KIWI: A Dataset of Knowledge-Intensive Writing Instructions for Answering Research Questions
Mar 06, 2024Fangyuan Xu, Kyle Lo, Luca Soldaini, Bailey Kuehl, Eunsol Choi, David Wadden
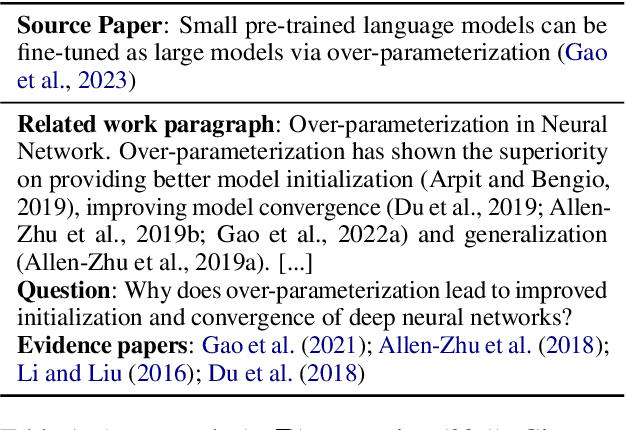
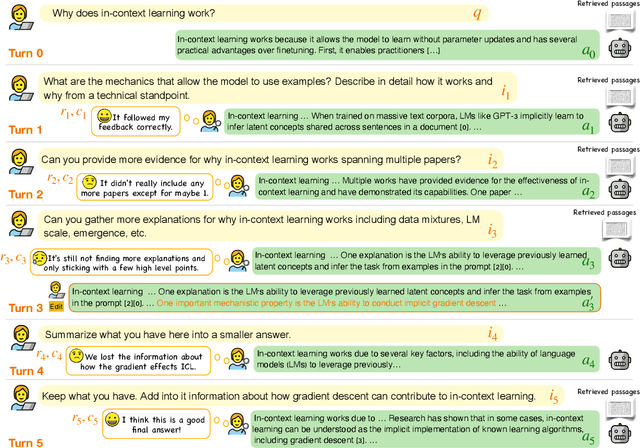

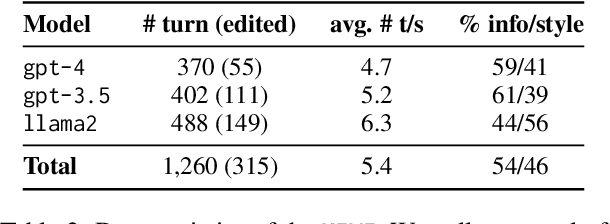
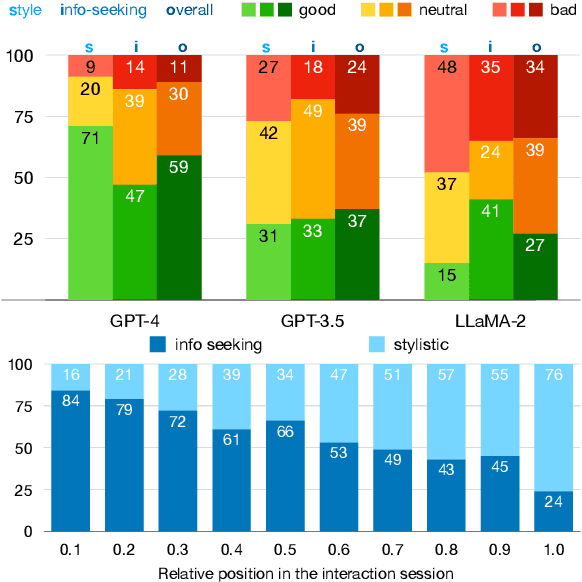
Large language models (LLMs) adapted to follow user instructions are now widely deployed as conversational agents. In this work, we examine one increasingly common instruction-following task: providing writing assistance to compose a long-form answer. To evaluate the capabilities of current LLMs on this task, we construct KIWI, a dataset of knowledge-intensive writing instructions in the scientific domain. Given a research question, an initial model-generated answer and a set of relevant papers, an expert annotator iteratively issues instructions for the model to revise and improve its answer. We collect 1,260 interaction turns from 234 interaction sessions with three state-of-the-art LLMs. Each turn includes a user instruction, a model response, and a human evaluation of the model response. Through a detailed analysis of the collected responses, we find that all models struggle to incorporate new information into an existing answer, and to perform precise and unambiguous edits. Further, we find that models struggle to judge whether their outputs successfully followed user instructions, with accuracy at least 10 points short of human agreement. Our findings indicate that KIWI will be a valuable resource to measure progress and improve LLMs' instruction-following capabilities for knowledge intensive writing tasks.
CARE: Extracting Experimental Findings From Clinical Literature
Nov 16, 2023Aakanksha Naik, Bailey Kuehl, Erin Bransom, Doug Downey, Tom Hope
Extracting fine-grained experimental findings from literature can provide massive utility for scientific applications. Prior work has focused on developing annotation schemas and datasets for limited aspects of this problem, leading to simpler information extraction datasets which do not capture the real-world complexity and nuance required for this task. Focusing on biomedicine, this work presents CARE (Clinical Aggregation-oriented Result Extraction) -- a new IE dataset for the task of extracting clinical findings. We develop a new annotation schema capturing fine-grained findings as n-ary relations between entities and attributes, which includes phenomena challenging for current IE systems such as discontinuous entity spans, nested relations, and variable arity n-ary relations. Using this schema, we collect extensive annotations for 700 abstracts from two sources: clinical trials and case reports. We also benchmark the performance of various state-of-the-art IE systems on our dataset, including extractive models and generative LLMs in fully supervised and limited data settings. Our results demonstrate the difficulty of our dataset -- even SOTA models such as GPT4 struggle, particularly on relation extraction. We release our annotation schema and CARE to encourage further research on extracting and aggregating scientific findings from literature.
ARIES: A Corpus of Scientific Paper Edits Made in Response to Peer Reviews
Jun 21, 2023Mike D'Arcy, Alexis Ross, Erin Bransom, Bailey Kuehl, Jonathan Bragg, Tom Hope, Doug Downey
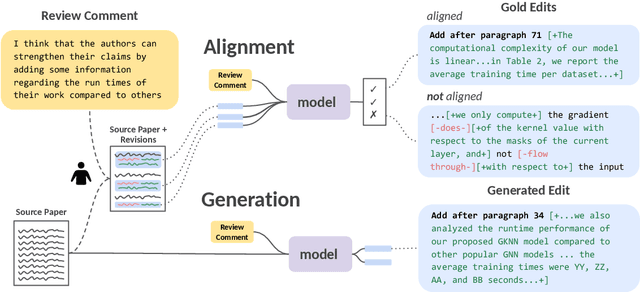
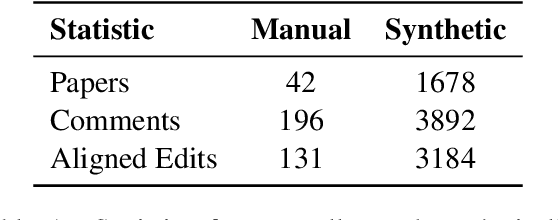

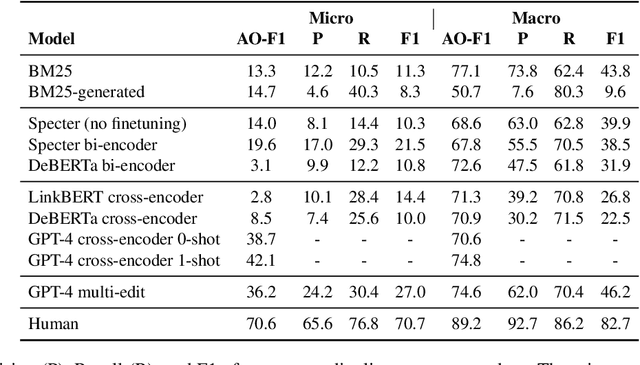
Revising scientific papers based on peer feedback is a challenging task that requires not only deep scientific knowledge and reasoning, but also the ability to recognize the implicit requests in high-level feedback and to choose the best of many possible ways to update the manuscript in response. We introduce this task for large language models and release ARIES, a dataset of review comments and their corresponding paper edits, to enable training and evaluating models. We study two versions of the task: comment-edit alignment and edit generation, and evaluate several baselines, including GPT-4. We find that models struggle even to identify the edits that correspond to a comment, especially in cases where the comment is phrased in an indirect way or where the edit addresses the spirit of a comment but not the precise request. When tasked with generating edits, GPT-4 often succeeds in addressing comments on a surface level, but it rigidly follows the wording of the feedback rather than the underlying intent, and includes fewer technical details than human-written edits. We hope that our formalization, dataset, and analysis will form a foundation for future work in this area.
S2abEL: A Dataset for Entity Linking from Scientific Tables
Apr 30, 2023Yuze Lou, Bailey Kuehl, Erin Bransom, Sergey Feldman, Aakanksha Naik, Doug Downey
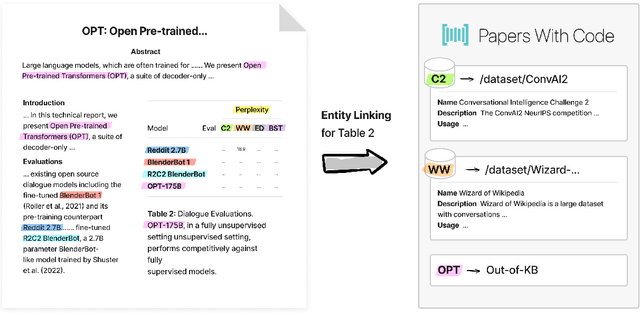
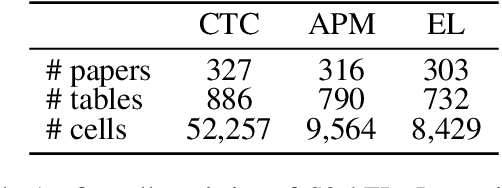
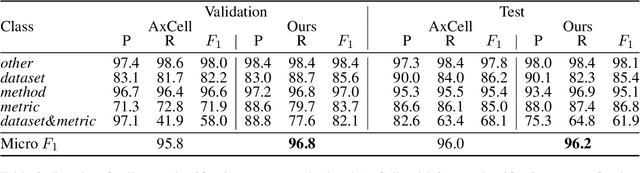
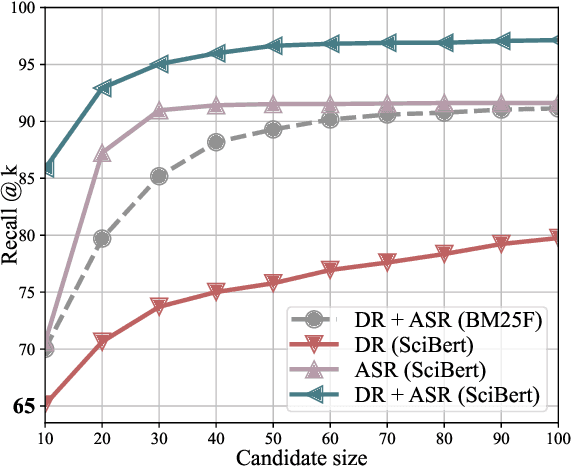
Entity linking (EL) is the task of linking a textual mention to its corresponding entry in a knowledge base, and is critical for many knowledge-intensive NLP applications. When applied to tables in scientific papers, EL is a step toward large-scale scientific knowledge bases that could enable advanced scientific question answering and analytics. We present the first dataset for EL in scientific tables. EL for scientific tables is especially challenging because scientific knowledge bases can be very incomplete, and disambiguating table mentions typically requires understanding the papers's tet in addition to the table. Our dataset, S2abEL, focuses on EL in machine learning results tables and includes hand-labeled cell types, attributed sources, and entity links from the PaperswithCode taxonomy for 8,429 cells from 732 tables. We introduce a neural baseline method designed for EL on scientific tables containing many out-of-knowledge-base mentions, and show that it significantly outperforms a state-of-the-art generic table EL method. The best baselines fall below human performance, and our analysis highlights avenues for improvement.
The Semantic Reader Project: Augmenting Scholarly Documents through AI-Powered Interactive Reading Interfaces
Mar 25, 2023Kyle Lo, Joseph Chee Chang, Andrew Head, Jonathan Bragg, Amy X. Zhang, Cassidy Trier, Chloe Anastasiades, Tal August, Russell Authur, Danielle Bragg, Erin Bransom, Isabel Cachola, Stefan Candra, Yoganand Chandrasekhar, Yen-Sung Chen, Evie Yu-Yen Cheng, Yvonne Chou, Doug Downey, Rob Evans, Raymond Fok, Fangzhou Hu, Regan Huff, Dongyeop Kang, Tae Soo Kim, Rodney Kinney, Aniket Kittur, Hyeonsu Kang, Egor Klevak, Bailey Kuehl, Michael Langan, Matt Latzke, Jaron Lochner, Kelsey MacMillan, Eric Marsh, Tyler Murray, Aakanksha Naik, Ngoc-Uyen Nguyen, Srishti Palani, Soya Park, Caroline Paulic, Napol Rachatasumrit, Smita Rao, Paul Sayre, Zejiang Shen, Pao Siangliulue, Luca Soldaini, Huy Tran, Madeleine van Zuylen, Lucy Lu Wang, Christopher Wilhelm, Caroline Wu, Jiangjiang Yang, Angele Zamarron, Marti A. Hearst, Daniel S. Weld
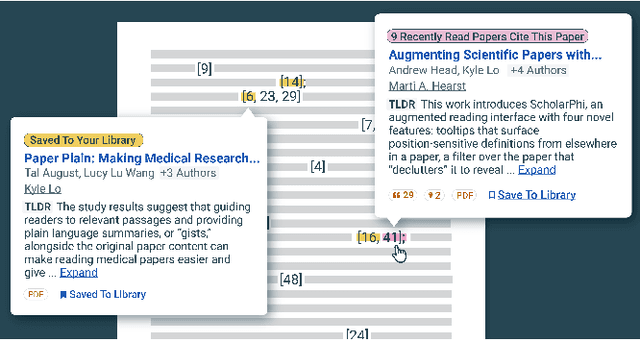


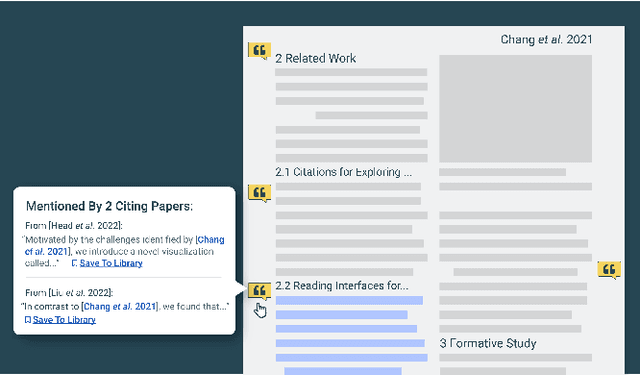
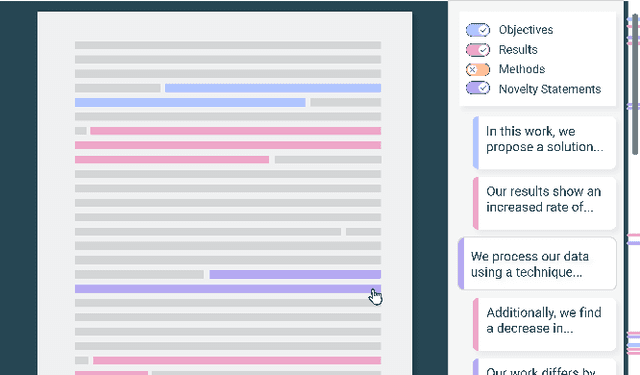
Scholarly publications are key to the transfer of knowledge from scholars to others. However, research papers are information-dense, and as the volume of the scientific literature grows, the need for new technology to support the reading process grows. In contrast to the process of finding papers, which has been transformed by Internet technology, the experience of reading research papers has changed little in decades. The PDF format for sharing research papers is widely used due to its portability, but it has significant downsides including: static content, poor accessibility for low-vision readers, and difficulty reading on mobile devices. This paper explores the question "Can recent advances in AI and HCI power intelligent, interactive, and accessible reading interfaces -- even for legacy PDFs?" We describe the Semantic Reader Project, a collaborative effort across multiple institutions to explore automatic creation of dynamic reading interfaces for research papers. Through this project, we've developed ten research prototype interfaces and conducted usability studies with more than 300 participants and real-world users showing improved reading experiences for scholars. We've also released a production reading interface for research papers that will incorporate the best features as they mature. We structure this paper around challenges scholars and the public face when reading research papers -- Discovery, Efficiency, Comprehension, Synthesis, and Accessibility -- and present an overview of our progress and remaining open challenges.
LongEval: Guidelines for Human Evaluation of Faithfulness in Long-form Summarization
Jan 30, 2023Kalpesh Krishna, Erin Bransom, Bailey Kuehl, Mohit Iyyer, Pradeep Dasigi, Arman Cohan, Kyle Lo
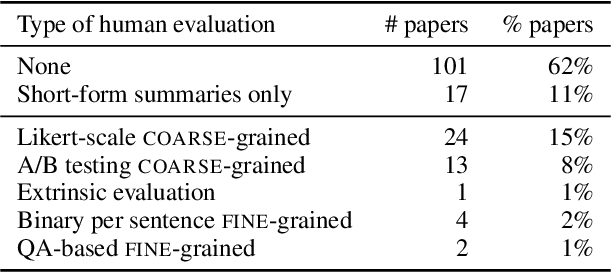

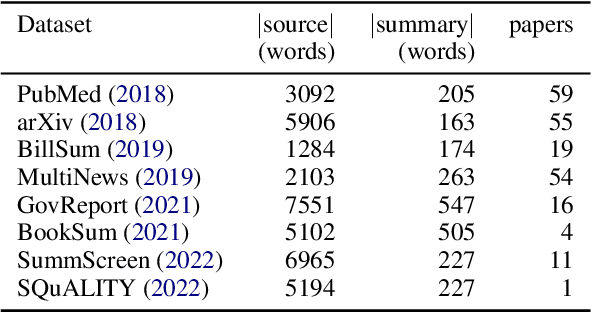

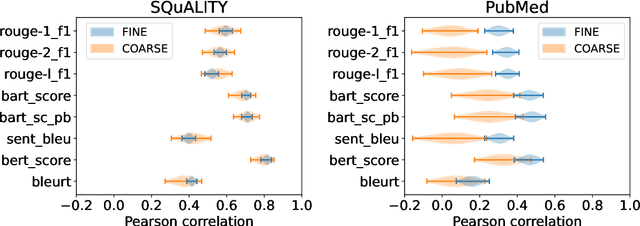
While human evaluation remains best practice for accurately judging the faithfulness of automatically-generated summaries, few solutions exist to address the increased difficulty and workload when evaluating long-form summaries. Through a survey of 162 papers on long-form summarization, we first shed light on current human evaluation practices surrounding long-form summaries. We find that 73% of these papers do not perform any human evaluation on model-generated summaries, while other works face new difficulties that manifest when dealing with long documents (e.g., low inter-annotator agreement). Motivated by our survey, we present LongEval, a set of guidelines for human evaluation of faithfulness in long-form summaries that addresses the following challenges: (1) How can we achieve high inter-annotator agreement on faithfulness scores? (2) How can we minimize annotator workload while maintaining accurate faithfulness scores? and (3) Do humans benefit from automated alignment between summary and source snippets? We deploy LongEval in annotation studies on two long-form summarization datasets in different domains (SQuALITY and PubMed), and we find that switching to a finer granularity of judgment (e.g., clause-level) reduces inter-annotator variance in faithfulness scores (e.g., std-dev from 18.5 to 6.8). We also show that scores from a partial annotation of fine-grained units highly correlates with scores from a full annotation workload (0.89 Kendall's tau using 50% judgments). We release our human judgments, annotation templates, and our software as a Python library for future research.
The Semantic Scholar Open Data Platform
Jan 24, 2023Rodney Kinney, Chloe Anastasiades, Russell Authur, Iz Beltagy, Jonathan Bragg, Alexandra Buraczynski, Isabel Cachola, Stefan Candra, Yoganand Chandrasekhar, Arman Cohan, Miles Crawford, Doug Downey, Jason Dunkelberger, Oren Etzioni, Rob Evans, Sergey Feldman, Joseph Gorney, David Graham, Fangzhou Hu, Regan Huff, Daniel King, Sebastian Kohlmeier, Bailey Kuehl, Michael Langan, Daniel Lin, Haokun Liu, Kyle Lo, Jaron Lochner, Kelsey MacMillan, Tyler Murray, Chris Newell, Smita Rao, Shaurya Rohatgi, Paul Sayre, Zejiang Shen, Amanpreet Singh, Luca Soldaini, Shivashankar Subramanian, Amber Tanaka, Alex D. Wade, Linda Wagner, Lucy Lu Wang, Chris Wilhelm, Caroline Wu, Jiangjiang Yang, Angele Zamarron, Madeleine Van Zuylen, Daniel S. Weld
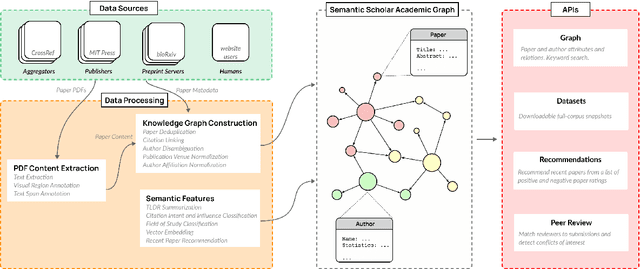
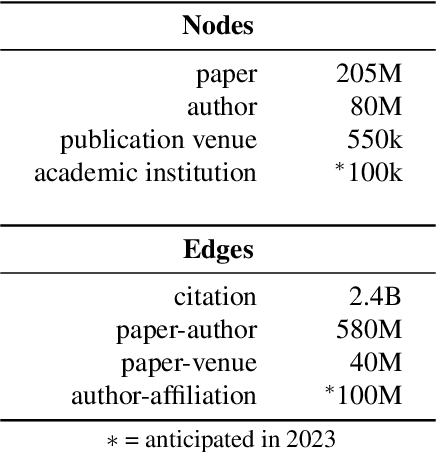
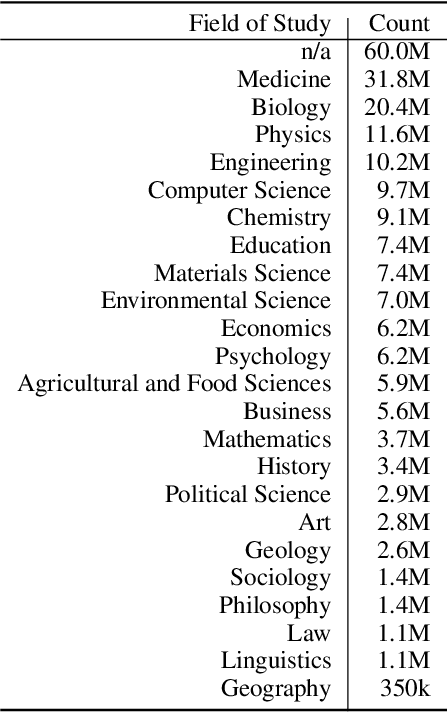
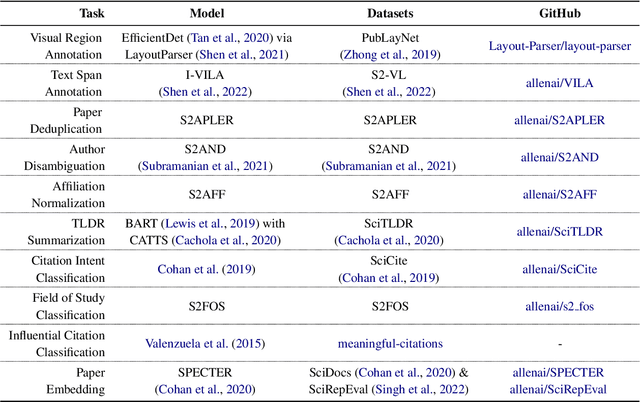
The volume of scientific output is creating an urgent need for automated tools to help scientists keep up with developments in their field. Semantic Scholar (S2) is an open data platform and website aimed at accelerating science by helping scholars discover and understand scientific literature. We combine public and proprietary data sources using state-of-the-art techniques for scholarly PDF content extraction and automatic knowledge graph construction to build the Semantic Scholar Academic Graph, the largest open scientific literature graph to-date, with 200M+ papers, 80M+ authors, 550M+ paper-authorship edges, and 2.4B+ citation edges. The graph includes advanced semantic features such as structurally parsed text, natural language summaries, and vector embeddings. In this paper, we describe the components of the S2 data processing pipeline and the associated APIs offered by the platform. We will update this living document to reflect changes as we add new data offerings and improve existing services.
SciFact-Open: Towards open-domain scientific claim verification
Oct 25, 2022David Wadden, Kyle Lo, Bailey Kuehl, Arman Cohan, Iz Beltagy, Lucy Lu Wang, Hannaneh Hajishirzi
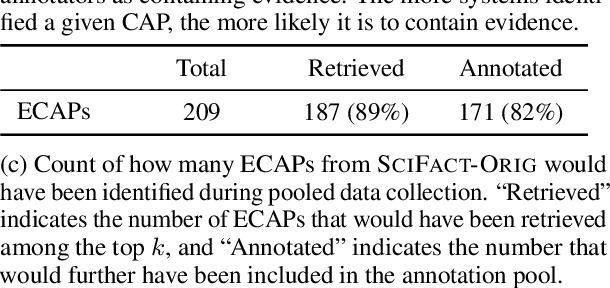
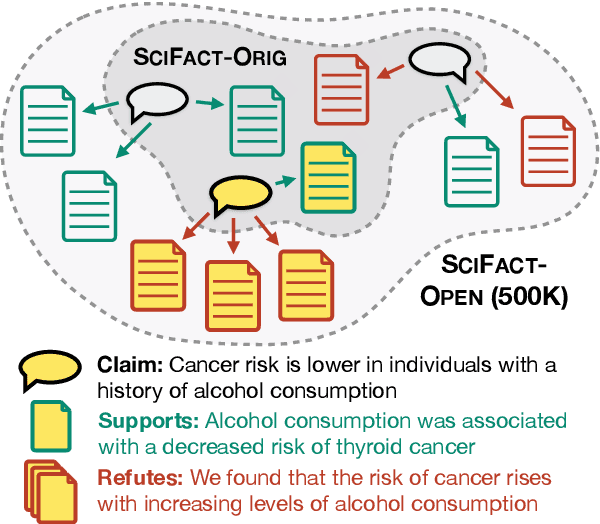
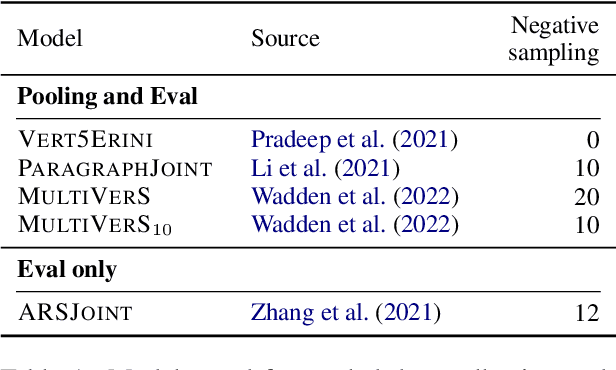


While research on scientific claim verification has led to the development of powerful systems that appear to approach human performance, these approaches have yet to be tested in a realistic setting against large corpora of scientific literature. Moving to this open-domain evaluation setting, however, poses unique challenges; in particular, it is infeasible to exhaustively annotate all evidence documents. In this work, we present SciFact-Open, a new test collection designed to evaluate the performance of scientific claim verification systems on a corpus of 500K research abstracts. Drawing upon pooling techniques from information retrieval, we collect evidence for scientific claims by pooling and annotating the top predictions of four state-of-the-art scientific claim verification models. We find that systems developed on smaller corpora struggle to generalize to SciFact-Open, exhibiting performance drops of at least 15 F1. In addition, analysis of the evidence in SciFact-Open reveals interesting phenomena likely to appear when claim verification systems are deployed in practice, e.g., cases where the evidence supports only a special case of the claim. Our dataset is available at https://github.com/dwadden/scifact-open.
ACCoRD: A Multi-Document Approach to Generating Diverse Descriptions of Scientific Concepts
May 14, 2022Sonia K. Murthy, Kyle Lo, Daniel King, Chandra Bhagavatula, Bailey Kuehl, Sophie Johnson, Jonathan Borchardt, Daniel S. Weld, Tom Hope, Doug Downey
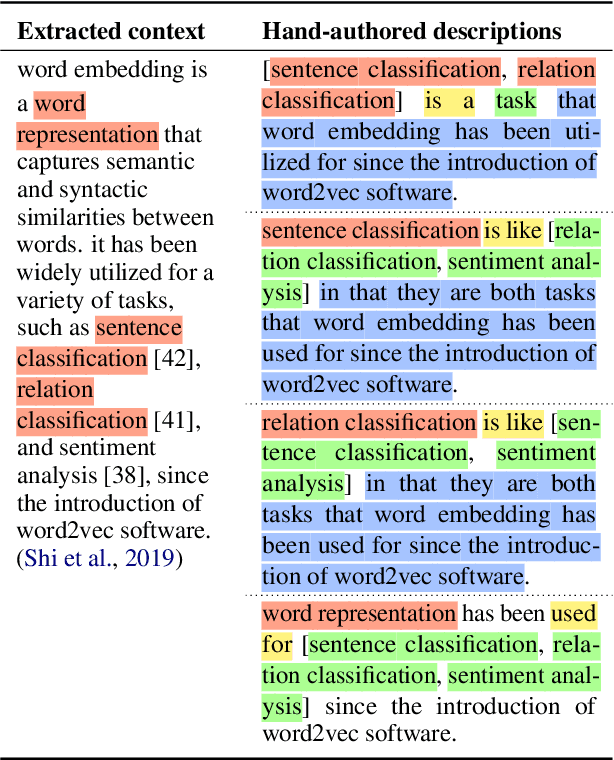
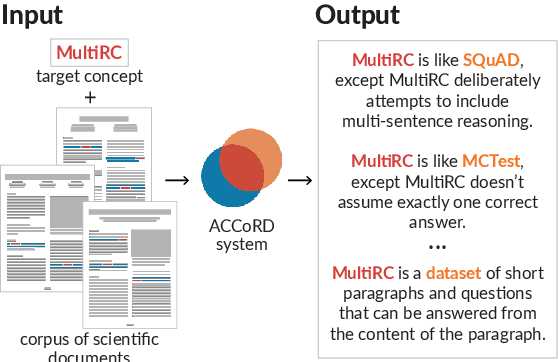


Systems that can automatically define unfamiliar terms hold the promise of improving the accessibility of scientific texts, especially for readers who may lack prerequisite background knowledge. However, current systems assume a single "best" description per concept, which fails to account for the many potentially useful ways a concept can be described. We present ACCoRD, an end-to-end system tackling the novel task of generating sets of descriptions of scientific concepts. Our system takes advantage of the myriad ways a concept is mentioned across the scientific literature to produce distinct, diverse descriptions of target scientific concepts in terms of different reference concepts. To support research on the task, we release an expert-annotated resource, the ACCoRD corpus, which includes 1,275 labeled contexts and 1,787 hand-authored concept descriptions. We conduct a user study demonstrating that (1) users prefer descriptions produced by our end-to-end system, and (2) users prefer multiple descriptions to a single "best" description.
Generating Scientific Claims for Zero-Shot Scientific Fact Checking
Mar 24, 2022Dustin Wright, David Wadden, Kyle Lo, Bailey Kuehl, Arman Cohan, Isabelle Augenstein, Lucy Lu Wang
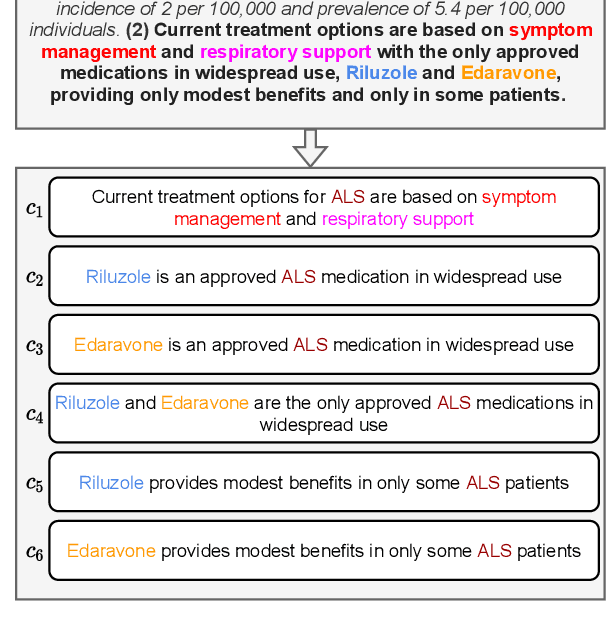
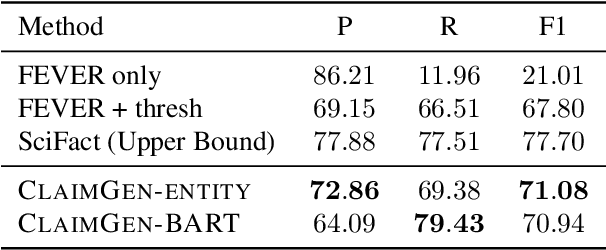
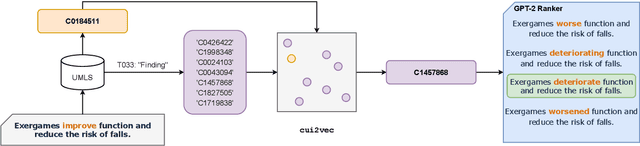

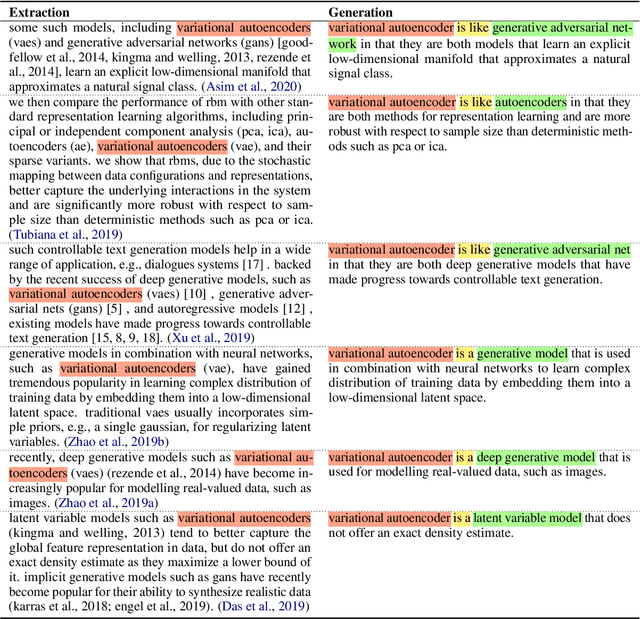
Automated scientific fact checking is difficult due to the complexity of scientific language and a lack of significant amounts of training data, as annotation requires domain expertise. To address this challenge, we propose scientific claim generation, the task of generating one or more atomic and verifiable claims from scientific sentences, and demonstrate its usefulness in zero-shot fact checking for biomedical claims. We propose CLAIMGEN-BART, a new supervised method for generating claims supported by the literature, as well as KBIN, a novel method for generating claim negations. Additionally, we adapt an existing unsupervised entity-centric method of claim generation to biomedical claims, which we call CLAIMGEN-ENTITY. Experiments on zero-shot fact checking demonstrate that both CLAIMGEN-ENTITY and CLAIMGEN-BART, coupled with KBIN, achieve up to 90% performance of fully supervised models trained on manually annotated claims and evidence. A rigorous evaluation study demonstrates significant improvement in generated claim and negation quality over existing baselines
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge