Generating QM1B with PySCF$_{\text{IPU}}$
Nov 02, 2023Alexander Mathiasen, Hatem Helal, Kerstin Klaser, Paul Balanca, Josef Dean, Carlo Luschi, Dominique Beaini, Andrew Fitzgibbon, Dominic Masters
The emergence of foundation models in Computer Vision and Natural Language Processing have resulted in immense progress on downstream tasks. This progress was enabled by datasets with billions of training examples. Similar benefits are yet to be unlocked for quantum chemistry, where the potential of deep learning is constrained by comparatively small datasets with 100k to 20M training examples. These datasets are limited in size because the labels are computed using the accurate (but computationally demanding) predictions of Density Functional Theory (DFT). Notably, prior DFT datasets were created using CPU supercomputers without leveraging hardware acceleration. In this paper, we take a first step towards utilising hardware accelerators by introducing the data generator PySCF$_{\text{IPU}}$ using Intelligence Processing Units (IPUs). This allowed us to create the dataset QM1B with one billion training examples containing 9-11 heavy atoms. We demonstrate that a simple baseline neural network (SchNet 9M) improves its performance by simply increasing the amount of training data without additional inductive biases. To encourage future researchers to use QM1B responsibly, we highlight several limitations of QM1B and emphasise the low-resolution of our DFT options, which also serves as motivation for even larger, more accurate datasets. Code and dataset are available on Github: http://github.com/graphcore-research/pyscf-ipu
Towards Foundational Models for Molecular Learning on Large-Scale Multi-Task Datasets
Oct 18, 2023Dominique Beaini, Shenyang Huang, Joao Alex Cunha, Zhiyi Li, Gabriela Moisescu-Pareja, Oleksandr Dymov, Samuel Maddrell-Mander, Callum McLean, Frederik Wenkel, Luis Müller, Jama Hussein Mohamud, Ali Parviz, Michael Craig, Michał Koziarski, Jiarui Lu, Zhaocheng Zhu, Cristian Gabellini, Kerstin Klaser, Josef Dean, Cas Wognum, Maciej Sypetkowski, Guillaume Rabusseau, Reihaneh Rabbany, Jian Tang, Christopher Morris, Ioannis Koutis, Mirco Ravanelli, Guy Wolf, Prudencio Tossou, Hadrien Mary, Therence Bois, Andrew Fitzgibbon, Błażej Banaszewski, Chad Martin, Dominic Masters
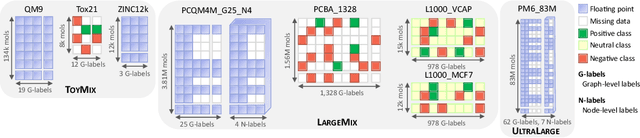
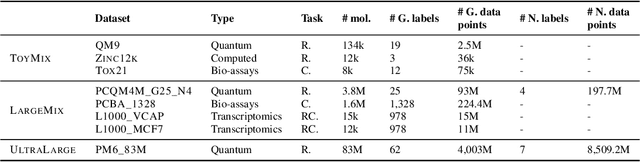
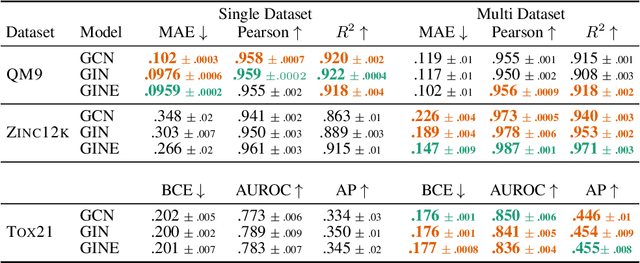
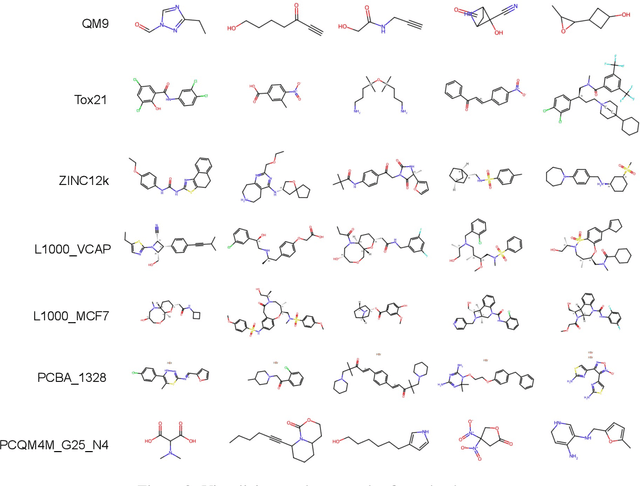
Recently, pre-trained foundation models have enabled significant advancements in multiple fields. In molecular machine learning, however, where datasets are often hand-curated, and hence typically small, the lack of datasets with labeled features, and codebases to manage those datasets, has hindered the development of foundation models. In this work, we present seven novel datasets categorized by size into three distinct categories: ToyMix, LargeMix and UltraLarge. These datasets push the boundaries in both the scale and the diversity of supervised labels for molecular learning. They cover nearly 100 million molecules and over 3000 sparsely defined tasks, totaling more than 13 billion individual labels of both quantum and biological nature. In comparison, our datasets contain 300 times more data points than the widely used OGB-LSC PCQM4Mv2 dataset, and 13 times more than the quantum-only QM1B dataset. In addition, to support the development of foundational models based on our proposed datasets, we present the Graphium graph machine learning library which simplifies the process of building and training molecular machine learning models for multi-task and multi-level molecular datasets. Finally, we present a range of baseline results as a starting point of multi-task and multi-level training on these datasets. Empirically, we observe that performance on low-resource biological datasets show improvement by also training on large amounts of quantum data. This indicates that there may be potential in multi-task and multi-level training of a foundation model and fine-tuning it to resource-constrained downstream tasks.
GPS++: Reviving the Art of Message Passing for Molecular Property Prediction
Feb 06, 2023Dominic Masters, Josef Dean, Kerstin Klaser, Zhiyi Li, Sam Maddrell-Mander, Adam Sanders, Hatem Helal, Deniz Beker, Andrew Fitzgibbon, Shenyang Huang, Ladislav Rampášek, Dominique Beaini

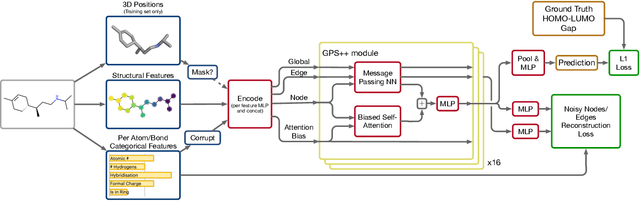
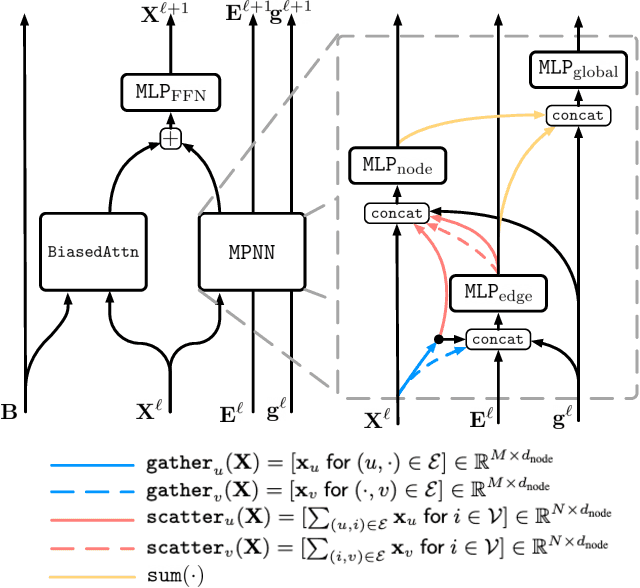
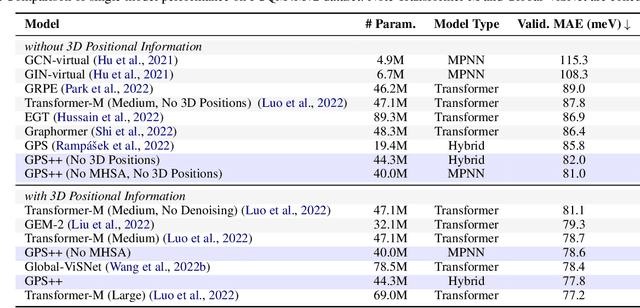
We present GPS++, a hybrid Message Passing Neural Network / Graph Transformer model for molecular property prediction. Our model integrates a well-tuned local message passing component and biased global attention with other key ideas from prior literature to achieve state-of-the-art results on large-scale molecular dataset PCQM4Mv2. Through a thorough ablation study we highlight the impact of individual components and, contrary to expectations set by recent trends, find that nearly all of the model's performance can be maintained without any use of global self-attention. We also show that our approach is significantly more accurate than prior art when 3D positional information is not available.
Efficient and Sound Differentiable Programming in a Functional Array-Processing Language
Dec 20, 2022Amir Shaikhha, Mathieu Huot, Shabnam Ghasemirad, Andrew Fitzgibbon, Simon Peyton Jones, Dimitrios Vytiniotis
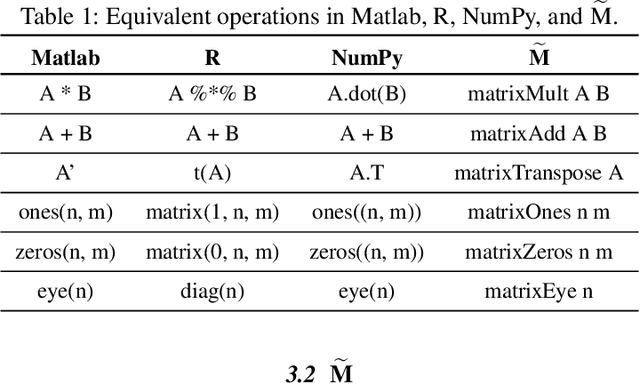
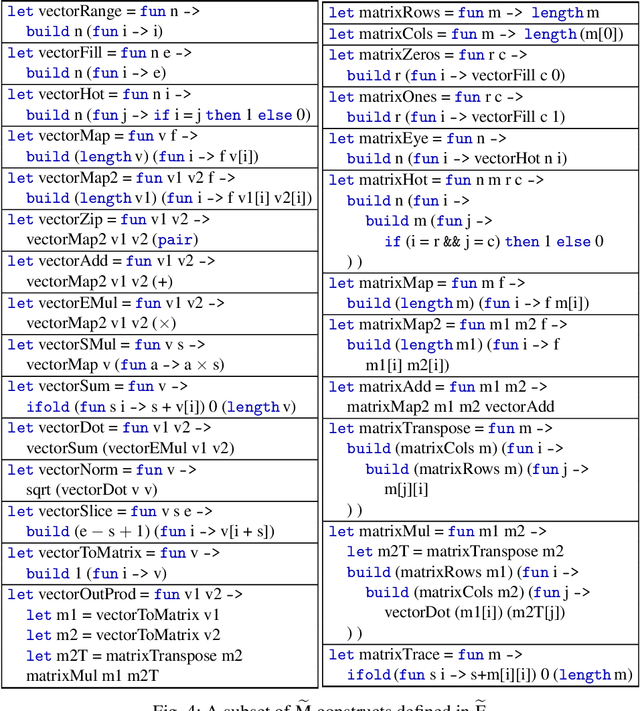
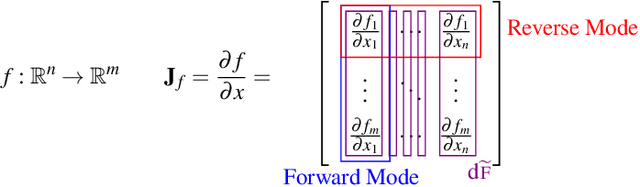



Automatic differentiation (AD) is a technique for computing the derivative of a function represented by a program. This technique is considered as the de-facto standard for computing the differentiation in many machine learning and optimisation software tools. Despite the practicality of this technique, the performance of the differentiated programs, especially for functional languages and in the presence of vectors, is suboptimal. We present an AD system for a higher-order functional array-processing language. The core functional language underlying this system simultaneously supports both source-to-source forward-mode AD and global optimisations such as loop transformations. In combination, gradient computation with forward-mode AD can be as efficient as reverse mode, and the Jacobian matrices required for numerical algorithms such as Gauss-Newton and Levenberg-Marquardt can be efficiently computed.
BESS: Balanced Entity Sampling and Sharing for Large-Scale Knowledge Graph Completion
Nov 22, 2022Alberto Cattaneo, Daniel Justus, Harry Mellor, Douglas Orr, Jerome Maloberti, Zhenying Liu, Thorin Farnsworth, Andrew Fitzgibbon, Blazej Banaszewski, Carlo Luschi
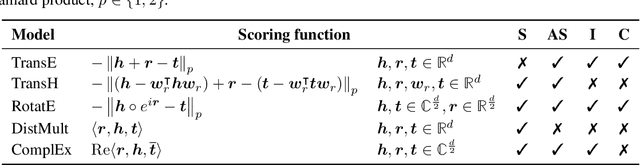

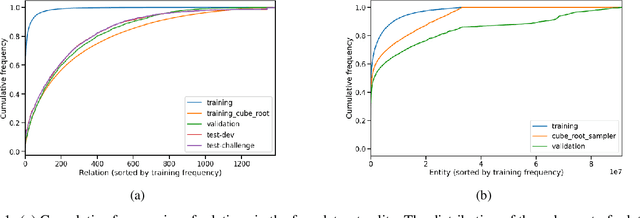

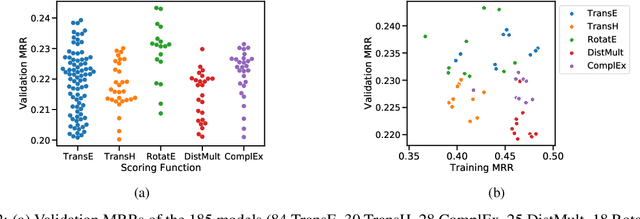
We present the award-winning submission to the WikiKG90Mv2 track of OGB-LSC@NeurIPS 2022. The task is link-prediction on the large-scale knowledge graph WikiKG90Mv2, consisting of 90M+ nodes and 600M+ edges. Our solution uses a diverse ensemble of $85$ Knowledge Graph Embedding models combining five different scoring functions (TransE, TransH, RotatE, DistMult, ComplEx) and two different loss functions (log-sigmoid, sampled softmax cross-entropy). Each individual model is trained in parallel on a Graphcore Bow Pod$_{16}$ using BESS (Balanced Entity Sampling and Sharing), a new distribution framework for KGE training and inference based on balanced collective communications between workers. Our final model achieves a validation MRR of 0.2922 and a test-challenge MRR of 0.2562, winning the first place in the competition. The code is publicly available at: https://github.com/graphcore/distributed-kge-poplar/tree/2022-ogb-submission.
FLAG: Flow-based 3D Avatar Generation from Sparse Observations
Mar 11, 2022Sadegh Aliakbarian, Pashmina Cameron, Federica Bogo, Andrew Fitzgibbon, Thomas J. Cashman
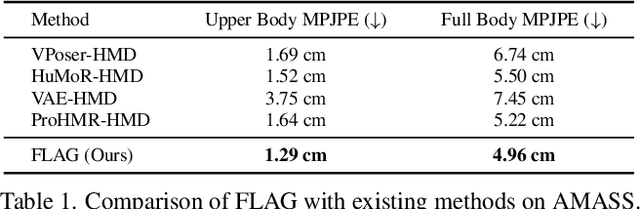
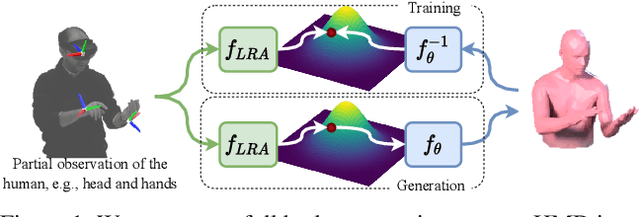

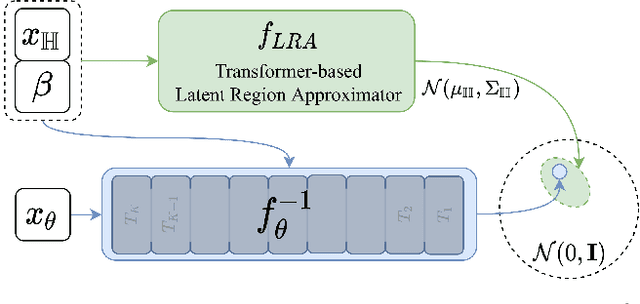
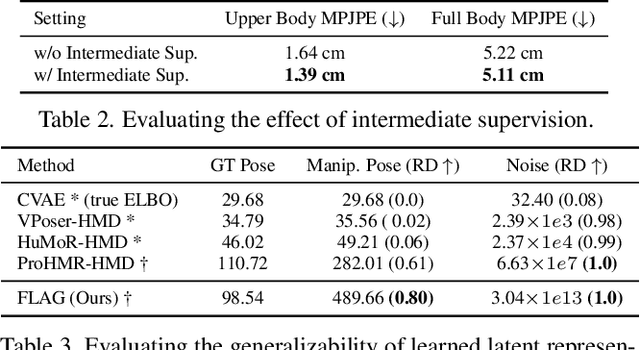
To represent people in mixed reality applications for collaboration and communication, we need to generate realistic and faithful avatar poses. However, the signal streams that can be applied for this task from head-mounted devices (HMDs) are typically limited to head pose and hand pose estimates. While these signals are valuable, they are an incomplete representation of the human body, making it challenging to generate a faithful full-body avatar. We address this challenge by developing a flow-based generative model of the 3D human body from sparse observations, wherein we learn not only a conditional distribution of 3D human pose, but also a probabilistic mapping from observations to the latent space from which we can generate a plausible pose along with uncertainty estimates for the joints. We show that our approach is not only a strong predictive model, but can also act as an efficient pose prior in different optimization settings where a good initial latent code plays a major role.
Who Left the Dogs Out? 3D Animal Reconstruction with Expectation Maximization in the Loop
Jul 21, 2020Benjamin Biggs, Oliver Boyne, James Charles, Andrew Fitzgibbon, Roberto Cipolla
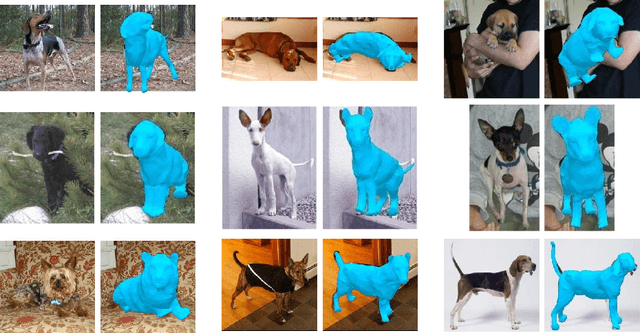

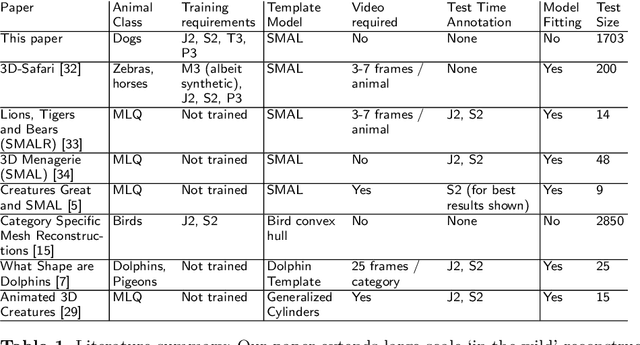
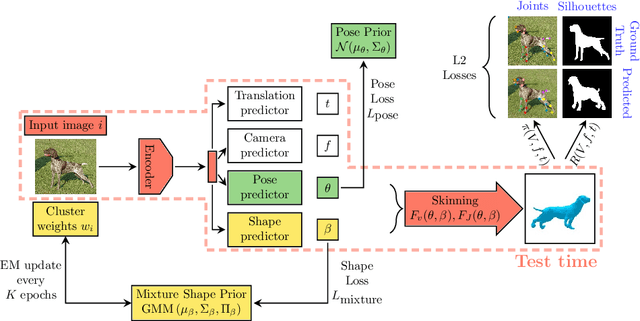
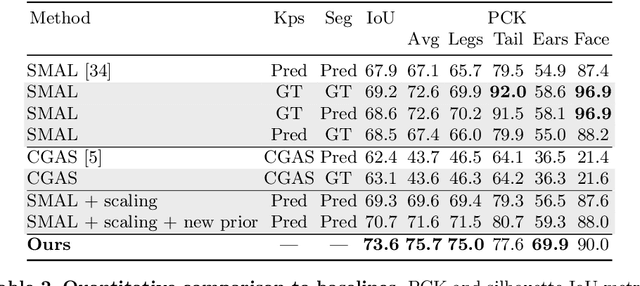
We introduce an automatic, end-to-end method for recovering the 3D pose and shape of dogs from monocular internet images. The large variation in shape between dog breeds, significant occlusion and low quality of internet images makes this a challenging problem. We learn a richer prior over shapes than previous work, which helps regularize parameter estimation. We demonstrate results on the Stanford Dog dataset, an 'in the wild' dataset of 20,580 dog images for which we have collected 2D joint and silhouette annotations to split for training and evaluation. In order to capture the large shape variety of dogs, we show that the natural variation in the 2D dataset is enough to learn a detailed 3D prior through expectation maximization (EM). As a by-product of training, we generate a new parameterized model (including limb scaling) SMBLD which we release alongside our new annotation dataset StanfordExtra to the research community.
Inverse Graphics GAN: Learning to Generate 3D Shapes from Unstructured 2D Data
Feb 28, 2020Sebastian Lunz, Yingzhen Li, Andrew Fitzgibbon, Nate Kushman
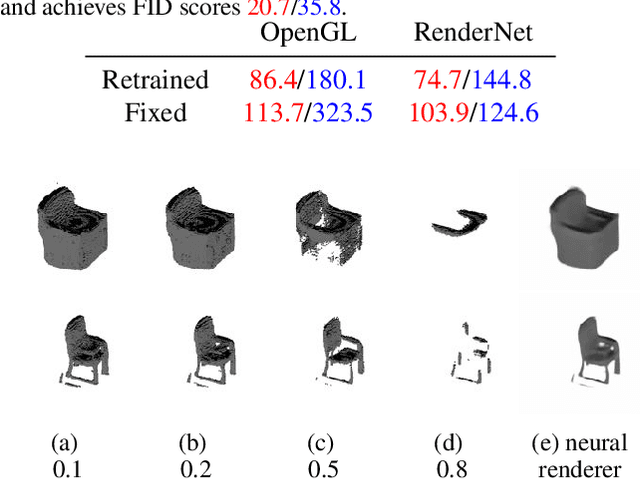

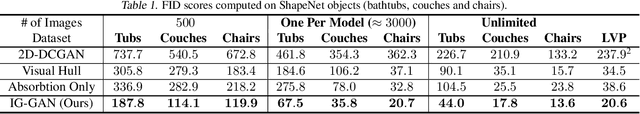
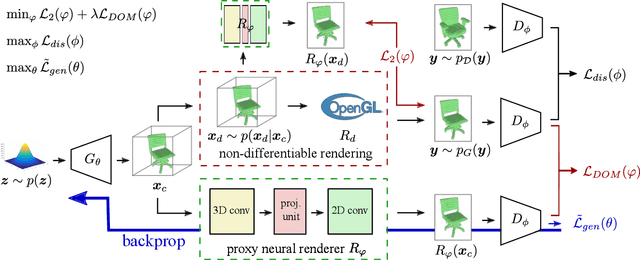

Recent work has shown the ability to learn generative models for 3D shapes from only unstructured 2D images. However, training such models requires differentiating through the rasterization step of the rendering process, therefore past work has focused on developing bespoke rendering models which smooth over this non-differentiable process in various ways. Such models are thus unable to take advantage of the photo-realistic, fully featured, industrial renderers built by the gaming and graphics industry. In this paper we introduce the first scalable training technique for 3D generative models from 2D data which utilizes an off-the-shelf non-differentiable renderer. To account for the non-differentiability, we introduce a proxy neural renderer to match the output of the non-differentiable renderer. We further propose discriminator output matching to ensure that the neural renderer learns to smooth over the rasterization appropriately. We evaluate our model on images rendered from our generated 3D shapes, and show that our model can consistently learn to generate better shapes than existing models when trained with exclusively unstructured 2D images.
Creatures great and SMAL: Recovering the shape and motion of animals from video
Nov 14, 2018Benjamin Biggs, Thomas Roddick, Andrew Fitzgibbon, Roberto Cipolla
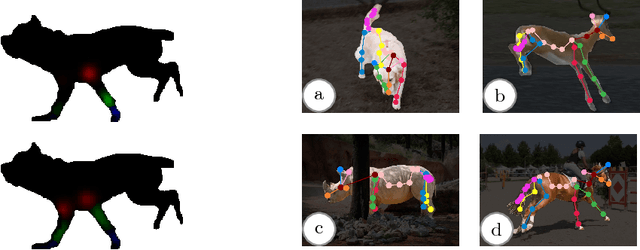
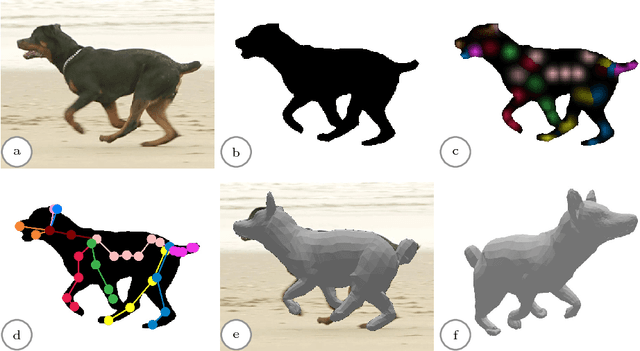
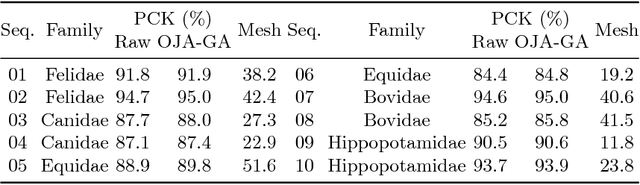

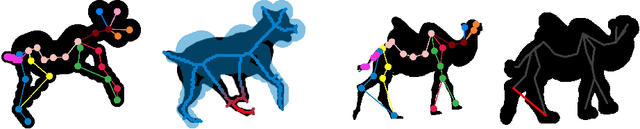
We present a system to recover the 3D shape and motion of a wide variety of quadrupeds from video. The system comprises a machine learning front-end which predicts candidate 2D joint positions, a discrete optimization which finds kinematically plausible joint correspondences, and an energy minimization stage which fits a detailed 3D model to the image. In order to overcome the limited availability of motion capture training data from animals, and the difficulty of generating realistic synthetic training images, the system is designed to work on silhouette data. The joint candidate predictor is trained on synthetically generated silhouette images, and at test time, deep learning methods or standard video segmentation tools are used to extract silhouettes from real data. The system is tested on animal videos from several species, and shows accurate reconstructions of 3D shape and pose.
A Benchmark of Selected Algorithmic Differentiation Tools on Some Problems in Computer Vision and Machine Learning
Jul 26, 2018Filip Šrajer, Zuzana Kukelova, Andrew Fitzgibbon
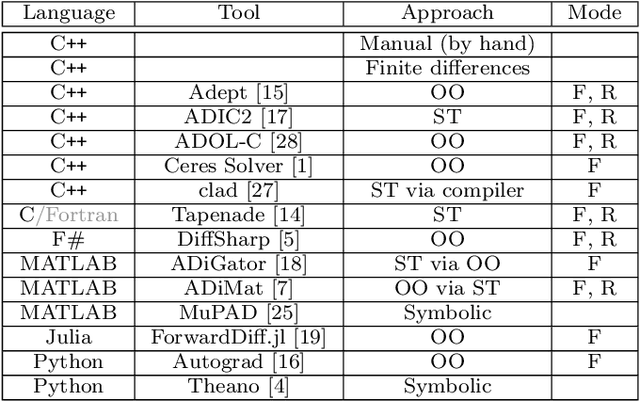
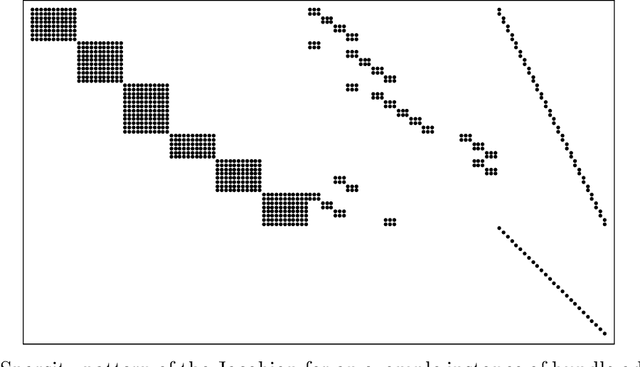
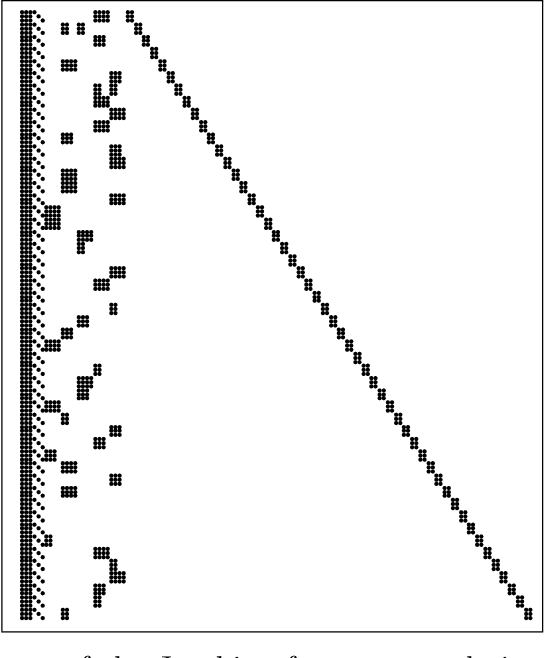
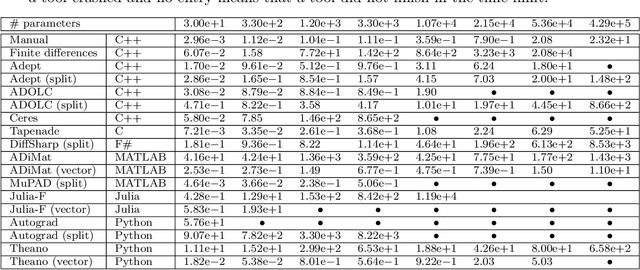
Algorithmic differentiation (AD) allows exact computation of derivatives given only an implementation of an objective function. Although many AD tools are available, a proper and efficient implementation of AD methods is not straightforward. The existing tools are often too different to allow for a general test suite. In this paper, we compare fifteen ways of computing derivatives including eleven automatic differentiation tools implementing various methods and written in various languages (C++, F#, MATLAB, Julia and Python), two symbolic differentiation tools, finite differences, and hand-derived computation. We look at three objective functions from computer vision and machine learning. These objectives are for the most part simple, in the sense that no iterative loops are involved, and conditional statements are encapsulated in functions such as {\tt abs} or {\tt logsumexp}. However, it is important for the success of algorithmic differentiation that such `simple' objective functions are handled efficiently, as so many problems in computer vision and machine learning are of this form. Of course, our results depend on programmer skill, and familiarity with the tools. However, we contend that this paper presents an important datapoint: a skilled programmer devoting roughly a week to each tool produced the timings we present. We have made our implementations available as open source to allow the community to replicate and update these benchmarks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge