels in a semi-supervised manner. Further, a self-attentive fusion module is developed for feature fusion, sample selection, and emotion recognition, which highlights EEG features more relevant to emotions and data samples in the labeled source domain that are closer to the target domain. Extensive experiments conducted on two benchmark databases (SEED and SEED-IV) using a semi-supervised cross-subject leave-one-subject-out cross-validation evaluation scheme show that the proposed model outperforms existing methods under different incomplete label conditions (with an average improvement of 5.83% on SEED and 6.99% on SEED-IV), demonstrating its effectiveness in addressing the label scarcity problem in cross-subject EEG-based emotion recognition.
An Automata-Theoretic Approach to Synthesizing Binarized Neural Networks
Jul 29, 2023Ye Tao, Wanwei Liu, Fu Song, Zhen Liang, Ji Wang, Hongxu Zhu
Deep neural networks, (DNNs, a.k.a. NNs), have been widely used in various tasks and have been proven to be successful. However, the accompanied expensive computing and storage costs make the deployments in resource-constrained devices a significant concern. To solve this issue, quantization has emerged as an effective way to reduce the costs of DNNs with little accuracy degradation by quantizing floating-point numbers to low-width fixed-point representations. Quantized neural networks (QNNs) have been developed, with binarized neural networks (BNNs) restricted to binary values as a special case. Another concern about neural networks is their vulnerability and lack of interpretability. Despite the active research on trustworthy of DNNs, few approaches have been proposed to QNNs. To this end, this paper presents an automata-theoretic approach to synthesizing BNNs that meet designated properties. More specifically, we define a temporal logic, called BLTL, as the specification language. We show that each BLTL formula can be transformed into an automaton on finite words. To deal with the state-explosion problem, we provide a tableau-based approach in real implementation. For the synthesis procedure, we utilize SMT solvers to detect the existence of a model (i.e., a BNN) in the construction process. Notably, synthesis provides a way to determine the hyper-parameters of the network before training.Moreover, we experimentally evaluate our approach and demonstrate its effectiveness in improving the individual fairness and local robustness of BNNs while maintaining accuracy to a great extent.
Verifying Safety of Neural Networks from Topological Perspectives
Jun 27, 2023Zhen Liang, Dejin Ren, Bai Xue, Ji Wang, Wenjing Yang, Wanwei Liu
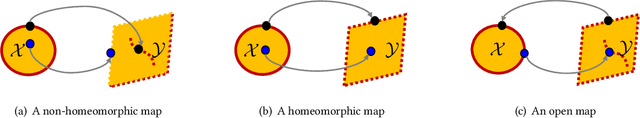
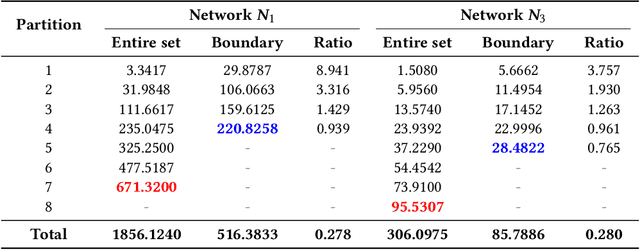
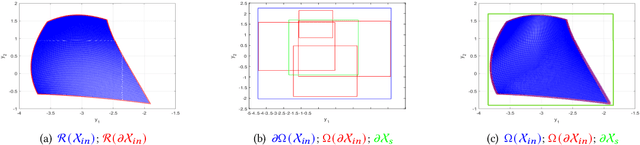
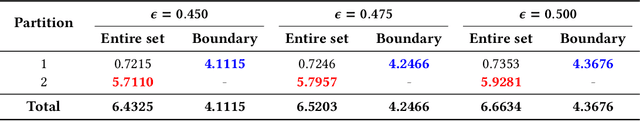
Neural networks (NNs) are increasingly applied in safety-critical systems such as autonomous vehicles. However, they are fragile and are often ill-behaved. Consequently, their behaviors should undergo rigorous guarantees before deployment in practice. In this paper, we propose a set-boundary reachability method to investigate the safety verification problem of NNs from a topological perspective. Given an NN with an input set and a safe set, the safety verification problem is to determine whether all outputs of the NN resulting from the input set fall within the safe set. In our method, the homeomorphism property and the open map property of NNs are mainly exploited, which establish rigorous guarantees between the boundaries of the input set and the boundaries of the output set. The exploitation of these two properties facilitates reachability computations via extracting subsets of the input set rather than the entire input set, thus controlling the wrapping effect in reachability analysis and facilitating the reduction of computation burdens for safety verification. The homeomorphism property exists in some widely used NNs such as invertible residual networks (i-ResNets) and Neural ordinary differential equations (Neural ODEs), and the open map is a less strict property and easier to satisfy compared with the homeomorphism property. For NNs establishing either of these properties, our set-boundary reachability method only needs to perform reachability analysis on the boundary of the input set. Moreover, for NNs that do not feature these properties with respect to the input set, we explore subsets of the input set for establishing the local homeomorphism property and then abandon these subsets for reachability computations. Finally, some examples demonstrate the performance of the proposed method.
Semi-Supervised Learning for Multi-Label Cardiovascular Diseases Prediction:A Multi-Dataset Study
Jun 18, 2023Rushuang Zhou, Lei Lu, Zijun Liu, Ting Xiang, Zhen Liang, David A. Clifton, Yining Dong, Yuan-Ting Zhang
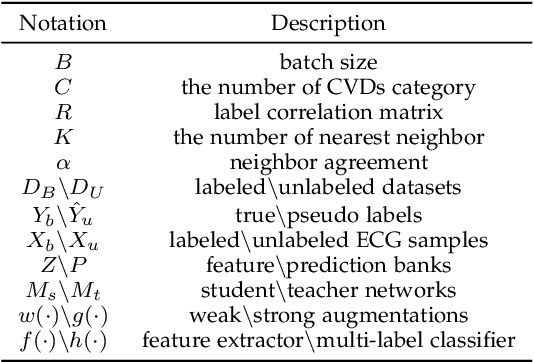

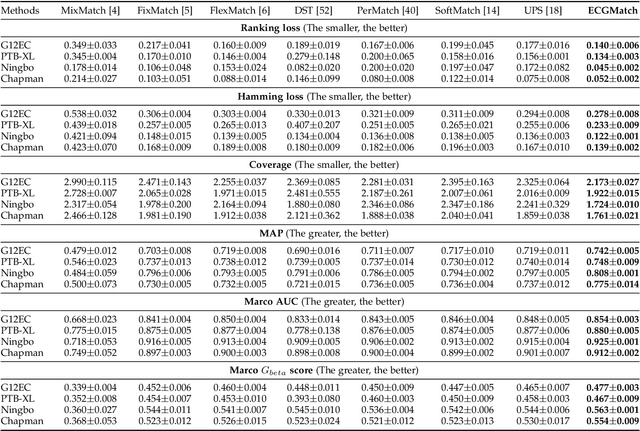
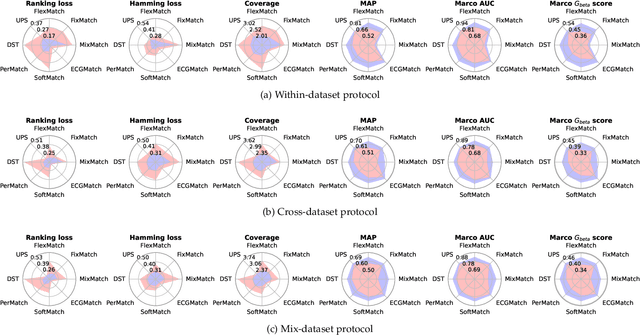
Electrocardiography (ECG) is a non-invasive tool for predicting cardiovascular diseases (CVDs). Current ECG-based diagnosis systems show promising performance owing to the rapid development of deep learning techniques. However, the label scarcity problem, the co-occurrence of multiple CVDs and the poor performance on unseen datasets greatly hinder the widespread application of deep learning-based models. Addressing them in a unified framework remains a significant challenge. To this end, we propose a multi-label semi-supervised model (ECGMatch) to recognize multiple CVDs simultaneously with limited supervision. In the ECGMatch, an ECGAugment module is developed for weak and strong ECG data augmentation, which generates diverse samples for model training. Subsequently, a hyperparameter-efficient framework with neighbor agreement modeling and knowledge distillation is designed for pseudo-label generation and refinement, which mitigates the label scarcity problem. Finally, a label correlation alignment module is proposed to capture the co-occurrence information of different CVDs within labeled samples and propagate this information to unlabeled samples. Extensive experiments on four datasets and three protocols demonstrate the effectiveness and stability of the proposed model, especially on unseen datasets. As such, this model can pave the way for diagnostic systems that achieve robust performance on multi-label CVDs prediction with limited supervision.
Repairing Deep Neural Networks Based on Behavior Imitation
May 05, 2023Zhen Liang, Taoran Wu, Changyuan Zhao, Wanwei Liu, Bai Xue, Wenjing Yang, Ji Wang
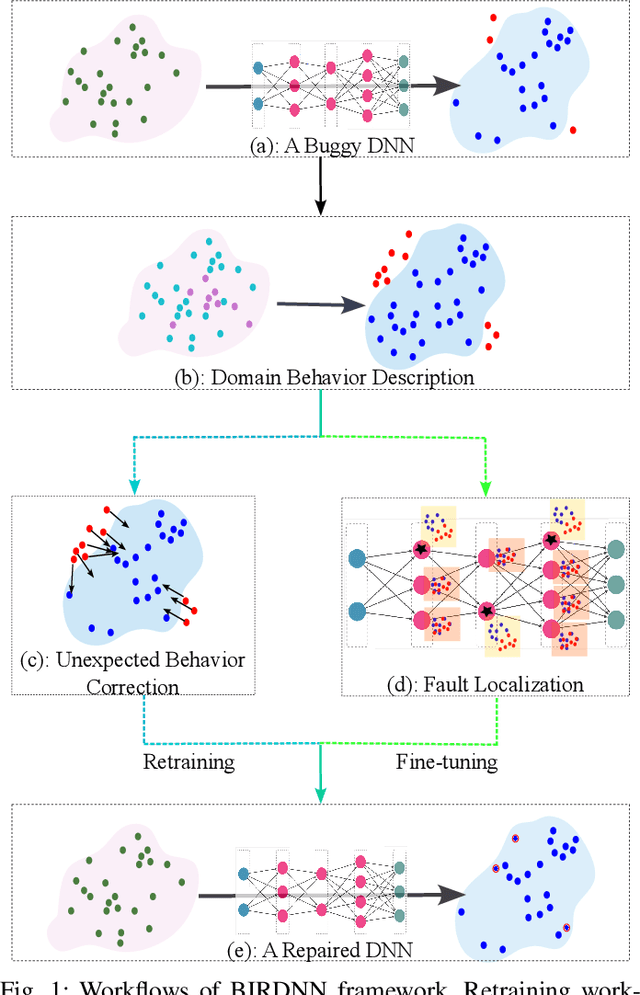
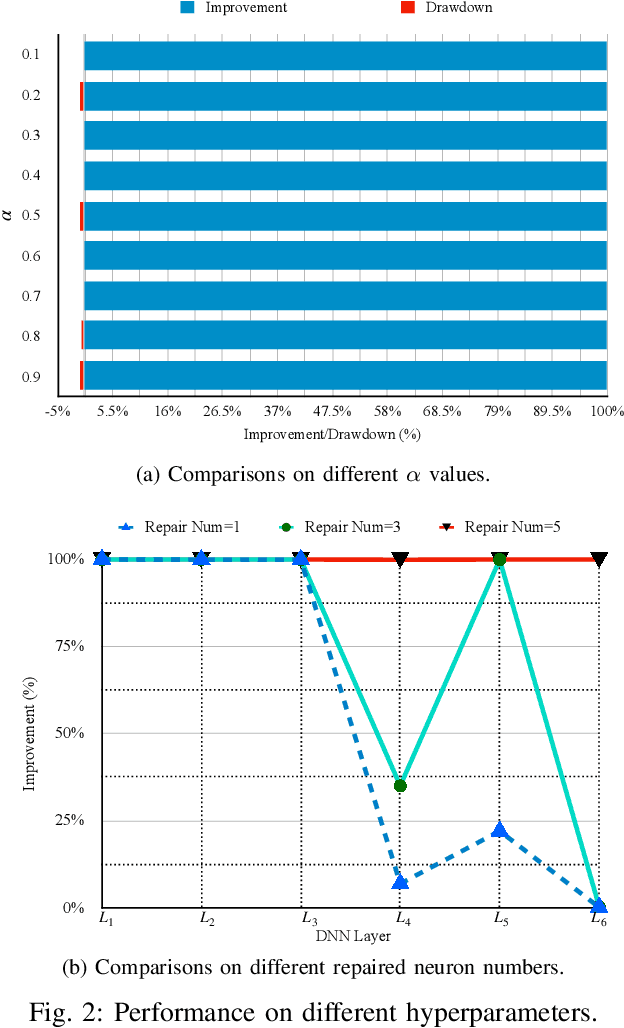
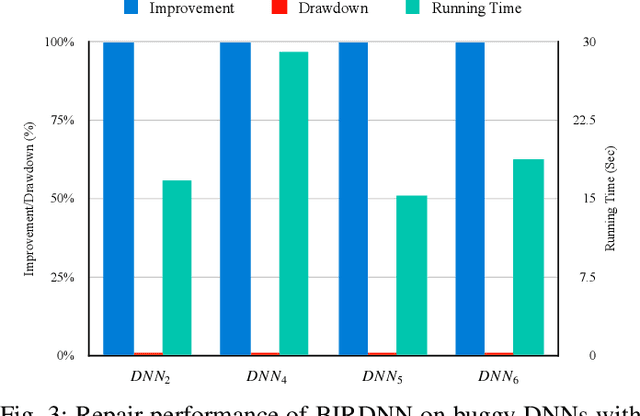
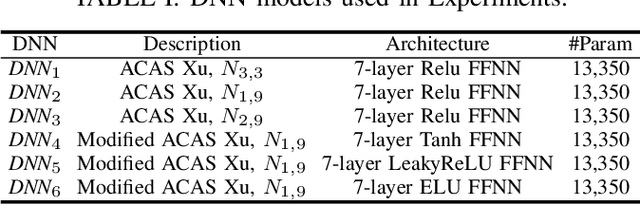
The increasing use of deep neural networks (DNNs) in safety-critical systems has raised concerns about their potential for exhibiting ill-behaviors. While DNN verification and testing provide post hoc conclusions regarding unexpected behaviors, they do not prevent the erroneous behaviors from occurring. To address this issue, DNN repair/patch aims to eliminate unexpected predictions generated by defective DNNs. Two typical DNN repair paradigms are retraining and fine-tuning. However, existing methods focus on the high-level abstract interpretation or inference of state spaces, ignoring the underlying neurons' outputs. This renders patch processes computationally prohibitive and limited to piecewise linear (PWL) activation functions to great extent. To address these shortcomings, we propose a behavior-imitation based repair framework, BIRDNN, which integrates the two repair paradigms for the first time. BIRDNN corrects incorrect predictions of negative samples by imitating the closest expected behaviors of positive samples during the retraining repair procedure. For the fine-tuning repair process, BIRDNN analyzes the behavior differences of neurons on positive and negative samples to identify the most responsible neurons for the erroneous behaviors. To tackle more challenging domain-wise repair problems (DRPs), we synthesize BIRDNN with a domain behavior characterization technique to repair buggy DNNs in a probably approximated correct style. We also implement a prototype tool based on BIRDNN and evaluate it on ACAS Xu DNNs. Our experimental results show that BIRDNN can successfully repair buggy DNNs with significantly higher efficiency than state-of-the-art repair tools. Additionally, BIRDNN is highly compatible with different activation functions.
EEGMatch: Learning with Incomplete Labels for Semi-Supervised EEG-based Cross-Subject Emotion Recognition
Mar 27, 2023Rushuang Zhou, Weishan Ye, Zhiguo Zhang, Yanyang Luo, Li Zhang, Linling Li, Gan Huang, Yining Dong, Yuan-Ting Zhang, Zhen Liang




Electroencephalography (EEG) is an objective tool for emotion recognition and shows promising performance. However, the label scarcity problem is a main challenge in this field, which limits the wide application of EEG-based emotion recognition. In this paper, we propose a novel semi-supervised learning framework (EEGMatch) to leverage both labeled and unlabeled EEG data. First, an EEG-Mixup based data augmentation method is developed to generate more valid samples for model learning. Second, a semi-supervised two-step pairwise learning method is proposed to bridge prototype-wise and instance-wise pairwise learning, where the prototype-wise pairwise learning measures the global relationship between EEG data and the prototypical representation of each emotion class and the instance-wise pairwise learning captures the local intrinsic relationship among EEG data. Third, a semi-supervised multi-domain adaptation is introduced to align the data representation among multiple domains (labeled source domain, unlabeled source domain, and target domain), where the distribution mismatch is alleviated. Extensive experiments are conducted on two benchmark databases (SEED and SEED-IV) under a cross-subject leave-one-subject-out cross-validation evaluation protocol. The results show the proposed EEGmatch performs better than the state-of-the-art methods under different incomplete label conditions (with 6.89% improvement on SEED and 1.44% improvement on SEED-IV), which demonstrates the effectiveness of the proposed EEGMatch in dealing with the label scarcity problem in emotion recognition using EEG signals. The source code is available at https://github.com/KAZABANA/EEGMatch.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge